-
PDF
- Split View
-
Views
-
Cite
Cite
Chiara Papetti, Antonio Di Franco, Lorenzo Zane, Paolo Guidetti, Valeria De Simone, Marianna Spizzotin, Barbara Zorica, Vanja Čikeš Keč, Carlotta Mazzoldi, Single population and common natal origin for Adriatic Scomber scombrus stocks: evidence from an integrated approach, ICES Journal of Marine Science, Volume 70, Issue 2, March 2013, Pages 387–398, https://doi.org/10.1093/icesjms/fss201
Close - Share Icon Share
Abstract
In order to implement proper fishery management strategies aimed at avoiding stock declines, information about connectivity among stocks and populations is critically required. In this perspective, the present study investigated population structure of the Atlantic mackerel Scomber scombrus in the northern–central Adriatic Sea by integrating multiple approaches (analysis of fisheries data, population genetics, and otolith chemistry). Monthly data of fishery landings indicate a latitudinal trend along the western Adriatic coast, with Atlantic mackerel disappearing from the northern waters in winter, corresponding to the reproductive season. Population genetic analyses by genotyping of eight microsatellites clearly point to the presence of a single panmictic population in the northern–central Adriatic Sea. Otolith cores of samples from the northern–central Adriatic were chemically homogeneous, suggesting a common spawning ground. These results strongly suggest that Atlantic mackerel perform an autumn–winter migration in the northern–central Adriatic Sea, from the northern to the central sector, to reach a single spawning ground, and that a single population is present in this area. Considering that S. scombrus has shown a marked decline in the last 40 years in the Adriatic, this study highlights a potential high vulnerability to collapse by overfishing for the Atlantic mackerel stocks in this geographic area.Papetti, C., Di Franco, A., Zane, L., Guidetti, P., De Simone, V., Spizzotin, M., Zorica, B., Čikeš Keč, V. and Mazzoldi, C. 2013. Single population and common natal origin for Adriatic Scomber scombrus stocks: evidence from an integrated approach – ICES Journal of Marine Science, 70: 387–398.
Introduction
Failing to detect the population structure of marine species targeted by fishing can lead to local overfishing and severe stock decline (Ying et al., 2011). This issue has often been underestimated based on the untested assumption of equality between the genetic concept of population (a group of conspecific individuals interbreeding and living in the same place at the same time) and the management concept of stock (identifying a management unit, i.e. a semi-discrete group of fish, with some definable attributes that are of interest to fishery managers regardless of strict genetic and/or biological assumptions; Jennings et al., 2001). This has resulted in neglecting both that a stock could be replenished by multiple populations and that multiple stocks could belong to a single population (Hauser and Carvalho, 2008; Ying et al., 2011). Proper fish stock management thus requires knowledge of the stock structure (i.e. identification of the populations replenishing the stock) and its variation with respect to environmental and ecological conditions (Allison et al., 2009; Rose et al., 2011). In particular, a better understanding of the factors influencing connectivity among spawning groups and estimation of the potential for replenishment of depleted grounds are critically needed (Rose et al., 2011). In this framework, a multidisciplinary approach integrating, for example, fisheries data, population genetics, and otolith microchemistry analyses could provide essential insights into population demographic processes, connectivity, and migration patterns (Hauser and Seeb, 2008), which represent critical pieces of information needed to answer relevant fishery science questions. Data from a specific fishing fleet operating in a definite area may allow the highlighting of fish migration patterns through periodic and predictable changes in catches (Walsh et al., 1995). Otolith microchemistry provides one of the most valuable methods to identify homogeneity/heterogeneity in the natal origin of individuals belonging to single or multiple stocks and potentially to make inferences regarding patterns of dispersal and connectivity. Otoliths can be used as a natural biological tag for natal origin of fish (Green et al., 2009), because they incorporate, into the calcium carbonate matrix of their core (i.e. the portion of the otolith originating at birth and thus related to the natal origin of fish; Green et al., 2009), chemical elements at rates related to environment concentrations and conditions at the moment of spawning. Similarly to otolith chemical data, genetics can provide connectivity information difficult to obtain with traditional methods, allowing a fine-scale picture of spatio-temporal boundaries of populations (Hauser and Carvalho, 2008; Rose et al., 2011).
In the present study, we integrated multiple approaches (i.e. fisheries data, otolith microchemistry, and genetics) to investigate population structure of Atlantic mackerel (Scomber scombrus) in the Adriatic Sea, where this species represents a valuable target for local fisheries (IREPA, 2011). Scomber scombrus is a pelagic species widely distributed in the Atlantic Ocean, Black Sea, and Mediterranean Sea (Studholme et al., 1999; Froese and Pauly, 2012). Atlantic mackerel form large shoals (Lockwood, 1988) that undergo migrations over great distances (i.e. 1200 km in 13 d in the Northeast Atlantic; Lockwood, 1988) mainly for reproductive and trophic needs (Walsh et al., 1995). Spawning grounds and/or migration routes are well known for the various stocks in the Atlantic, e.g. along the European shelf around Britain (Walsh et al., 1995; Reid et al., 1997), in the western Atlantic (Studholme et al., 1999), and in the eastern Atlantic (Uriarte and Lucio, 2001). Migration routes are usually known and exploited by local fisheries, with the start of fishing activities coinciding with the spawning migration (Punzón and Villamor, 2009).
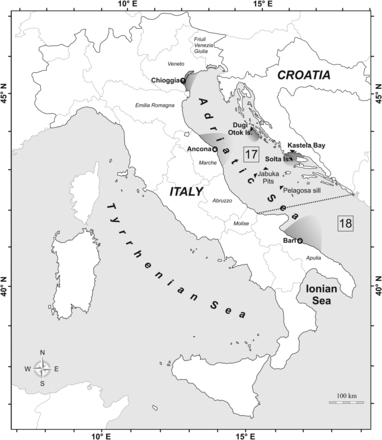
The Adriatic Sea is a semi-enclosed, elongated basin of the Mediterranean Sea (Russo and Artegiani, 1996; Figure 1) encompassing two Geographical Subareas (GSA 17 for the northern–central Adriatic and GSA 18 for the southern Adriatic; General Fisheries Commission for the Mediterranean, www.gfcm.org; Figure 1). Three sub-basins have been recognized: (i) the northern one extending from the northernmost part to the 100 m bathymetric line; (ii) the middle Adriatic from the limit of the northern sub-basin to the Pelagosa Sill (∼170 m depth) characterized by two depressions, the Jabuka (Pomo) Pits, reaching a maximum depth of ∼270 m; and (iii) the southern sub-basin extending from the Pelagosa Sill to the border of the Ionian Sea, with a depth up to 1000 m (Russo and Artegiani, 1996). The Adriatic Sea is one of the most exploited basins in the Mediterranean and has been historically characterized by an intense fishery on demersal and pelagic fish (Barausse et al., 2009). In the Adriatic Sea, mackerel is a target of multiple fisheries, namely bottom and pelagic trawling, purse-seine, and a small artisanal fishery (IREPA, 2011). This species has shown a massive decline in catches in the 1970s in different areas of the Adriatic, with no signs of recovery (Sinovčić, 2001; Azzurro et al., 2011; Clodia Database, 2012).
The Adriatic Sea. Italian regions, sampling sites (shaded areas), and the main localities cited in the text are indicated. The dotted line indicates the boundary between Geographical Subareas (GSA) 17 and 18 following the General Fisheries Commission for the Mediterranean.
The combination of analysis of fisheries data, population genetics, and otolith chemistry was used in this study to investigate population and stock structure of Atlantic mackerel in the northern–central Adriatic (i.e. GSA 17). In particular, landings data from the different Italian regions of the Adriatic Sea were used to highlight recurrent monthly–seasonal variations in catch, potentially providing suggestions about stock migrations. Population genetics analyses were aimed at verifying the presence of differentiation and possibly the existence of a panmictic population in the northern–central Adriatic Sea, as suggested by the potential for long-distance dispersal of this species. Otolith chemical analyses were aimed at identifying the occurrence of one or multiple spawning ground/s.
Material and methods
Fisheries data
To explore the monthly variation in Atlantic mackerel landings along the western Italian coast, official fisheries statistics from IREPA (Institute for Economic Research in Fishery and Aquaculture, http://www.irepa.org) were used. Mackerel landings were available for each Adriatic Italian region (Figure 1) from 2004 to 2009 (from 2005 to 2009 for Molise). Apulia data included only fisheries data from the Adriatic Sea.
Sample collection
Samples were collected within the GSA 17 from three areas of the Adriatic Sea located in the northern sub-basin and at the border between the northern and central sub-basins. In order to illustrate the latitudinal location of the three sites, they will be referred as: northern–western (off Chioggia), central–western (off Ancona), and central–eastern (Kastela Bay, open sea off Solta and Dugi Otok islands; Figure 1). Collections were made from April to July 2010, outside the breeding season that lasts from January to March (Bottari et al., 2004; C. Mazzoldi, unpubl. data). A total of 172 samples (Table 1) were obtained by local fishers and processed fresh in the laboratory. Total length (TL to the nearest 0.1 cm) and sex were recorded for each specimen. In order to avoid any bias in the results due to temporal or sex differences in genetic structure or otolith chemistry, samples were chosen to maintain a sex ratio and age ratio (1:2 years, C. Mazzoldi, unpubl. data) close to 1 for each sampling site. Age was assessed by otolith reading (C. Mazzoldi, unpubl. data). When this was not possible, the size range of individuals was used to infer age. All TLs were within the size range of individuals presenting age 1 or 2 (C. Mazzoldi, unpubl. data). A finclip was taken, preserved in 95% ethanol, and refrigerated at 4°C for genetic analysis. Otoliths (sagittae) were removed, cleaned of soft tissue using plastic dissecting pins, and dry preserved.
Samples of Scomber scombrus analysed in this study.
| Sampling area (site, GSA) . | Year . | Tot ind. . | Size range (mm TL) . | n genetic . | n otolith . |
|---|---|---|---|---|---|
| Northern–western (off Chioggia, 17) | 2010 | 60 | 240–356 | 60 | 53 |
| Central–western (off Ancona, 17) | 2010 | 59 | 228–300 | 59 | 39 |
| Central–eastern (Kastela Bay and off Solta and Dugi Otok, 17)a | 2010 | 53 | 237–333 | 53 | 39 |
| Southern–western (off Bari, 18) | 2001 | 50 | 265–317 | 50 | NA |
| Sampling area (site, GSA) . | Year . | Tot ind. . | Size range (mm TL) . | n genetic . | n otolith . |
|---|---|---|---|---|---|
| Northern–western (off Chioggia, 17) | 2010 | 60 | 240–356 | 60 | 53 |
| Central–western (off Ancona, 17) | 2010 | 59 | 228–300 | 59 | 39 |
| Central–eastern (Kastela Bay and off Solta and Dugi Otok, 17)a | 2010 | 53 | 237–333 | 53 | 39 |
| Southern–western (off Bari, 18) | 2001 | 50 | 265–317 | 50 | NA |
All individuals were genotyped for eight microsatellite loci, while population samples from the northern and central Adriatic Sea were used for otolith reading.
GSA, Geographic Subarea following the General Fisheries Commission for the Mediterranean nomenclature; year, year of collection; tot ind., total number of individuals analysed for each site; size range, total length in mm; n genetic, number of samples used for genetic analyses; n otolith, number of samples used for otolith analyses; NA, not available.
aSex was not determined for three specimens
Samples of Scomber scombrus analysed in this study.
| Sampling area (site, GSA) . | Year . | Tot ind. . | Size range (mm TL) . | n genetic . | n otolith . |
|---|---|---|---|---|---|
| Northern–western (off Chioggia, 17) | 2010 | 60 | 240–356 | 60 | 53 |
| Central–western (off Ancona, 17) | 2010 | 59 | 228–300 | 59 | 39 |
| Central–eastern (Kastela Bay and off Solta and Dugi Otok, 17)a | 2010 | 53 | 237–333 | 53 | 39 |
| Southern–western (off Bari, 18) | 2001 | 50 | 265–317 | 50 | NA |
| Sampling area (site, GSA) . | Year . | Tot ind. . | Size range (mm TL) . | n genetic . | n otolith . |
|---|---|---|---|---|---|
| Northern–western (off Chioggia, 17) | 2010 | 60 | 240–356 | 60 | 53 |
| Central–western (off Ancona, 17) | 2010 | 59 | 228–300 | 59 | 39 |
| Central–eastern (Kastela Bay and off Solta and Dugi Otok, 17)a | 2010 | 53 | 237–333 | 53 | 39 |
| Southern–western (off Bari, 18) | 2001 | 50 | 265–317 | 50 | NA |
All individuals were genotyped for eight microsatellite loci, while population samples from the northern and central Adriatic Sea were used for otolith reading.
GSA, Geographic Subarea following the General Fisheries Commission for the Mediterranean nomenclature; year, year of collection; tot ind., total number of individuals analysed for each site; size range, total length in mm; n genetic, number of samples used for genetic analyses; n otolith, number of samples used for otolith analyses; NA, not available.
aSex was not determined for three specimens
Sample processing for population genetics
Genetic analyses were performed on all collected specimens and on one additional “historical” sample (n = 50) collected in the southern–western Adriatic (in front of Bari, GSA 18) in June 2001.
Genomic DNA was extracted from 10–50 mg of finclip tissue following a standard salting-out protocol (Patwary et al., 1994). All individuals were genotyped for eight microsatellite loci (GenBank accession numbers JQ219865, JQ219866, JQ219869, JQ219871, JQ219872, JQ219874, JQ219876, and JQ219878). Polymerase chain reaction (PCR) products were obtained in a GeneAmp PCR System 9700 thermal cycler (Applied Biosystems) following described conditions (Molecular Ecology Resources Primer Development Consortium et al., 2012). Amplified fragments were screened for microsatellite polymorphism using an ABI 3130 XL automatic capillary sequencer (Applied Biosystems; service provided by BMR Genomics, http://bmr-genomics.com). Alleles were sized using a LIZ 500 (50–500 bp) size ladder (Applied Biosystems) and analysed using GeneMarker v. 1.71 (SoftGenetics); binning was automated with the software FlexiBin 2 (Amos et al., 2007) and refined by eye. All samples with unsuccessful PCR were reamplified; in addition, ∼10% of all individuals were reamplified and rerun to ensure repeatability of allele scoring and to check for genotyping artefacts. Micro-Checker ver. 2.2.3 (van Oosterhout et al., 2004) was used to identify possible genotyping errors (i.e. stuttering, large allele dropout, and null alleles, 1000 randomizations). Correction for null alleles was performed with FreeNA (Chapuis and Estoup, 2007).
Analysis of otolith chemistry
Otoliths were mounted sulcus side up on a glass slide using crystalbond, previously tested to ensure it was not a source of contamination under the adopted analysis settings. Otoliths were polished with 40, 3, and 1 μm Imperial lapping film to expose inner growth layers for analysis. We chose not to polish the otolith to the core, but to leave material above the core to ensure it was not removed during pre-ablation procedures, which potentially allowed us to sample all the material associated with the core. After polishing with lapping film, otoliths were rinsed and sonicated for 10 min in ultrapure water. After drying, otoliths were mounted in random order on clean petrographic slides.
All otoliths were analysed using a Thermo Elemental X7 inductively coupled plasma mass spectrometer (ICP-MS) coupled to a NewWave Research UP213 with an aperture imaging laser ablation (LA) system. External calibration was performed with two Standard Reference Materials (SRMs) from the National Institute of Standards and Technology: NIST 610 and NIST 612. Calcium was used as an internal standard to account for variation in ablation and aerosol efficiency. All nine elements analysed (24Mg, 55Mn, 66Zn, 88Sr, 138Ba, 208Pb, 7Li, 57Fe, and 59Co) were expressed as ratios relative to 44Ca.
Limits of detection (LOD) were calculated from the concentration of analyte, yielding a signal equivalent to three times the standard deviation of the blank signal for each of the elements, and are described in Table 2.
Limit of detection (LOD, given in mmol mol−1) for each trace element in Scomber scombrus otoliths and percentage of otolith ablations above the LOD for each element in both margins and cores.
| Element . | LOD . | % ablations > LOD (otolith core) . | % ablations > LOD (otolith margins) . |
|---|---|---|---|
| 7Li | 0.09695 | 0.00 | 0.44 |
| 24Mg | 0.00599 | 100.00 | 100.00 |
| 55Mn | 0.00971 | 1.94 | 0.44 |
| 57Fe | 0.30301 | 0.86 | 0.88 |
| 59Co | 0.00873 | 0.00 | 0.44 |
| 66Zn | 0.00818 | 14.62 | 30.68 |
| 88Sr | 0.00457 | 100.00 | 100.00 |
| 138Ba | 0.00026 | 100.00 | 100.00 |
| 208Pb | 0.00024 | 3.87 | 6.18 |
| Element . | LOD . | % ablations > LOD (otolith core) . | % ablations > LOD (otolith margins) . |
|---|---|---|---|
| 7Li | 0.09695 | 0.00 | 0.44 |
| 24Mg | 0.00599 | 100.00 | 100.00 |
| 55Mn | 0.00971 | 1.94 | 0.44 |
| 57Fe | 0.30301 | 0.86 | 0.88 |
| 59Co | 0.00873 | 0.00 | 0.44 |
| 66Zn | 0.00818 | 14.62 | 30.68 |
| 88Sr | 0.00457 | 100.00 | 100.00 |
| 138Ba | 0.00026 | 100.00 | 100.00 |
| 208Pb | 0.00024 | 3.87 | 6.18 |
Limit of detection (LOD, given in mmol mol−1) for each trace element in Scomber scombrus otoliths and percentage of otolith ablations above the LOD for each element in both margins and cores.
| Element . | LOD . | % ablations > LOD (otolith core) . | % ablations > LOD (otolith margins) . |
|---|---|---|---|
| 7Li | 0.09695 | 0.00 | 0.44 |
| 24Mg | 0.00599 | 100.00 | 100.00 |
| 55Mn | 0.00971 | 1.94 | 0.44 |
| 57Fe | 0.30301 | 0.86 | 0.88 |
| 59Co | 0.00873 | 0.00 | 0.44 |
| 66Zn | 0.00818 | 14.62 | 30.68 |
| 88Sr | 0.00457 | 100.00 | 100.00 |
| 138Ba | 0.00026 | 100.00 | 100.00 |
| 208Pb | 0.00024 | 3.87 | 6.18 |
| Element . | LOD . | % ablations > LOD (otolith core) . | % ablations > LOD (otolith margins) . |
|---|---|---|---|
| 7Li | 0.09695 | 0.00 | 0.44 |
| 24Mg | 0.00599 | 100.00 | 100.00 |
| 55Mn | 0.00971 | 1.94 | 0.44 |
| 57Fe | 0.30301 | 0.86 | 0.88 |
| 59Co | 0.00873 | 0.00 | 0.44 |
| 66Zn | 0.00818 | 14.62 | 30.68 |
| 88Sr | 0.00457 | 100.00 | 100.00 |
| 138Ba | 0.00026 | 100.00 | 100.00 |
| 208Pb | 0.00024 | 3.87 | 6.18 |
Otoliths were analysed for chemical composition of both the core (in order to acquire information about natal origin) and the margin (i.e. close to the otolith edge, reflecting exposure of an individual to environmental conditions at the site where the individual was sampled).
Otoliths were placed in the ablation chamber and viewed remotely on a computer screen where the area for ablation was selected. The laser was focused on the sample surface and fired through the microscope objective lens using a spot size of 30 μm. Each run generally consisted of 40 s acquisition: (i) 10 s blank to correct for background, which was subtracted from each sample; (ii) 10 s ablation (laser at 65% power, ∼6 J cm−2) resulting in a pit ∼10 μm deep; and (iii) 20 s for washout. Prior to analysis, samples were pre-ablated to remove any surface contamination (laser at 50% power). Helium gas was flushed into the ablation cell to reduce the deposition of ablated aerosols and to improve signal intensities. The ablated aerosol was then mixed with argon before entering the ICP torch.
Elemental analyses of otolith cores and margins produced concentrations that were greater than the LOD in 100% of the samples for Sr, Ba, and Mg (Table 2). Concentrations of other elements considered were predominantly less than the LOD; for this reason, these elements were excluded from subsequent analyses.
We sampled the material associated with the core using three discrete vertical pits 30 μm (identified previously as the approximate size of the cores) from the surface of the otolith through the visible core. Due to the exclusion of Mn from the analyses (because it was consistently below the LOD) and due to the fact that the LOD of Mn in the present study was similar to or lower than those from other studies where a spike in Mn:Ca was adopted as an indicator of the core location (e.g. Di Franco et al., 2012, and references therein), we hypothesized that, in the studied species, a spike in Mn:Ca could not be an effective “core localizer”. From this perspective, we chose to consider for the core analyses all the three replicates sampled. In the present work, the core identifies the area laid down at egg fecundation and very early larval stages (as in Miller and Shanks, 2004).
In the otolith margin, we ablated three horizontal pits, and all three were considered in subsequent analysis in order to account for within-otolith variability (see Di Franco et al., 2011 for further details).
Data analyses
Monthly fisheries data for each Italian region were analysed applying the Kruskal–Wallis non-parametric analysis of variance (ANOVA), excluding August data since midwater and bottom trawling is banned in this month in several regions.
Microsatellite genotypes were examined to infer genetic variability and population structure patterns. The online version of Genepop (Raymond and Rousset, 1995) was used to identify deviations from Hardy–Weinberg equilibrium (HWE) by exact tests (1000 iterations) and genotypic disequilibrium (1000 iterations). Significance levels for multiple comparisons were adjusted using the standard Bonferroni technique (Bonferroni, 1936; Miller, 1981).
Genetix ver. 4.05.2 (Belkhir et al., 2001) was used to calculate the total number of alleles (TNA) and the observed and expected heterozygosities (Ho and He, respectively). Allelic richness (AR) was calculated by Fstat 2.9.3.2 (Goudet, 1995). A Bayesian algorithm implemented in the software Structure ver. 2.3.3 (Pritchard et al., 2000) was used to identify genetically homogeneous groups of individuals on the basis of their genotypes at multiple loci.
Differences in allele and genotype frequencies among samples (sampling site, sex, and age/size) were assessed using Fisher's exact test as implemented in Genepop. Significance levels for multiple simultaneous comparisons were adjusted using the Bonferroni technique, as described above. Population structure was explored by calculating pairwise FST between samples in Arlequin 3.5.1.2 (Excoffier and Lischer, 2010). Finally, the statistical power to detect various true levels of divergence for our sample sizes, number of loci, and their allele frequencies was evaluated by means of Fisher's exact test using the simulation method of Ryman and Palm (2006) implemented in the software Powsim ver. 4.1. Simulations were run using default parameter values for dememorizations (1000), batches (100), and iterations per batch (1000) for a scenario involving four subpopulations with FST ranging from 0.001 to 0.02, using an effective population size (Ne of 6600) calculated with a heterozygosity-based method (Ohta and Kimura, 1973). This method predicts that, at mutation–drift equilibrium under the stepwise mutation model (SMM), Ne should equal [(1/1 – He)2 – 1]/8μ, where He is the observed averaged heterozygosity and μ is the mutation rate. μ was assumed to be 5 × 10−4, according to Estoup and Angers (1998). The statistical power was estimated after 2000 replicates as the proportion of statistically significant test (p < 0.05). The probability of obtaining false positives when the true FST = 0 was also obtained at generation t = 0 as a measure of α error rate. To assess consistency, all simulations were repeated at least three times.
For otolith chemistry analyses, to determine the number of potential natal origins, the otoliths' core elemental concentrations (as a proxy for identifying the existence of single or multiple areas of origin) were analysed by cluster analysis. The similarity profile permutation test (SIMPROF) procedure was used to determine which clusters were significantly different at the 5% level. For each specimen, the centroid from the three replicates was considered.
Because homogeneity in otolith chemical composition may simply reflect environmental similarity, we evaluated potential spatial variability in otolith chemical composition. For this purpose, we used unbalanced permutational multivariate analysis of variance (PERMANOVA) to test for differences between the three sampling sites by analysing the otolith margin (i.e. the otolith portion laid down just before capture). “Site” (Si) was treated as a random factor (three levels), and “Otolith” (Ot) as a random factor nested in (Si) (39–59 levels). There were three replicate ablations for each otolith (n = 453): the assessment of intraotolith variability is, in fact, instrumental to then assess the “among-otoliths variability” (Di Franco et al., 2011). Fish TL was set as the covariate. The test for covariate effect was performed to prevent a fish size (possibly different from site to site and considered as a proxy of fish age) effect on spatial comparisons. In other words, only conditioning on fish size, any observed difference is attributable to “pure” spatial patterns. Elemental/Ca ratios that contribute to the significant differences among sites were identified using similarity percentage (SIMPER).
Multivariate components of variation (VC), %TV [% of total variation, i.e. (variation of each factor/total variation) in %], and the ratio “estimated magnitude of variance for each factor/estimated residual variance” (ø) were calculated for the two random factors considered in the PERMANOVA analyses (Gray et al., 2009).
Statistical analyses were run using the Primer 6 PERMANOVA + software package and STATISTICA 10 software.
Results
Fisheries data
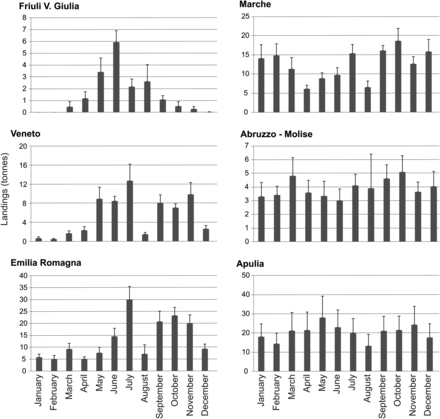
Average landing of Atlantic mackerel showed significant differences between months in Friuli Venezia Giulia (H10 = 48.70; p < 0.0001), Veneto (H10 = 46.52; p < 0.0001), Emilia Romagna (H10 = 40.91; p < 0.0001), and Marche (H10 = 19.62; p = 0.033). Landings showed a seasonal trend shifted along a north–south gradient: the northern part, Friuli Venezia Giulia, presented the lowest landings in autumn–winter, Veneto in winter, and Emilia Romagna in winter–spring. Marche did not show any clear pattern. No significant differences were found in Abruzzo (H10 = 4.69; p = 0.911), Molise (H10 = 6.57; p = 0.765), and Apulia (H10 = 3.21; p = 0.976) (Figure 2).
Monthly landings of Scomber scombrus in the Italian regions (data from IREPA, 2004–2009). Bars represent standard errors. Data from Abruzzo and Molise were pooled.
Population genetics
The total number of alleles ranged between 6 (locus Ss235 in the central–eastern Adriatic) and 30 (locus Ss48 in the central–western Adriatic), and allelic richness varied from 5.984 (locus Ss235 in the central–eastern Adriatic) to 28.856 (locus Ss48 in the southern–western Adriatic). Average observed heterozygosity ranged from 0.746 in the northern–western sample to 0.789 in the central–western Adriatic, whereas average expected heterozygosity varied from 0.794 in the central–eastern to 0.817 in the southern–western sample (Table 3). All comparisons across samples using a non-parametric Friedman test were statistically not significant (p > 0.5) except for allelic richness comparison (p = 0.014), due to higher AR in the GSA 18 sample.
Summary of genetic variability for all samples of Scomber scombrus in four sampling sites at eight microsatellite loci, including allelic richness (AR) based on a minimum sample size of 48 individuals, total number of alleles (TNA), observed (Ho), and expected heterozygosity (He).
| Locus . | Parameter . | Sampling area . | Total . | |||
|---|---|---|---|---|---|---|
| Northern–western . | Central–western . | Central–eastern . | Southern–western . | |||
| Ss25 | AR | 7.588 | 9.409 | 9.708 | 11.000 | 10.010 |
| TNA | 8 | 10 | 10 | 11 | 16 | |
| Ho | 0.644 | 0.786 | 0.679 | 0.750 | 0.714 | |
| He | 0.707 | 0.738 | 0.731 | 0.844 | 0.755 | |
| p(HWE) | 0.1439 | 0.9777 | 0.0484 | 0.1299 | 0.0802 | |
| Ss48 | AR | 27.196 | 28.848 | 26.755 | 28.856 | 28.926 |
| TNA | 28 | 30 | 27 | 29 | 36 | |
| Ho | 0.898 | 0.898 | 0.900 | 0.898 | 0.898 | |
| He | 0.951 | 0.963 | 0.958 | 0.962 | 0.958 | |
| p(HWE) | 0.0307 | 0.1019 | 0.3636 | 0.2108 | 0.0337 | |
| Ss142 | AR | 24.169 | 22.685 | 21.373 | 25.554 | 23.366 |
| TNA | 26 | 24 | 22 | 26 | 35 | |
| Ho | 0.746 | 0.763 | 0.846 | 0.880 | 0.808 | |
| He | 0.930 | 0.926 | 0.913 | 0.932 | 0.925 | |
| p(HWE) | <0.0001 | 0.0001 | 0.0989 | 0.0700 | <0.0001 | |
| Ss180 | AR | 21.827 | 15.999 | 16.683 | 24.734 | 20.151 |
| TNA | 24 | 17 | 17 | 25 | 35 | |
| Ho | 0.767 | 0.814 | 0.849 | 0.878 | 0.827 | |
| He | 0.844 | 0.850 | 0.821 | 0.858 | 0.843 | |
| p(HWE) | 0.0377 | 0.4255 | 0.5693 | 0.8519 | 0.2858 | |
| Ss186 | AR | 24.238 | 23.190 | 23.580 | 24.676 | 23.983 |
| TNA | 25 | 24 | 24 | 25 | 28 | |
| Ho | 0.917 | 0.898 | 0.906 | 0.840 | 0.890 | |
| He | 0.949 | 0.952 | 0.938 | 0.950 | 0.947 | |
| p(HWE) | 0.5223 | 0.0497 | 0.1114 | 0.0410 | 0.0206 | |
| Ss212 | AR | 9.161 | 8.713 | 9.528 | 12.796 | 9.925 |
| TNA | 10 | 9 | 10 | 13 | 17 | |
| Ho | 0.617 | 0.564 | 0.604 | 0.640 | 0.606 | |
| He | 0.660 | 0.601 | 0.557 | 0.639 | 0.614 | |
| p(HWE) | 0.0774 | 0.1418 | 0.2178 | 0.3494 | 0.0773 | |
| Ss224 | AR | 7.778 | 7.621 | 8.803 | 8.958 | 8.443 |
| TNA | 8 | 8 | 9 | 9 | 11 | |
| Ho | 0.712 | 0.695 | 0.698 | 0.580 | 0.671 | |
| He | 0.698 | 0.698 | 0.734 | 0.627 | 0.689 | |
| p(HWE) | 0.1050 | 0.1854 | 0.3821 | 0.1972 | 0.1102 | |
| Ss235 | AR | 8.677 | 9.533 | 5.984 | 7.879 | 8.801 |
| TNA | 9 | 10 | 6 | 8 | 13 | |
| Ho | 0.667 | 0.893 | 0.793 | 0.680 | 0.758 | |
| He | 0.757 | 0.782 | 0.699 | 0.727 | 0.741 | |
| p(HWE) | 0.0117 | 0.0644 | 0.0383 | 0.5497 | 0.0047 | |
| Overall | AR | 16.329 | 15.750 | 15.302 | 18.057 | 16.701 |
| TNA (mean) | 17.250 | 16.500 | 15.625 | 18.250 | 23.875 | |
| Ho | 0.746 | 0.789 | 0.784 | 0.768 | 0.771 | |
| He | 0.812 | 0.814 | 0.794 | 0.817 | 0.809 | |
| p(HWE) | <0.0001 | 0.0002 | 0.0195 | 0.0574 | <0.0001 | |
| Locus . | Parameter . | Sampling area . | Total . | |||
|---|---|---|---|---|---|---|
| Northern–western . | Central–western . | Central–eastern . | Southern–western . | |||
| Ss25 | AR | 7.588 | 9.409 | 9.708 | 11.000 | 10.010 |
| TNA | 8 | 10 | 10 | 11 | 16 | |
| Ho | 0.644 | 0.786 | 0.679 | 0.750 | 0.714 | |
| He | 0.707 | 0.738 | 0.731 | 0.844 | 0.755 | |
| p(HWE) | 0.1439 | 0.9777 | 0.0484 | 0.1299 | 0.0802 | |
| Ss48 | AR | 27.196 | 28.848 | 26.755 | 28.856 | 28.926 |
| TNA | 28 | 30 | 27 | 29 | 36 | |
| Ho | 0.898 | 0.898 | 0.900 | 0.898 | 0.898 | |
| He | 0.951 | 0.963 | 0.958 | 0.962 | 0.958 | |
| p(HWE) | 0.0307 | 0.1019 | 0.3636 | 0.2108 | 0.0337 | |
| Ss142 | AR | 24.169 | 22.685 | 21.373 | 25.554 | 23.366 |
| TNA | 26 | 24 | 22 | 26 | 35 | |
| Ho | 0.746 | 0.763 | 0.846 | 0.880 | 0.808 | |
| He | 0.930 | 0.926 | 0.913 | 0.932 | 0.925 | |
| p(HWE) | <0.0001 | 0.0001 | 0.0989 | 0.0700 | <0.0001 | |
| Ss180 | AR | 21.827 | 15.999 | 16.683 | 24.734 | 20.151 |
| TNA | 24 | 17 | 17 | 25 | 35 | |
| Ho | 0.767 | 0.814 | 0.849 | 0.878 | 0.827 | |
| He | 0.844 | 0.850 | 0.821 | 0.858 | 0.843 | |
| p(HWE) | 0.0377 | 0.4255 | 0.5693 | 0.8519 | 0.2858 | |
| Ss186 | AR | 24.238 | 23.190 | 23.580 | 24.676 | 23.983 |
| TNA | 25 | 24 | 24 | 25 | 28 | |
| Ho | 0.917 | 0.898 | 0.906 | 0.840 | 0.890 | |
| He | 0.949 | 0.952 | 0.938 | 0.950 | 0.947 | |
| p(HWE) | 0.5223 | 0.0497 | 0.1114 | 0.0410 | 0.0206 | |
| Ss212 | AR | 9.161 | 8.713 | 9.528 | 12.796 | 9.925 |
| TNA | 10 | 9 | 10 | 13 | 17 | |
| Ho | 0.617 | 0.564 | 0.604 | 0.640 | 0.606 | |
| He | 0.660 | 0.601 | 0.557 | 0.639 | 0.614 | |
| p(HWE) | 0.0774 | 0.1418 | 0.2178 | 0.3494 | 0.0773 | |
| Ss224 | AR | 7.778 | 7.621 | 8.803 | 8.958 | 8.443 |
| TNA | 8 | 8 | 9 | 9 | 11 | |
| Ho | 0.712 | 0.695 | 0.698 | 0.580 | 0.671 | |
| He | 0.698 | 0.698 | 0.734 | 0.627 | 0.689 | |
| p(HWE) | 0.1050 | 0.1854 | 0.3821 | 0.1972 | 0.1102 | |
| Ss235 | AR | 8.677 | 9.533 | 5.984 | 7.879 | 8.801 |
| TNA | 9 | 10 | 6 | 8 | 13 | |
| Ho | 0.667 | 0.893 | 0.793 | 0.680 | 0.758 | |
| He | 0.757 | 0.782 | 0.699 | 0.727 | 0.741 | |
| p(HWE) | 0.0117 | 0.0644 | 0.0383 | 0.5497 | 0.0047 | |
| Overall | AR | 16.329 | 15.750 | 15.302 | 18.057 | 16.701 |
| TNA (mean) | 17.250 | 16.500 | 15.625 | 18.250 | 23.875 | |
| Ho | 0.746 | 0.789 | 0.784 | 0.768 | 0.771 | |
| He | 0.812 | 0.814 | 0.794 | 0.817 | 0.809 | |
| p(HWE) | <0.0001 | 0.0002 | 0.0195 | 0.0574 | <0.0001 | |
Probabilities of deviation from Hardy–Weinberg equilibrium [p(HWE)] are presented at each locus. Values in bold indicate significant HWE deviations after Bonferroni correction (p < 0.00125).
Summary of genetic variability for all samples of Scomber scombrus in four sampling sites at eight microsatellite loci, including allelic richness (AR) based on a minimum sample size of 48 individuals, total number of alleles (TNA), observed (Ho), and expected heterozygosity (He).
| Locus . | Parameter . | Sampling area . | Total . | |||
|---|---|---|---|---|---|---|
| Northern–western . | Central–western . | Central–eastern . | Southern–western . | |||
| Ss25 | AR | 7.588 | 9.409 | 9.708 | 11.000 | 10.010 |
| TNA | 8 | 10 | 10 | 11 | 16 | |
| Ho | 0.644 | 0.786 | 0.679 | 0.750 | 0.714 | |
| He | 0.707 | 0.738 | 0.731 | 0.844 | 0.755 | |
| p(HWE) | 0.1439 | 0.9777 | 0.0484 | 0.1299 | 0.0802 | |
| Ss48 | AR | 27.196 | 28.848 | 26.755 | 28.856 | 28.926 |
| TNA | 28 | 30 | 27 | 29 | 36 | |
| Ho | 0.898 | 0.898 | 0.900 | 0.898 | 0.898 | |
| He | 0.951 | 0.963 | 0.958 | 0.962 | 0.958 | |
| p(HWE) | 0.0307 | 0.1019 | 0.3636 | 0.2108 | 0.0337 | |
| Ss142 | AR | 24.169 | 22.685 | 21.373 | 25.554 | 23.366 |
| TNA | 26 | 24 | 22 | 26 | 35 | |
| Ho | 0.746 | 0.763 | 0.846 | 0.880 | 0.808 | |
| He | 0.930 | 0.926 | 0.913 | 0.932 | 0.925 | |
| p(HWE) | <0.0001 | 0.0001 | 0.0989 | 0.0700 | <0.0001 | |
| Ss180 | AR | 21.827 | 15.999 | 16.683 | 24.734 | 20.151 |
| TNA | 24 | 17 | 17 | 25 | 35 | |
| Ho | 0.767 | 0.814 | 0.849 | 0.878 | 0.827 | |
| He | 0.844 | 0.850 | 0.821 | 0.858 | 0.843 | |
| p(HWE) | 0.0377 | 0.4255 | 0.5693 | 0.8519 | 0.2858 | |
| Ss186 | AR | 24.238 | 23.190 | 23.580 | 24.676 | 23.983 |
| TNA | 25 | 24 | 24 | 25 | 28 | |
| Ho | 0.917 | 0.898 | 0.906 | 0.840 | 0.890 | |
| He | 0.949 | 0.952 | 0.938 | 0.950 | 0.947 | |
| p(HWE) | 0.5223 | 0.0497 | 0.1114 | 0.0410 | 0.0206 | |
| Ss212 | AR | 9.161 | 8.713 | 9.528 | 12.796 | 9.925 |
| TNA | 10 | 9 | 10 | 13 | 17 | |
| Ho | 0.617 | 0.564 | 0.604 | 0.640 | 0.606 | |
| He | 0.660 | 0.601 | 0.557 | 0.639 | 0.614 | |
| p(HWE) | 0.0774 | 0.1418 | 0.2178 | 0.3494 | 0.0773 | |
| Ss224 | AR | 7.778 | 7.621 | 8.803 | 8.958 | 8.443 |
| TNA | 8 | 8 | 9 | 9 | 11 | |
| Ho | 0.712 | 0.695 | 0.698 | 0.580 | 0.671 | |
| He | 0.698 | 0.698 | 0.734 | 0.627 | 0.689 | |
| p(HWE) | 0.1050 | 0.1854 | 0.3821 | 0.1972 | 0.1102 | |
| Ss235 | AR | 8.677 | 9.533 | 5.984 | 7.879 | 8.801 |
| TNA | 9 | 10 | 6 | 8 | 13 | |
| Ho | 0.667 | 0.893 | 0.793 | 0.680 | 0.758 | |
| He | 0.757 | 0.782 | 0.699 | 0.727 | 0.741 | |
| p(HWE) | 0.0117 | 0.0644 | 0.0383 | 0.5497 | 0.0047 | |
| Overall | AR | 16.329 | 15.750 | 15.302 | 18.057 | 16.701 |
| TNA (mean) | 17.250 | 16.500 | 15.625 | 18.250 | 23.875 | |
| Ho | 0.746 | 0.789 | 0.784 | 0.768 | 0.771 | |
| He | 0.812 | 0.814 | 0.794 | 0.817 | 0.809 | |
| p(HWE) | <0.0001 | 0.0002 | 0.0195 | 0.0574 | <0.0001 | |
| Locus . | Parameter . | Sampling area . | Total . | |||
|---|---|---|---|---|---|---|
| Northern–western . | Central–western . | Central–eastern . | Southern–western . | |||
| Ss25 | AR | 7.588 | 9.409 | 9.708 | 11.000 | 10.010 |
| TNA | 8 | 10 | 10 | 11 | 16 | |
| Ho | 0.644 | 0.786 | 0.679 | 0.750 | 0.714 | |
| He | 0.707 | 0.738 | 0.731 | 0.844 | 0.755 | |
| p(HWE) | 0.1439 | 0.9777 | 0.0484 | 0.1299 | 0.0802 | |
| Ss48 | AR | 27.196 | 28.848 | 26.755 | 28.856 | 28.926 |
| TNA | 28 | 30 | 27 | 29 | 36 | |
| Ho | 0.898 | 0.898 | 0.900 | 0.898 | 0.898 | |
| He | 0.951 | 0.963 | 0.958 | 0.962 | 0.958 | |
| p(HWE) | 0.0307 | 0.1019 | 0.3636 | 0.2108 | 0.0337 | |
| Ss142 | AR | 24.169 | 22.685 | 21.373 | 25.554 | 23.366 |
| TNA | 26 | 24 | 22 | 26 | 35 | |
| Ho | 0.746 | 0.763 | 0.846 | 0.880 | 0.808 | |
| He | 0.930 | 0.926 | 0.913 | 0.932 | 0.925 | |
| p(HWE) | <0.0001 | 0.0001 | 0.0989 | 0.0700 | <0.0001 | |
| Ss180 | AR | 21.827 | 15.999 | 16.683 | 24.734 | 20.151 |
| TNA | 24 | 17 | 17 | 25 | 35 | |
| Ho | 0.767 | 0.814 | 0.849 | 0.878 | 0.827 | |
| He | 0.844 | 0.850 | 0.821 | 0.858 | 0.843 | |
| p(HWE) | 0.0377 | 0.4255 | 0.5693 | 0.8519 | 0.2858 | |
| Ss186 | AR | 24.238 | 23.190 | 23.580 | 24.676 | 23.983 |
| TNA | 25 | 24 | 24 | 25 | 28 | |
| Ho | 0.917 | 0.898 | 0.906 | 0.840 | 0.890 | |
| He | 0.949 | 0.952 | 0.938 | 0.950 | 0.947 | |
| p(HWE) | 0.5223 | 0.0497 | 0.1114 | 0.0410 | 0.0206 | |
| Ss212 | AR | 9.161 | 8.713 | 9.528 | 12.796 | 9.925 |
| TNA | 10 | 9 | 10 | 13 | 17 | |
| Ho | 0.617 | 0.564 | 0.604 | 0.640 | 0.606 | |
| He | 0.660 | 0.601 | 0.557 | 0.639 | 0.614 | |
| p(HWE) | 0.0774 | 0.1418 | 0.2178 | 0.3494 | 0.0773 | |
| Ss224 | AR | 7.778 | 7.621 | 8.803 | 8.958 | 8.443 |
| TNA | 8 | 8 | 9 | 9 | 11 | |
| Ho | 0.712 | 0.695 | 0.698 | 0.580 | 0.671 | |
| He | 0.698 | 0.698 | 0.734 | 0.627 | 0.689 | |
| p(HWE) | 0.1050 | 0.1854 | 0.3821 | 0.1972 | 0.1102 | |
| Ss235 | AR | 8.677 | 9.533 | 5.984 | 7.879 | 8.801 |
| TNA | 9 | 10 | 6 | 8 | 13 | |
| Ho | 0.667 | 0.893 | 0.793 | 0.680 | 0.758 | |
| He | 0.757 | 0.782 | 0.699 | 0.727 | 0.741 | |
| p(HWE) | 0.0117 | 0.0644 | 0.0383 | 0.5497 | 0.0047 | |
| Overall | AR | 16.329 | 15.750 | 15.302 | 18.057 | 16.701 |
| TNA (mean) | 17.250 | 16.500 | 15.625 | 18.250 | 23.875 | |
| Ho | 0.746 | 0.789 | 0.784 | 0.768 | 0.771 | |
| He | 0.812 | 0.814 | 0.794 | 0.817 | 0.809 | |
| p(HWE) | <0.0001 | 0.0002 | 0.0195 | 0.0574 | <0.0001 | |
Probabilities of deviation from Hardy–Weinberg equilibrium [p(HWE)] are presented at each locus. Values in bold indicate significant HWE deviations after Bonferroni correction (p < 0.00125).
Significant departure from HWE was found at Ss142 in the population samples from the central–western and northern–western Adriatic, with a corresponding heterozygote deficiency in both samples. Micro-Checker provided no indications that genotyping errors affected allele scoring (e.g. allele dropouts or stuttering) at any marker in any sample, whereas it identified null alleles at locus Ss142. However, departure from HWE remained significant after correction for null alleles with FreeNA (Chapuis and Estoup, 2007), and samples that failed to amplify were <1%, indicating that null homozygotes were not common. In addition, no sample failed to amplify at more than one locus (data not shown), and this makes it unlikely that poor DNA quality affected our results. For these reasons, we maintained the uncorrected genotypes of locus Ss142 in our analyses, and we cross-checked differentiation results removing the locus from the dataset when appropriate. Non-significant genotypic disequilibrium was found after Bonferroni correction for multiple tests.
When investigating the genetic structure of the populations in GSA 17 (therefore excluding the historical sample from the southern–western Adriatic in order to exclude any potential bias due to the wide difference in collection time, i.e. 9 years), the software Structure inferred one single population as most likely. Indeed, no significant values were obtained in all the comparisons involving northern–western, central–western, and central–eastern samples by Fisher's exact test (p > 0.05) and pairwise FST (p > 0.05, Table 4). In contrast, both approaches identified a weak but significant difference between the 2010 northern–central Adriatic samples and the sample collected in the southern–western Adriatic in 2001, with an average FST of 0.009 (p < 0.05) (Table 4). While this comparison does not allow discrimination between geographic and temporal variation, it clearly shows that our set of markers is adequate to detect genetic differentiation, as also demonstrated by statistical power detection. In fact, our simulation demonstrated that our markers would detect a true FST of ≥0.005 with a probability approaching 100% (Table 5). The α error (when true FST = 0, t = 0) was consistently close to 5% in all simulations (data not reported). The data in Tables 4 and 5 were obtained over the entire dataset of eight loci. Similar conclusions were obtained after removing locus Ss142 (data not shown). No differentiation was detected between males and females, among age/size classes and otolith clusters depicted in Figure 3 (data not reported) either considering or excluding locus Ss142.
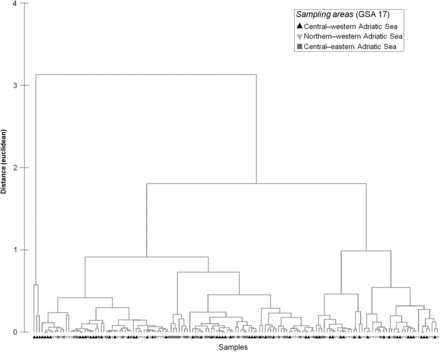
Classification of sample otolith cores of Scomber scombrus. No statistically different group was detected from cluster analysis (indicated by grey lines).
Pairwise FST estimates (above diagonal) and homogeneity test probability (below diagonal) among all samples of Scomber scombrus collected in the Adriatic Sea.
| . | Northern–western . | Central–western . | Central–eastern . | Southern–western . |
|---|---|---|---|---|
| Northern–western | – | –0.001 | 0.004 | 0.011 |
| Central–western | 0.703 | – | 0.001 | 0.007 |
| Central–eastern | 0.063 | 0.333 | – | 0.008 |
| Southern–western | <0.001 | <0.001 | <0.001 | – |
| . | Northern–western . | Central–western . | Central–eastern . | Southern–western . |
|---|---|---|---|---|
| Northern–western | – | –0.001 | 0.004 | 0.011 |
| Central–western | 0.703 | – | 0.001 | 0.007 |
| Central–eastern | 0.063 | 0.333 | – | 0.008 |
| Southern–western | <0.001 | <0.001 | <0.001 | – |
Values in bold are significant after Bonferroni correction (p < 0.0083).
Pairwise FST estimates (above diagonal) and homogeneity test probability (below diagonal) among all samples of Scomber scombrus collected in the Adriatic Sea.
| . | Northern–western . | Central–western . | Central–eastern . | Southern–western . |
|---|---|---|---|---|
| Northern–western | – | –0.001 | 0.004 | 0.011 |
| Central–western | 0.703 | – | 0.001 | 0.007 |
| Central–eastern | 0.063 | 0.333 | – | 0.008 |
| Southern–western | <0.001 | <0.001 | <0.001 | – |
| . | Northern–western . | Central–western . | Central–eastern . | Southern–western . |
|---|---|---|---|---|
| Northern–western | – | –0.001 | 0.004 | 0.011 |
| Central–western | 0.703 | – | 0.001 | 0.007 |
| Central–eastern | 0.063 | 0.333 | – | 0.008 |
| Southern–western | <0.001 | <0.001 | <0.001 | – |
Values in bold are significant after Bonferroni correction (p < 0.0083).
Estimates of statistical power for detecting various true levels of population structure (FST) in Scomber scombrus by means of Fisher's exact test when using the present loci, allele frequencies, and sample sizes from the four sampling localities.
| Expected FST . | Average FST . | Fisher's test . | t . |
|---|---|---|---|
| 0.0010 | 0.0010 | 0.4185 | 13 |
| 0.0025 | 0.0025 | 0.9420 | 33 |
| 0.0050 | 0.0050 | 1.0000 | 66 |
| 0.0100 | 0.0100 | 1.0000 | 132 |
| 0.0200 | 0.0200 | 1.0000 | 266 |
| Expected FST . | Average FST . | Fisher's test . | t . |
|---|---|---|---|
| 0.0010 | 0.0010 | 0.4185 | 13 |
| 0.0025 | 0.0025 | 0.9420 | 33 |
| 0.0050 | 0.0050 | 1.0000 | 66 |
| 0.0100 | 0.0100 | 1.0000 | 132 |
| 0.0200 | 0.0200 | 1.0000 | 266 |
The power is expressed as the proportion of simulations that provide statistical significance at the 0.05 level after a number of generations (t) which are necessary to reach an average FST equal to the expected values.
Estimates of statistical power for detecting various true levels of population structure (FST) in Scomber scombrus by means of Fisher's exact test when using the present loci, allele frequencies, and sample sizes from the four sampling localities.
| Expected FST . | Average FST . | Fisher's test . | t . |
|---|---|---|---|
| 0.0010 | 0.0010 | 0.4185 | 13 |
| 0.0025 | 0.0025 | 0.9420 | 33 |
| 0.0050 | 0.0050 | 1.0000 | 66 |
| 0.0100 | 0.0100 | 1.0000 | 132 |
| 0.0200 | 0.0200 | 1.0000 | 266 |
| Expected FST . | Average FST . | Fisher's test . | t . |
|---|---|---|---|
| 0.0010 | 0.0010 | 0.4185 | 13 |
| 0.0025 | 0.0025 | 0.9420 | 33 |
| 0.0050 | 0.0050 | 1.0000 | 66 |
| 0.0100 | 0.0100 | 1.0000 | 132 |
| 0.0200 | 0.0200 | 1.0000 | 266 |
The power is expressed as the proportion of simulations that provide statistical significance at the 0.05 level after a number of generations (t) which are necessary to reach an average FST equal to the expected values.
Otolith chemistry
SIMPROF on the core samples analysed (Table 1) did not detect statistically different groups (Figure 3) from cluster analysis.
No significant effect of the covariate “fish size” was detected on the chemical composition of the otolith margin, suggesting an absence of temporal variability (due to the fact that specimens of different ages were analysed). The chemical composition of the otolith margin was significantly different among the sampling sites (PERMANOVA, pseudo-f: 9.306, p < 0.01). Significant differences among otoliths were also found (pseudo-f: 7.651, p < 0.01), suggesting within-site differences among individuals.
The elemental:Ca ratio contributing most to differences among sampling sites was Sr:Ca (ranging from 62.47% to 98.60% of the total dissimilarity in pairwise comparison among sites, SIMPER analysis); Ba:Ca and Mg:Ca had little influence in determining differences between groups. The three sampling sites significantly differed for the elemental ratios of Sr:Ca in the otoliths (PERMANOVA p < 0.01) and not for Ba:Ca and Mg:Ca.
Components of variation show that most of the variability is associated with the factor “otolith” (VC = 0.98, %TV = 46.6, ø = 1.49), while the variability associated with the residuals (VC = 0.65, %TV = 31.1, ø = NA) and with the factor “site” (VC = 0.45, %TV = 21.6, ø = 0.69) was lower. TL had almost no influence on total variation (VC = 0.01, %TV = 0.7, ø = 0.02).
Discussion
Microsatellite analysis clearly indicated the presence of a single panmictic population in the northern–central Adriatic Sea. Moreover, otolith chemistry analysis suggested a common natal origin for individuals sampled in the three areas of the northern–central Adriatic Sea (i.e. GSA 17). These results, together with monthly shifts in fishery landings, suggest that S. scombrus could perform an autumn–winter migration from the northern to the central Adriatic to reach a spawning ground common to different stocks in the northern–central Adriatic.
Landings data highlighted a clear north–south trend in the seasonality of the Atlantic mackerel catches: the species almost completely disappeared in catches from the northern Italian regions Friuli Venezia Giulia and Veneto, and displayed a marked reduction in Emilia Romagna in the winter months. In the Marche region, the species showed an increase in the winter months, while no clear patterns emerged in the southern region. These data suggest a late autumn migration that follows the southward–western branch of the cyclonic current in the northern–central Adriatic Sea (Russo and Artegiani, 1996). Landings data present well-known limits; indeed changes in landing may not necessarily mirror changes in stock abundance, but may rather reflect changes in fishing effort or fishing areas (Pauly et al., 1998). However, seasonal variations in catches are quite consistent among years and are supported, at least for the northern Adriatic, by the knowledge of local fishers about the movement of the study species. Therefore, these seasonal variations in landings are expected to reflect variations in local stock abundance reliably. Moreover, other pelagic fish species in the Adriatic are known to perform a similar seasonal migration in the northern–central part, following the main currents, for trophic and reproductive needs (Morello and Arneri, 2009).
With regard to population genetics, the simulation approach used revealed a high power of our markers to detect population subdivision at FST values as low as 0.0025, well below the threshold under which any possibly undetected difference would have limited ecological and biological relevance (evolutionary population criterion EV3 < 5 and EV4 < 25 effective migrants per generation; Waples and Gaggiotti, 2006). Thus, the non-significant FST values close to zero found for the northern–central Adriatic gene pool clearly point to an overall panmixia of S. scombrus, which is in line with several findings for marine species collected at a similar geographical scale in GSA 17, including pelagic [European anchovy (Engraulis encrasicolus), Borsa (2002); European sardine (Sardina pilchardus), Tinti et al. (2002)] and demersal fish [common sole (Solea solea), Garoia et al. (2007); red mullet (Mullus barbatus), Maggio et al. (2009); marbled goby (Pomatoschistus marmoratus), Mejri et al. (2011)] as well as the nekton-benthic squids and cuttlefish (Garoia et al., 2004). The occurrence of a single panmictic population in the northern–central Adriatic is somewhat expected for the Atlantic mackerel, considering the high dispersal capacity of this species at both the larval and adult stages. In fact, larvae are able to disperse over tens of kilometres in just 10 d (Jansen et al., 2012), a time frame representing about a quarter of the entire larval stage that is reported to last up to 40 d in the northern Atlantic Ocean (Villamor et al., 2004). Moreover, adults are highly migratory and can cover up to 1200 km in 13 d (Lockwood, 1988).
In contrast, our microsatellite analysis showed highly significant values of differentiation when samples collected at the northern–central sampling sites were compared with the southern Adriatic sample. This comparison involves samples collected in 2010 at the GSA 17 locations and one single “historical” sample collected in 2001 in the southern part of the Adriatic basin, since, for this location, we were not able to obtain, from commercial fishers, new samples with reliable information about the collection site.
The presence of population subdivision was demonstrated in S. scombrus, at least at a large geographic scale. In fact, a restriction in gene flow was found between spawning stocks from the eastern and western Atlantic by Nesbø et al. (2000), based on mitochondrial marker variability. At our geographic scale, the differentiation between samples from GSA 17 and 18 is, indeed, supported by a previous study (Zardoya et al., 2004) that found genetic differentiation comparing samples of S. scombrus from Ancona (July 1997) and Bari (December 2000–January 2001). In that case, Bari was similar to an Aegean sample, and a general pattern of differentiation along the east–west axis of the Mediterranean Sea was reported. The significantly higher allelic richness found for our markers in the southern–western sample (2001) supports the hypothesis of Zardoya et al. (2004) that this sample receives and/or has received genetically different individuals. Genetic differentiation between the northern–central and southern part of the Adriatic was recorded for other pelagic species, such as the European anchovy (Bembo et al., 1996; Borsa, 2002). The presence of a stable gyre separating the northern and southern Adriatic waters, a bathymetric discontinuity corresponding to the deep Jabuka (Pomo) Pits (Figure 1; Russo and Artegiani, 1996), and the strong spatial and temporal variability of trophic conditions in the northern–central Adriatic (Fonda Umani et al., 1992) may constitute physical barriers to migrations and/or to larval dispersal. In addition, behavioural factors may be crucial in structuring populations (Nesbø et al., 2000). In the Atlantic mackerel, homing of adults to spawning grounds was demonstrated in the western Atlantic (Studholme et al., 1999). Similar homing behaviour and the possible occurrence of different spawning grounds between populations from the northern–central and southern Adriatic, as found for European anchovy and European sardine, may, therefore, further contribute to maintaining genetic differentiation. The differentiation pattern found between Atlantic mackerel from the northern–central and southern Adriatic may, indeed, be a result of a complex combination of environmental (hydrodynamics) and behavioural factors. Therefore, further analyses based on samples collected simultaneously from the two GSAs are needed to confirm the subtle, but significant, genetic divergence of the southern–western sample with respect to those from the northern–central Adriatic.
Otolith chemistry revealed homogeneity in core composition of samples belonging to the three different population samples investigated in the central–northern Adriatic Sea. This evidence is suggestive of a common natal origin (i.e. spawning area) for the individuals that, as adults, were collected from the three different sampling areas investigated. It has also to be taken into account that homogeneous chemical signatures from cores could emerge from potentially multiple natal origins (i.e. spawning sites) having similar if not almost identical environmental features.
The potential of otolith microchemistry analysis in detecting spatial variability relies on features related both to fish physiology and to environmental characteristics (e.g. seawater composition, salinity, and temperature; Chang and Geffen, 2012). From this perspective, it would be ecologically meaningful to compare our results with those arising from a species with a similar life cycle and inhabiting a geographic area close to the one investigated, but, as far as we know, no studies with these characteristics are available. In the Mediterranean, some examples are available of fish showing spatial variability in core chemical composition: some refer to species with a similar life cycle, but coming from other areas of the Mediterranean [e.g. Tanner et al. (2012) for a study on European hake (Merluccius merluccius) from the western Mediterranean at a spatial scale comparable with the one adopted in the present study; and Guidetti et al. (2013) for a study on anchovy from the Ligurian Sea], and another one arises from species with different life habits, but coming from the Adriatic Sea (e.g. Di Franco et al., 2012 focusing on a coastal fish, Diplodus sargus sargus, from the southern Adriatic). This evidence supports the hypothesis that the recorded homogeneity in core composition represents a single natal origin for S. scomber in our study area and is not just based on a failure to detect different natal origins characterized by similar environmental conditions. An additional factor supporting a single natal origin is provided by the high variability in otolith margins recorded, minimizing the idea of similar environmental features among the sampling sites. It has to be stressed that we tested whether otolith chemistry in S. scombrus is able to record environmental heterogeneity by analysing potential spatial variability in the otolith portion laid down before capture (i.e. the portion of the otolith “influenced” by the environmental condition of the sampling site), but this assessment of variability reflects divergence in water chemistry at the time of sampling that does not necessarily represent water conditions at the time the fish were spawned, due to temporal variability in otolith chemistry. Temporal stability of the otolith fingerprint can be a major concern for application to natal origin discrimination and connectivity studies. Studies dealing with temporal variability in elemental signature are controversial, with some identifying significant interannual variability and others highlighting non-significant differences or differences insufficient to preclude grouping together fish caught in the same location in different years (for a review, see Chang and Geffen, 2012). Moreover, in the case of a marine species such as Atlantic mackerel, the stability of the open marine environment compared with the greater year-to-year variability in, for example, estuarine or embayment conditions would render the accounting for temporal variation less critical (Fairclough et al., 2011).
Variability in chemical composition of the otolith marginal portion was mainly induced by variability in Sr:Ca. The incorporation of trace elements into otoliths is a complex process that is still not completely understood. It is potentially influenced by a number of factors such as salinity, temperature, water chemistry, physiology, and metabolism (see Green et al., 2009, and references therein). Otolith Sr concentration is the result of the interaction of concentrations of chemicals in the seawater, temperature, and salinity (Chang and Geffen, 2012). As far as we know, no data about Sr water concentration in the sampled area are available, but clear variability in temperature and salinity was demonstrated during the summer (i.e. the season when samples were collected; Cushman-Roisin et al., 2007), potentially driving the recorded difference in otolith chemistry.
Our data allow the depiction of a scenario characterized by a single and common spawning area of Atlantic mackerel in the central–northern Adriatic Sea, where adults migrate and converge at reproduction, a hypothesis supported by the disappearance of ripe males and females from the northern Adriatic during the breeding season (C. Mazzoldi, unpubl. data) which corresponds well with the hypothesis that individuals disperse after reproduction towards different areas (Sinovčić, 2001; Bottari et al., 2004). Atlantic mackerel eggs have been found in the eastern Adriatic (Hure, 1961) around the area where the European anchovy and European sardine are also known to spawn (Morello and Arneri, 2009). This area is characterized by upwelling phenomena (Morello and Arneri, 2009) and could, therefore, constitute a suitable spawning ground for S. scombrus as well.
Conclusions
Our study points out the presence of a single population of S. scombrus in the central–northern Adriatic that depends on a single spawning area. These results, together with evidence of a decline in maximum age (from 8 to 3 years) and size (from 420 to 360 mm TL), which occurred in the same area in the last 20 years (C. Mazzoldi, unpubl. data), definitively suggest that the Atlantic mackerel stocks are at high risk and should be managed as a single population. Our results regarding S. scombrus population connectivity, coupled with the knowledge of migration patterns and the precise location of the spawning ground, provide an ample basis for developing effective management strategies, possibly including a fishing ban during spawning periods in reproductive areas to reduce fishing pressure on spawning aggregations.
Acknowledgements
For sample collection, we thank M. La Mesa and F. Donato (Ancona, 2010), C. Meneghesso and E. Riginella (Chioggia, 2010), and S. Greco (Bari, 2001). Many thanks are due to B. Zannato for her support in microsatellite genotyping, and G. De Benedetto and A. Pennetta for their invaluable help in the otolith chemistry analyses. We wish to thank the Editor and two anonymous reviewers for their constructive criticisms and comments that helped in improving the manuscript. This work was part of the project CLODIA funded by the Veneto Region (Italy) Law 15/2007 (DGR n. 4069).
References
Author notes
These authors contributed equally to this work.
Handling editor: Emory Anderson