-
PDF
- Split View
-
Views
-
Cite
Cite
Laura Orellana, Johan Gustavsson, Cathrine Bergh, Ozge Yoluk, Erik Lindahl, eBDIMS server: protein transition pathways with ensemble analysis in 2D-motion spaces, Bioinformatics, Volume 35, Issue 18, September 2019, Pages 3505–3507, https://doi.org/10.1093/bioinformatics/btz104
Close - Share Icon Share
Abstract
Understanding how proteins transition between different conformers, and how conformers relate to each other in terms of structure and function, is not trivial. Here, we present an online tool for transition pathway generation between two protein conformations using Elastic Network Driven Brownian Dynamics Importance Sampling, a coarse-grained simulation algorithm, which spontaneously predicts transition intermediates trapped experimentally. In addition to path-generation, the server provides an interactive 2D-motion landscape graphical representation of the transitions or any additional conformers to explore their structural relationships.
eBDIMS is available online: http://ebdims.biophysics.se/ or as standalone software: https://github.com/laura-orellana/eBDIMS, https://github.com/cabergh/eBDIMS.
Supplementary data are available at Bioinformatics online.
1 Introduction
Protein conformational changes are essential to understand biological function, but are difficult to tackle both experimentally and computationally. Cryo-EM and X-ray techniques are however providing a wealth of new structures trapped in different conformations. Yet, understanding how different conformers relate to each other is often non-trivial and requires information on the actual pathways connecting experimental end-states. Public web servers that calculate transitions are therefore common (Zheng and Wen, 2017) but at most evaluate the generated pathways in terms of stereochemical quality. Typically, validation of the biological feasibility of in silico pathways is complex (Weiss and Levitt, 2009), and relies on ad hoc reaction coordinates (e.g. distances, angles, etc.). We have demonstrated that minimal but ‘structurally rich’ X-ray ensembles provide Principal Components (PCs) which are powerful intrinsic coordinates for path-validation (Orellana et al., 2016). Using projections onto the PC-space (Fig. 1), we showed that Elastic Network Driven Brownian Dynamics Importance Sampling (eBDIMS) coarse-grained simulations generate transitions along X-ray encoded motions, spontaneously predicting multiple on-pathway experimental intermediates. Unlike other methods, eBDIMS tracks both forward and reverse paths, which together define a low-energy area sampled by costly molecular dynamics (MD) simulations. Here we present an online tool for eBDIMS-pathways generation between two user-defined end-states, and its integrated 2D-motion projection, either in PC-space if an ensemble is provided, or by default in normal mode (NM) space, to facilitate path and conformer analysis.
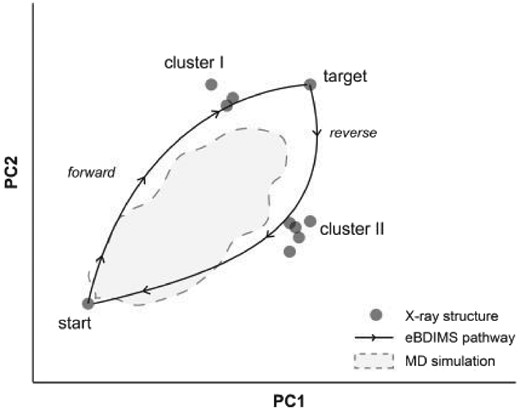
Scheme of eBDIMS path-projections onto the PC-motion space. eBDIMS generates two transitions between the start and target structures (forward and reverse). Projection onto the PCs of an experimental X-ray ensemble (dots) shows path asymmetry and how they visit different on-pathway intermediate clusters. Note that the area sampled by MD typically is contained within the low-energy area defined by both paths
2 Materials and methods
2.1 The eBDIMS algorithm
Until convergence (99.5%) or maximal time (4 h) is reached. Further ED-ENM/eBDIMS details are described elsewhere (Orellana et al., 2016).
2.2 Pathway projections onto 2D-motion spaces
3 Implementation
The eBDIMS code is written in C and parallelized with OpenMP for shared memory multi-threading. Jobs are scheduled linearly and executed on a server equipped with Quad-Core AMD Opteron 64-bit Processors 2374 HE at 2.20 GHz (8 cores) and 16 GB RAM. Interactive 2D- and 3D-plots are generated in the browser with d3.js and NGL viewer (Rose and Hildebrand, 2015), respectively. See scheme of the workflow in Supplementary Figure S2. Documentation, tutorials and examples including those in Orellana et al. (2016) with sample files (Supplementary Table S1) are provided online.
Input data
End-state structures (required): two conformations (referred to as start and target, respectively) are uploaded as coordinate files in strict PDB-format, or retrieved by their IDs (e.g. 4NPQ: A, chain A of PDB 4NPQ).
Conformational ensemble (optional): one file with coordinate sets in PDB-format and delimited by MODEL and ENDMDL labels that strictly match the reference end-states in residue number.
Output results
Successful job submission redirects to a unique result page displaying job status. Active jobs can be monitored in real-time charts reporting the transition progress (%) and RMSD to the target (Å). A typical run takes from minutes to hours depending on system size and transition complexity. Finished jobs present interactive 3D time-resolved molecular representations linked to PC or NM 2D-spaces that show eBDIMS transitions (forward: start → target; reverse: target → start) and all input structures.
Funding
This work was supported by the Sven and Lilly Lawskis Foundation (to L.O.); the Knut and Alice Wallenberg foundation (KAW), Vetenskapsrådet and the Swedish e-Science Research Center (to E.L.), and computational resources from the Swedish National Infrastructure for Computing (SNIC).
Conflict of Interest: none declared.
References