-
PDF
- Split View
-
Views
-
Cite
Cite
Hualin Liu, Jinshui Zheng, Dexin Bo, Yun Yu, Weixing Ye, Donghai Peng, Ming Sun, BtToxin_Digger: a comprehensive and high-throughput pipeline for mining toxin protein genes from Bacillus thuringiensis, Bioinformatics, Volume 38, Issue 1, January 2022, Pages 250–251, https://doi.org/10.1093/bioinformatics/btab506
Close - Share Icon Share
Abstract
Bacillus thuringiensis (Bt) has been used as the most successful microbial pesticide for decades. Its toxin genes are used for the development of genetically modified crops against pests. We previously developed a web-based insecticidal gene mining tool BtToxin_scanner. It has been frequently used by many researchers worldwide. However, it can only handle the genome one by one online. To facilitate efficiently mining toxin genes from large-scale sequence data, we re-designed this tool with a new workflow and the novel bacterial pesticidal protein database. Here, we present BtToxin_Digger, a comprehensive and high-throughput Bt toxin mining tool. It can be used to predict Bt toxin genes from thousands of raw genome and metagenome data, and provides accurate results for downstream analysis and experiment testing. Moreover, it can also be used to mine other targeting genes from large-scale genome and metagenome data with the replacement of the database.
The BtToxin_Digger codes and web services are freely available at https://github.com/BMBGenomics/BtToxin_Digger and https://bcam.hzau.edu.cn/BtToxin_Digger, respectively.
Supplementary data are available at Bioinformatics online.
1 Introduction
The toxins produced by Bacillus thuringiensis (Bt) have insecticidal activity against many agricultural and forestry pests. Bt can produce several kinds of insect-targeting toxins, such as insecticidal crystal protein (Cry), vegetative insecticidal protein (Vip), cytotoxic protein (Cyt), etc. The reported target insects of these toxins include those from Lepidoptera, Diptera, Coleoptera, etc. The cry and vip genes are among the most important ones used for the development of genetically modified (GM) crops targeting insect pests. From 1996 to 2016, the planting of Bt maize and cotton had delivered $50.6 billion and $54 billion of extra farm income, respectively (Brookes and Barfoot, 2018). To fight against the Bt toxin resistant insects and the new emerging pests, the discovery of new Bt strains and novel toxin genes is one of the most important strategies (Sanahuja et al., 2011). Previously, we developed an on-line tool BtToxin_scanner (Ye et al., 2012) to predict cry genes from Bt genome sequences, and it was frequently used by researchers including those who are interested in plant protection, GM crops development or sustainable agriculture (Adang et al., 2014; Carroll et al., 2020; Prado et al., 2014; Reyaz et al., 2019, 2021). It can handle one assembled genome each time and provides comparative results between the predicted toxin and the reported ones. Here, we re-designed the previous tool to provide a novel and high-throughput software BtToxin_Digger which can be used to handle large-scale genomic and metagenomic data to predict all kinds of putative toxin genes that match the recently updated toxin database (Crickmore et al., 2020), as well as other virulence factors which contribute to the pathogenicity but not lethality of Bt against its target insects, such as Sip (Donovan et al., 2006), Chitinase (Zhang et al., 2014), InhA (Dalhammar and Steiner, 1984), Bmp1 (Luo et al., 2013), Enhancin (Fang et al., 2009) and ZwA (He et al., 1994). It also generates comprehensive and readable results to facilitate the downstream sequence analysis or experiment design.
2 Materials and methods
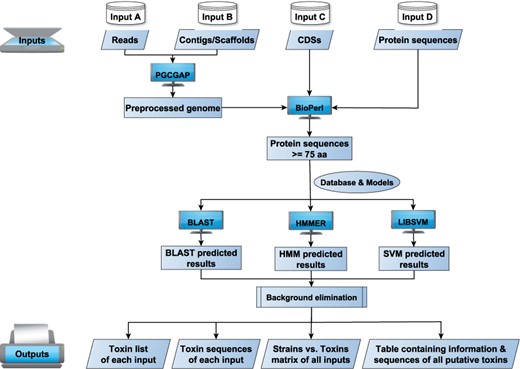
The types of input data supported by BtToxin_Digger include raw Reads data (pair-end reads generated by different platforms of Illumina, long-reads from PacBio and ONT or hybrid-reads), genome or metagenome assemblies, coding sequences (CDSs) and protein sequences. PGCGAP (Liu et al., 2020) is used for genome assembly. ORFs finding and translation are performed by BioPerl (Stajich et al., 2002). All protein sequences with a length above 115-aa are searched against the database and trained models by BLAST (Camacho et al., 2009), HMMER (Eddy, 2011) and LIBSVM (Chang and Lin, 2011), respectively. The candidate proteins are blasted against a background database to filter out the false-positive records. Then several Perl scripts are used to parse the results to get the putative target protein genes (Fig. 1).
3 Results
BtToxin_Digger can be used online and easily installed on Linux, macOS and Windows Subsystem for Linux (WSL) platforms by the conda package manager (Grüning et al., 2018) or docker container. We tested BtToxin_Digger on a laptop with an Intel CPU containing 8 threads of GHz-2.50 and 16 GB memory. It took about 14 min to process the 1.3-Gbp raw reads generated by Illumina Hiseq 2500 and less than 1 min for its assembled genome. In addition, BtToxin_Digger can be used to mine other interesting genes with the replacement of the toxin database by other target sequences.
Compared to the recent reported tool CryProcessor (Shikov et al., 2020), BtToxin_Digger presents the following advantages, more flexible input file types, more comprehensive and accurate results, more readable outputs (Supplementary Table S1). We tested BtToxin_Digger and CryProcessor using the protein sequences of 601 Bacillus thuringiensis genomes retrieved from GenBank. Our tool identified 18 types of interesting genes, while CryProcessor just predicted one type (Supplementary Table S2). For Cry toxins, BtToxin_Digger output not only the 874 ones with 3-domain structure predicted by CryProcessor but also other 371 Crys with at least one domain.
Funding
This work was supported by the National Key R&D Program of China [2017YFD0201201] and National Natural Science Foundation of China [31670085, 31970003 and 31770003].
Conflict of Interest: none declared.