-
PDF
- Split View
-
Views
-
Cite
Cite
Marco Maccaferri, Maria Corinna Sanguineti, Andrea Demontis, Ahmed El-Ahmed, Luis Garcia del Moral, Fouad Maalouf, Miloudi Nachit, Nasserlehaq Nserallah, Hassan Ouabbou, Sayar Rhouma, Conxita Royo, Dolors Villegas, Roberto Tuberosa, Association mapping in durum wheat grown across a broad range of water regimes, Journal of Experimental Botany, Volume 62, Issue 2, January 2011, Pages 409–438, https://doi.org/10.1093/jxb/erq287
Close - Share Icon Share
Abstract
Association mapping was used to dissect the genetic basis of drought-adaptive traits and grain yield (GY) in a collection of 189 elite durum wheat accessions evaluated in 15 environments highly different for water availability during the crop cycle (from 146 to 711 mm) and GY (from 9.9 to 67.3 q ha−1). For highly heritable traits (e.g. heading date, kernel weight, etc.) several significant experiment-wise marker-trait associations were detected across five or more (up to 13 for kernel weight) environments, with R2 values ranging from ca. 5 to 10%. As to GY, significant associations (R2 from 2.5 to 4.2%) were mostly detected in one environment only (56 markers), while decreasing rapidly from two to five environments (from 20 to three markers, respectively) and with only one marker (Xbarc197 on chr. 5A) found significant in six environments (ranging from low- to high-yielding). These results are probably due to the complex genetic basis of GY and its interaction with environmental conditions. The number of markers significantly affecting GY decreased considerably under drought conditions, suggesting a limited effectiveness of association mapping to identify loci for GY under low-moisture conditions, most likely because different genotypes can attain similar phenotypes via different morpho-physiological traits and corresponding gene networks. Our study confirmed the role of major loci for phenology previously described in biparental mapping populations, highlighted a novel set of loci for drought-adaptive traits, and provided information on the agronomic value of the alleles at such loci across a broad range of soil moisture conditions.
Introduction
Coping with the consequences of climate change on agriculture will require unprecedented efforts toward the release of improved cultivars (cvs.) more resilient to weather vagaries, particularly drought (Semenov and Halford, 2009). In this context, a better knowledge of the genetic make-up of yield and the main traits that underlie the adaptive response of crops across a broad range of soil moisture is an essential prerequisite toward more effective and targeted breeding activities (Araus et al., 2002; Salekdeh et al., 2009). Accordingly, genomics approaches enable the identification and selection of chromosome (chr.) regions harbouring genes/QTLs (Quantitative Trait Loci) controlling agronomic traits and yield in crops (Tuberosa and Salvi, 2006; Collins et al., 2008; Cooper et al., 2009). Among such approaches, association mapping (AM) is increasingly being adopted as a method complementary to traditional bi-parental linkage mapping to identify genotype–phenotype associations (i.e. molecular marker–trait associations; Breseghello and Sorrells, 2006; Ravel et al., 2006; Ersoz et al., 2008; Sorrells and Yu, 2009; Waugh et al., 2009). Panels of accessions characterized by medium to high linkage disequilibrium (LD) levels are suitable to carry out genome-wide searches for chr. regions harbouring loci with allelic variants that regulate the phenotypic expression of target traits. Importantly, genome-wide scans can be conducted at a reasonably low marker density for those species currently lacking high density maps, such as wheat (Chao et al., 2007; Horvath et al., 2009; Sorrells and Yu, 2009; Rafalski, 2010). From a biological standpoint, the combined phenotypic and molecular characterization of the natural variation present in core collections of germplasm is valuable for genetic dissection of complex traits in general (Alonso-Blanco et al., 2009) and, more specifically, for traits involved in the adaptive response to different water regimes and, consequently, yield (Araus et al., 2008; Annicchiarico et al., 2009; Habash et al., 2009; Reynolds et al., 2009; Tardieu and Tuberosa, 2010). Further, on a more applicative side, the results of AM experiments are valuable for marker-assisted breeding programmes as they allow for the identification of the most desirable allelic variants at the major loci controlling the target traits (Heffner et al., 2009).
While AM has recently been suggested as a suitable approach for investigating the genetic basis of agronomically relevant traits in improved germplasm (Reynolds et al., 2009), its potential has not yet been tested in conditions of varying water regimes. In durum wheat (Triticum durum Desf.), the use of whole-genome AM has been limited to the study of the genetic control under non-stressed conditions of highly heritable traits related to kernel quality (Reimer et al., 2008) and disease resistance (Maccaferri et al., 2009). In addition, also in bread wheat (Triticum aestivum L.), the AM studies so far reported have all been conducted under rather favourable conditions, at least in terms of soil moisture (Breseghello and Sorrells, 2006; Crossa et al., 2007; Yao et al., 2009).
Durum wheat is well-adapted to the Mediterranean regions (Belaid, 2000), where ca. 75% and 50% of the worldwide acreage and production, respectively, are concentrated (Elias and Manthey, 2005). Its range of cultivation spans areas with different levels of fertility, ranging from favourable conditions (e.g. temperate regions, with medium to high annual rainfall rate and/or irrigation) to dryland areas characterized by drought and high temperature stresses, with an increased occurrence of water shortage and heat waves in late spring, i.e. during grain filling (Loss and Siddique, 1994; Katerji et al., 2008). Therefore, terminal drought, the stress most frequently experienced by wheat grown in the Mediterranean Basin, negatively affects grain weight and grain volume weight (Araus et al., 2003; Slafer et al., 2005), two very important yield-related traits for the milling industry. Furthermore, due to the high variability in rainfall pattern, water shortage in Mediterranean environments (envs.) can also occur from sowing time (e.g. late autumn) to early spring. In such cases, drought exerts even stronger negative effects on yield, impacting yield potential at the sink level, for example by decreasing the number of fertile spikes per unit area at the crop establishment and tillering phases, as well as the number of grains per spike (Garcia del Moral et al., 2003; Annicchiarico et al., 2005; Alvaro et al., 2008). Under such conditions, adaptation mechanisms, based on traits that allow the plant to escape (e.g. flowering date), avoid (e.g. root depth), and/or tolerate (e.g. osmolyte accumulation) the negative effects of drought, influence the balance and competition between vegetative and reproductive organs, thus playing a major role in determining final crop performance (Blum, 1988; Passioura, 2002; Tuberosa and Salvi, 2004; Araus et al., 2008; Reynolds and Tuberosa, 2008). It should be noted that the Mediterranean Basin is one of the geographical regions most densely populated where agriculture has been forecast to be at greater risk due to an increase in the frequency and severity of drought episodes (Reynolds et al., 2007).
In this study, AM was used to identify QTLs influencing the agronomic performance of durum wheat grown across a broad range of soil moisture conditions. A series of chr. regions was found to significantly affect the variability of traits related to plant adaptation (heading date), plant response to water shortage (plant height, ear peduncle elongation, and grain volume weight), yield components (ear fertility and grain weight), and grain yield is reported here. Further, the association results have been analysed in order to ascertain the presence of different marker–trait association patterns across widely differing environments as to water availability and, consequently, grain yield.
Materials and methods
Plant material
This study evaluated a collection of 189 durum wheat elite accessions (mainly cvs. and advanced lines) bred in Mediterranean countries (Italy, Morocco, Spain, Syria, and Tunisia), Southwestern USA, and Mexico. Most of the accessions are semi-dwarf elite materials released from the early 1970s to the late 1990s. The collection, reported hereafter as IDuWUE panel, also includes a number of important ‘founder genotypes’ widely used as parents in breeding programmes throughout the Mediterranean Basin. The detailed list of genotypes and their origins are presented in Supplementary Table S1 at JXB online. The accessions included in the collection were chosen from 330 accessions evaluated in a comparative field trial carried out in 2003 in Cadriano, near Bologna, Italy (M Maccaferri et al., unpublished data). The choice was mainly based on the analysis of the pedigree data and the crop cycle length: highly related accessions as well as excessively early or late heading accessions were excluded from the IDuWUE panel.
Field trials and phenotypic traits
Fifteen field trials were carried out in two growing seasons (seven and eight trials in 2003/2004 and 2004/2005, respectively) in Italy, Lebanon, Morocco, Spain, Syria, and Tunisia. Field trials (hereafter considered as single envs.) were coded according to the country (first three letters of each code), the water regime (with ‘r’ and ‘i’ standing for rainfed and irrigated trial, respectively) and the year (with 04 and 05 standing for 2004 and 2005, respectively). Trials were conducted in Italy (Itl1-r04 in Cadriano, 44° 33' N and 11° 24' E; Itl2-r04 and Itl2-r05 in Cerignola, 41° 28' N and 15° 84' E), Lebanon (Lbn-r04, Lbn-i04, Lbn-r05, and Lbn-i05 in Rayack, 33° 51' N and 35° 59' E), Morocco (Mrc-r04 and Mrc-i04 in Sidi Elaydi, 31° 15' N and 7° 30' W), Spain (Spn1-r04 in Gimenells, 38° 56' N and 0° 29' E; Spn2-r05 in Granada, 37° 15' N and 3° 46' E), Syria (Syr-r05 and Syr-i05 in Tel Hadya, 36° 56' N and 36° 04' E), and Tunisia (Tns-r05 and Tns-i05 in Kef, 36° 14' N and 8° 27' E).
The following environmental factors, which are known to be most critical for durum wheat performance under Mediterranean conditions, have been considered: (i) temperatures from emergence to heading and from heading to harvest (maximum and mean temperatures across the two phases), (ii) water availability to the crop (considered as water input, Wi: rainfall plus supplementary irrigation) from emergence to heading and from heading to harvest, (iii) average soil moisture (Sm, from 0 to 30 cm deep) evaluated as area under the soil water content curve measured in four critical periods: from jointing (i.e. the stage at which the internodal tissue in the grass leaf begins to elongate) to heading, from 1 week before heading to 3 weeks after heading, in the whole period from jointing to maturity, and from heading to maturity (see Supplementary Table S2 at JXB online).
The accessions were tested in unreplicated plots of 4 m2 (with eight rows, 0.20 m apart and 2.5 m long). Experimental layouts were prepared according to a modified augmented design with three checks (cvs. Simeto and Vitron as checks common to all field trials, plus one additional check as the best locally adapted cv.) randomly distributed in each row of the field schemes. Simeto (pedigree: Capeiti 8/Valnova, from the Italian germplasm) has been the first-ranking cv. in cultivation acreage in Italy since the early 1980s and is still widely cultivated throughout the Mediterranean Basin. Vitron is a high-yielding cv. developed from the CIMMYT germplasm, released in 1985 and characterized by high yield stability in the Mediterranean Basin (Pfeiffer et al., 2000). Seed of the accessions and checks was increased during 2003 in a single location (Cadriano, Bologna). Seeding rates were adjusted according to the results of a seed germination test to obtain a plant density from 400–450 viable seeds m−2, depending on the plant density locally adopted. Fields were fertilized and managed following the standard agricultural practices adopted in each location; weeds were chemically and mechanically controlled and, where necessary, the trials were protected against the main pathogens. In five trials, supplementary irrigation was provided with one to three applications mainly at stem elongation, booting, and/or flowering (irrigation volumes from 300 m3 ha−1 in Syr-i05 to 1000 m3 ha−1 in Lbn-i05). Water availability (rainfall plus supplementary irrigation) from emergence to harvest ranged from 170 mm (Mrc-r04) to 711 mm (Lbn-i04).
The following traits were considered for the AM analysis: heading date (HD), plant height (PH), ear peduncle length (PdL), grain yield (GY), thousand kernel weight (TKW), grain volume weight or test weight (TW), and number of grains per square metre (Kpsm). HD is reported as the number of days from seedling emergence to the date when ca. 50% of the ears had emerged from the flag leaf sheath for half of their length (stage 55 in the Zadoks scale; Zadoks et al., 1974). PH (cm) was measured at maturity from the ground to the ear tip (excluding awns) on three main stems per plot. PdL (cm) was measured as the distance from the flag leaf collar to the base of the ear on the same main stems used to measure PH. TKW (g) was measured on two samples of 100 seeds per plot. TW (kg hl−1) was estimated on a seed sample of ca. 300 g using the Grain Analysis Computer GAC 2100 (Dickey-John Corporation, Minneapolis, MN, USA). GY (q ha−1) and TKW were adjusted at 14% moisture. Kpsm data were calculated from GY and TKW data.
For each environment, phenotypic data were analysed by restricted maximum likelihood (REML) to fit a mixed model with checks as a fixed effect and rows, columns, and unreplicated entries as random effects (Littel et al., 1996). The REML model produced best linear unbiased predictors (BLUPs) for the phenotypic data of each genotype at each env. to be used in subsequent analyses. The analysis was performed using the MIXED procedure of the SAS statistical package (SAS, 2001).
BLUPs were then tested for normality using the Kolmogorov–Smirnov goodness of fit test in Minitab 15 (http://www.minitab.com/en-IT/products/minitab/) and used to perform a combined analysis considering environments as replicates.
Molecular profiling
For each accession, genomic DNA was extracted from a bulk of ca. 20 freeze-dried young leaves per accession following the method described in Saghai-Maroof et al. (1984). In our study, the panel of accessions investigated by Maccaferri et al. (2006) was profiled with an additional set of 90 SSRs, including: (i) the publicly available sets catalogued in the GrainGenes database (http://wheat.pw.usda.gov) as BARC (Xbarc marker loci; Song et al., 2005), CFA and CFD (Xcfa and Xcfd; Sourdille et al., 2001), WMC (Xwmc; Gupta et al., 2002), and WMS (Xgwm; Röder et al., 1998) sets, and (ii) a small additional subset of private genomic WMS primers developed and owned by TraitGenetics (supplied by Martin Ganal, TraitGenetics, Gatersleben, Germany). In total, 186 uniformly spaced genomic SSRs were used to profile the IDuWUE panel.
To explore LD and association in specific chr. regions harbouring important genes/QTLs on chrs. 2A and 5A, a few SSR loci were subsequently added (Xcfa2201 on chr. 2A; Xbarc117, Xwmc752, Xwmc705, and Xwmc805 on chr. 5A). The SSR markers used for genotyping were chosen among those that were polymorphic in two durum recombinant inbred line (RIL) populations used in previous studies (Maccaferri et al., 2008; Mantovani et al., 2008). The map position of the SSR loci used here was obtained by computing a combined map from the SSR data of the two above-mentioned RIL populations using JoinMap v. 4.0 (Van Ooijen, 2006). The SSR primers were further selected for producing clear and high-quality amplification banding patterns and for detecting polymorphisms in a subset of eight founder accessions (Cappelli, Capeiti 8, Valnova, Creso, Cham 1, Omrabi3, Karim=Yavaros 79, and Gallareta=Altar 84; the accessions are listed in chronological order of release from 1915 to 1984).
The SSR amplification was carried out according to Röder et al. (1998), except that forward primers were labelled with the IR700/IR800 fluorophore dyes. Reactions were performed in 2720 Perkin-Elmer thermocyclers (Norwalk, CT, USA) using amplification cycles according to the melting temperatures of the primers. Electrophoresis and data collection were carried out with the automated Li-Cor 4200 IR2 System (Li-Cor, Lincoln, NE, USA) using 25 cm-long and 0.4 mm-thick, 6% polyacrylamide gels and 64-well plastic combs. Samples were diluted 1 to 6 (v/v) in denaturing formamide loading buffer and 0.8 μl per sample of the diluted PCR product was loaded into the gel wells.
Polymorphism at Vrn-A1 (vernalization response), Ppd-A1 (photoperiod response), and RhT-B1 (semi-dwarfing) genes was also analysed. For Vrn-A1 (chr. 5A), the presence of polymorphism was analysed first in the promoter region (primer pair VRN1AF/VRN1R) of the gene, according to Yan et al. (2004); subsequently, the presence of the deletion in the first intron was investigated according to Fu et al. (2005) using the two primer pairs Ex1CF/Intr1AR3 and Intr1CF/Intr1ABR and the cv. Langdon as positive control. For Ppd-A1 (chr. 2A), the presence of wide deletions upstream of the coding region was investigated with a single PCR reaction carried out with Ag5del_F1, Ag5del_F2, and Ag5del_R2 primers (Wilhelm et al., 2009). This PCR assay amplified three fragments of different molecular weight associated to the three main Ppd-A1 alleles present in the germplasm, i.e. either the intact sequence (i.e. the wild type, photoperiod-sensitive allele, amplifying a DNA fragment of 452 bp) or two different promoter deletions of 1027 and 1117 bp, respectively (both associated with photoperiod insensitivity and tagged by DNA fragments of 380 bp and 290 bp, respectively; Wilhelm et al., 2009). At the Rht-B1 gene, the presence of either the Rht-B1b dwarfing allele or the Rht-B1a allele for tall stature was checked with the functional markers amplified using the primer pairs BF/MR1 and BF/WR1 according to Ellis et al. (2002).
Genetic structure and linkage disequilibrium analysis
The genetic structure of the IDuWUE panel has been investigated with a combination of model- and distance-based analyses. As a first step, the SSR data were used to estimate the LD values (r2 and D' statistics; Weir, 1996) among loci and their significance (P values) on a genome-wide basis. Rare alleles (less than 18 counts, i.e. with frequency <0.10) were excluded from the analysis, thus reducing their bias on LD estimates. Data-points showing residual allelic heterogeneity within accession (presence of two alleles) were considered as missing data. More in detail, the D' and r2 LD values for loci with multiple alleles (Farnir et al., 2000) were calculated using the software TASSEL (http://www.maizegenetics.net; Bradbury et al., 2007) and the LD significance (P values) was determined with 10 000 permutations. Based on the LD results, a subset of 96 loosely linked and evenly spread SSRs with null or limited LD (caused by true linkage) were chosen to investigate the population structure of the IDuWUE panel using the Bayesian model-based clustering method implemented in the software STRUCTURE v. 2.1 (http://pritch.bsd.uchicago.edu/structure.html; Pritchard et al., 2000). The admixture model with correlated allele frequencies (Falush et al., 2003) was used to test for the presence of genetic structure with a number of hypothetical subgroups ranging from two to ten. No prior origin or phenotypic information was used to define the subgroups. Burn-in period and simulation run length were set at 100 000 iterations. For each accession, the membership coefficients (Q population structure coefficients) to all subgroups were calculated. Five independent runs were carried out for each number of subgroups being tested to compare the consistency of results across independent runs; for each value, the run with the highest posterior probability of data was chosen as the representative run.
In the distance-based analysis, all SSR data (including the rare alleles) were used to calculate the pairwise genetic similarity values (GSij) using the simple matching coefficient for multi-state markers, i.e. the proportion of loci with shared alleles (Lu and Bernardo, 2001; Matsuoka et al., 2002; Maccaferri et al., 2003). Cluster analysis of the similarity matrix was carried out with SAHN (Sequential Agglomerative Hierarchical and Nested clustering method) and UPGMA (Unweighted Pair-Group Method with Arithmetic mean). The analysis was carried out in NTSYS-pc software as reported in Maccaferri et al. (2007a) using SIMQUAL for the calculation of the genetic distances, SAHN for the cluster analysis, and TREE PLOT for the graphic visualization of the dendrogram.
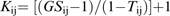
For the 857 SSR pairs of physically linked (syntenic) loci (i.e. on the same chr.), D' and r2 values were plotted against the corresponding map distances and locally weighted polynomial regression (LOESS) based-fitting curves were added to the plots. The distribution of the 14 896 pairs of unlinked markers was used to set the D' and r2 critical values (95th percentile of square root transformed data) to be considered as reliable evidence of true linkage between syntenic marker loci, with associations not due to main population genetic structure effects. The average extent of LD across all the linkage groups was thus estimated based on the intersections of the LOESS-curves with the LD significance baselines.
Association analysis
The influence of the genetic diversity structure present in the IDuWUE panel on the phenotypic variation of target traits was assessed by means of multiple regression. Significance and R2 values of the multiple regression of phenotypes on genetic diversity structure were calculated with Minitab.
Association analyses were carried out in TASSEL 2.0.1 software (http://www.maizegenetics.net/). The significance of marker-phenotype associations was tested using: (i) the fixed general linear model (GLM) including the Q population structure coefficients as covariates, (ii) the mixed linear model (MLM) including the Q population structure coefficients and the 189×189 K kinship matrices based on either Loiselle's kinship (Q+KLois) coefficient (MLMLois) or (iii) the identity-by-descent genetic similarity estimates (Q+KSM; MLMSM). Each env. was analysed separately.
Marker–trait associations were considered reliable when the significance was detected with the three association tests mentioned above. The following significance thresholds were considered: P 0.05 and P 0.01 marker-wise level and P 0.05 experiment-wise level (Bonferroni's correction).
For markers significantly associated with the phenotype, the R2 values of the marker and of the model (including both covariate and marker effects) as well as the least square means of the allelic variants present in the germplasm were calculated and, when significance was detected for loci with three or more non-rare alleles, the least significant difference between the allele means was calculated.
Results
Variation in phenology, grain yield, and yield components
The accessions were evaluated for phenological traits such as HD, PH, and PdL, GY and its components in 15 envs. across six Mediterranean countries. For all the investigated traits, a clear quantitative genetic basis was evidenced (see the phenotypic distributions reported as Supplementary Fig. S1 at JXB online). Considering HD values across all envs., it is worth noting that 167 out of 189 accessions (88%) reached heading in a 4 d interval, with all the accessions heading in a 9 d interval.
As shown in Table 1, the mean GY of the accessions comprised between a minimum of 9.9 q ha−1 (in Spn2-r05) and a maximum of 67.3 q ha−1 in Tns-i05. Within this broad range, the average GY was as follows: (i) less than 30 q ha−1 in three envs. characterized by severe water stress (hereafter identified as low-yielding) with GY equal to 9.9 (Spn2-r05), 22.2 (Lbn-r04), and 29.6 q ha−1 (Mrc-r04); (ii) from 35.3 to 45.7 q ha−1 in five envs. (medium-yielding: Mrc-i04, Lbn-r05, Lbn-i04, Tns-r-05, and Syr-r05), and (iii) from 54.3 to 67.3 q ha−1 in seven envs. (high-yielding: Lbn-i05, Itl2-r05, Syr-i05, Itl2-r04, Spn1-r04, Itl1-r04, and Tns-i05, listed following an increasing GY value). The very low GY in the Spn2-r05 trial can be attributed to low rainfall during the entire crop cycle (a total of 187 mm from emergence to harvest) and, in particular, from heading to harvest (12 mm), which led to very low average soil moisture (see Supplementary Table S2 at JXB online). The broad variation in GY (ca. 7-fold range) underlines that the field trials, in line with the main objective of the study, were managed under highly different environmental conditions, i.e. from favourable conditions in terms of water availability to severe drought- and/or heat-stressed conditions. In each env., the accessions showed widely different GY (e.g. from 31.5 to 87.1 q ha−1 and from 6.6 to 12.3 q ha−1 in the most and in the least productive env., respectively; Table 1), thus indicating that the panel considered here encompassed a broad range of genetic and functional diversity.
Mean grain yield values (GY, q ha−1) and range of variation for the 189 durum wheat accessions of the IDuWUE panel and the three checks evaluated in 15 field trials carried out in 2004 and 2005
| Environments | Accessions | Check cultivars | ||||
| Mean | Min | Max | Simeto | Vitron | Iride | |
| High-yielding | ||||||
| Tns-i05a | 67.32 | 31.50 | 87.05 | 69.11 | 68.88 | 71.17 |
| Itl1-r04 | 64.40 | 40.40 | 110.40 | 68.30 | 58.00 | 66.70 |
| Spn1-r04 | 64.17 | 41.12 | 80.59 | 67.75 | 57.79 | 72.75 |
| Itl2-r04 | 58.58 | 25.70 | 74.50 | 55.15 | 52.71 | 59.20 |
| Syr-i05 | 58.28 | 35.80 | 85.60 | 64.00 | 57.78 | 57.95 |
| Itl2-r05 | 55.96 | 41.20 | 68.45 | 60.53 | 54.86 | 62.60 |
| Lbn-i05 | 54.29 | 25.00 | 95.00 | 64.92 | 59.92 | 65.83 |
| Medium-yielding | ||||||
| Syr-r05 | 45.74 | 22.90 | 78.40 | 50.67 | 44.62 | 51.42 |
| Tns-r05 | 43.73 | 24.35 | 64.30 | 42.95 | 46.13 | 42.65 |
| Lbn-i04 | 40.88 | 9.73 | 74.80 | 41.72 | 36.51 | 44.95 |
| Lbn-r05 | 35.75 | 15.00 | 61.67 | 35.95 | 33.55 | 41.11 |
| Mrc-i04 | 35.30 | 10.98 | 69.64 | 37.13 | 38.93 | 39.93 |
| Low-yielding | ||||||
| Mrc-r04 | 29.58 | 9.14 | 51.25 | 27.62 | 32.50 | 34.33 |
| Lbn-r04 | 22.16 | 4.90 | 40.20 | 19.93 | 20.81 | 26.10 |
| Spn2-r05 | 9.93 | 6.56 | 12.28 | 9.93 | 9.88 | 9.98 |
| Environments | Accessions | Check cultivars | ||||
| Mean | Min | Max | Simeto | Vitron | Iride | |
| High-yielding | ||||||
| Tns-i05a | 67.32 | 31.50 | 87.05 | 69.11 | 68.88 | 71.17 |
| Itl1-r04 | 64.40 | 40.40 | 110.40 | 68.30 | 58.00 | 66.70 |
| Spn1-r04 | 64.17 | 41.12 | 80.59 | 67.75 | 57.79 | 72.75 |
| Itl2-r04 | 58.58 | 25.70 | 74.50 | 55.15 | 52.71 | 59.20 |
| Syr-i05 | 58.28 | 35.80 | 85.60 | 64.00 | 57.78 | 57.95 |
| Itl2-r05 | 55.96 | 41.20 | 68.45 | 60.53 | 54.86 | 62.60 |
| Lbn-i05 | 54.29 | 25.00 | 95.00 | 64.92 | 59.92 | 65.83 |
| Medium-yielding | ||||||
| Syr-r05 | 45.74 | 22.90 | 78.40 | 50.67 | 44.62 | 51.42 |
| Tns-r05 | 43.73 | 24.35 | 64.30 | 42.95 | 46.13 | 42.65 |
| Lbn-i04 | 40.88 | 9.73 | 74.80 | 41.72 | 36.51 | 44.95 |
| Lbn-r05 | 35.75 | 15.00 | 61.67 | 35.95 | 33.55 | 41.11 |
| Mrc-i04 | 35.30 | 10.98 | 69.64 | 37.13 | 38.93 | 39.93 |
| Low-yielding | ||||||
| Mrc-r04 | 29.58 | 9.14 | 51.25 | 27.62 | 32.50 | 34.33 |
| Lbn-r04 | 22.16 | 4.90 | 40.20 | 19.93 | 20.81 | 26.10 |
| Spn2-r05 | 9.93 | 6.56 | 12.28 | 9.93 | 9.88 | 9.98 |
Environments are listed in decreasing order according to the mean GY values. Codes for high-, medium- and low-yielding environments are in bold,italic, and regular font, respectively.
Details on the environment codes are provided in the Materials and methods.
Mean grain yield values (GY, q ha−1) and range of variation for the 189 durum wheat accessions of the IDuWUE panel and the three checks evaluated in 15 field trials carried out in 2004 and 2005
| Environments | Accessions | Check cultivars | ||||
| Mean | Min | Max | Simeto | Vitron | Iride | |
| High-yielding | ||||||
| Tns-i05a | 67.32 | 31.50 | 87.05 | 69.11 | 68.88 | 71.17 |
| Itl1-r04 | 64.40 | 40.40 | 110.40 | 68.30 | 58.00 | 66.70 |
| Spn1-r04 | 64.17 | 41.12 | 80.59 | 67.75 | 57.79 | 72.75 |
| Itl2-r04 | 58.58 | 25.70 | 74.50 | 55.15 | 52.71 | 59.20 |
| Syr-i05 | 58.28 | 35.80 | 85.60 | 64.00 | 57.78 | 57.95 |
| Itl2-r05 | 55.96 | 41.20 | 68.45 | 60.53 | 54.86 | 62.60 |
| Lbn-i05 | 54.29 | 25.00 | 95.00 | 64.92 | 59.92 | 65.83 |
| Medium-yielding | ||||||
| Syr-r05 | 45.74 | 22.90 | 78.40 | 50.67 | 44.62 | 51.42 |
| Tns-r05 | 43.73 | 24.35 | 64.30 | 42.95 | 46.13 | 42.65 |
| Lbn-i04 | 40.88 | 9.73 | 74.80 | 41.72 | 36.51 | 44.95 |
| Lbn-r05 | 35.75 | 15.00 | 61.67 | 35.95 | 33.55 | 41.11 |
| Mrc-i04 | 35.30 | 10.98 | 69.64 | 37.13 | 38.93 | 39.93 |
| Low-yielding | ||||||
| Mrc-r04 | 29.58 | 9.14 | 51.25 | 27.62 | 32.50 | 34.33 |
| Lbn-r04 | 22.16 | 4.90 | 40.20 | 19.93 | 20.81 | 26.10 |
| Spn2-r05 | 9.93 | 6.56 | 12.28 | 9.93 | 9.88 | 9.98 |
| Environments | Accessions | Check cultivars | ||||
| Mean | Min | Max | Simeto | Vitron | Iride | |
| High-yielding | ||||||
| Tns-i05a | 67.32 | 31.50 | 87.05 | 69.11 | 68.88 | 71.17 |
| Itl1-r04 | 64.40 | 40.40 | 110.40 | 68.30 | 58.00 | 66.70 |
| Spn1-r04 | 64.17 | 41.12 | 80.59 | 67.75 | 57.79 | 72.75 |
| Itl2-r04 | 58.58 | 25.70 | 74.50 | 55.15 | 52.71 | 59.20 |
| Syr-i05 | 58.28 | 35.80 | 85.60 | 64.00 | 57.78 | 57.95 |
| Itl2-r05 | 55.96 | 41.20 | 68.45 | 60.53 | 54.86 | 62.60 |
| Lbn-i05 | 54.29 | 25.00 | 95.00 | 64.92 | 59.92 | 65.83 |
| Medium-yielding | ||||||
| Syr-r05 | 45.74 | 22.90 | 78.40 | 50.67 | 44.62 | 51.42 |
| Tns-r05 | 43.73 | 24.35 | 64.30 | 42.95 | 46.13 | 42.65 |
| Lbn-i04 | 40.88 | 9.73 | 74.80 | 41.72 | 36.51 | 44.95 |
| Lbn-r05 | 35.75 | 15.00 | 61.67 | 35.95 | 33.55 | 41.11 |
| Mrc-i04 | 35.30 | 10.98 | 69.64 | 37.13 | 38.93 | 39.93 |
| Low-yielding | ||||||
| Mrc-r04 | 29.58 | 9.14 | 51.25 | 27.62 | 32.50 | 34.33 |
| Lbn-r04 | 22.16 | 4.90 | 40.20 | 19.93 | 20.81 | 26.10 |
| Spn2-r05 | 9.93 | 6.56 | 12.28 | 9.93 | 9.88 | 9.98 |
Environments are listed in decreasing order according to the mean GY values. Codes for high-, medium- and low-yielding environments are in bold,italic, and regular font, respectively.
Details on the environment codes are provided in the Materials and methods.
Similarly to GY, the mean values of the IDuWUE panel for yield components and phenological traits considered here also showed a wide variation (data not reported). Depending on the environment, the average HD ranged from 100 to 172 d, PH from 52 to 92 cm, PdL from 6 to 20 cm, Kpsm from 3500 to 18 378, TKW from 28.4 to 49.9 g, and TW from 70.3 to 82.8 kg hl−1.
The relationships between the critical environmental factors and the phenotypic traits were evaluated by means of correlation analysis (Table 2). The water input (Wi) in the vegetative phase was significantly related to HD only (r=0.46), while Wi in the critical phase from heading to maturity was significantly related to PH, PdL, and TKW, with the highest correlation observed for PdL (r=0.59). Higher associations were, in general, observed when the water available to the crop was estimated by considering the soil moisture (0–30 cm soil layer, Sm), rather than the raw water input. In fact, the soil moisture in the rapid-growth vegetative phase up to heading (Sm Jn-HD) was significantly and positively related to HD (r=0.69), TKW (r=0.46), TW (r=0.58), and GY (r=0.40). Sm in the reproductive phase from 1 week before heading up to maturity was positively associated with PdL, Kpsm, TW, and GY, with r values ranging from 0.45 to 0.57. Compared with PH, PdL always showed the highest and more significant correlations with the environmental factors which are critical to GY. Finally, the association between Sm and GY was higher from jointing to heading and at heading rather than in the period from heading to maturity.
Correlations between the environmental factors and the phenotypic traits (heading date, HD; plant height, PH; ear peduncle length, PdL; number of grains m−2, Kpsm; thousand kernel weight, TKW; test weight, TW; grain yield, GY) across environments (15 envs. for HD, Kpsm, TKW, and GY; 13 envs. for PH and PdL, and 11 envs. for TW)
| Environmental factorsa | HD | PH | PdL | Kpsm | TKW | TW | GY |
| Water input (WI) | |||||||
| Wi Em-HD | 0.46* | –0.11 ns | –0.15 ns | –0.18 ns | 0.11 ns | –0.06 ns | –0.12 ns |
| Wi HD-MD | –0.23 ns | 0.41* | 0.59* | 0.12 ns | 0.55* | 0.10 ns | 0.32 ns |
| Soil moisture (Sm) | |||||||
| Sm Jn-HD | 0.69** | 0.18 ns | 0.39 ns | 0.22 ns | 0.46* | 0.58* | 0.40* |
| Sm HDb | 0.45* | 0.26 ns | 0.46* | 0.49* | 0.36 ns | 0.13 ns | 0.57* |
| Sm Jn-MD | 0.43* | 0.20 ns | 0.43* | 0.14 ns | 0.38 ns | 0.54* | 0.28 ns |
| Sm HD-MD | –0.27 ns | 0.14 ns | 0.34 ns | –0.08 ns | 0.02 ns | –0.15 ns | –0.08 ns |
| Temperature (T) | |||||||
| Tmean Em-HD | –0.46* | –0.19 ns | –0.37 ns | –0.40* | –0.13 ns | 0.13 ns | –0.43* |
| Tmean HD-MD | 0.26 ns | –0.16 ns | –0.02 ns | 0.20 ns | –0.01 ns | –0.30 ns | 0.24 ns |
| Tmax Em-HD | –0.43* | –0.55* | –0.68** | –0.62** | –0.22 ns | 0.56* | –0.62** |
| Tmax HD-MD | 0.17 ns | –0.32 ns | –0.21 ns | 0.10 ns | –0.07 ns | –0.22 ns | –0.22 ns |
| Environmental factorsa | HD | PH | PdL | Kpsm | TKW | TW | GY |
| Water input (WI) | |||||||
| Wi Em-HD | 0.46* | –0.11 ns | –0.15 ns | –0.18 ns | 0.11 ns | –0.06 ns | –0.12 ns |
| Wi HD-MD | –0.23 ns | 0.41* | 0.59* | 0.12 ns | 0.55* | 0.10 ns | 0.32 ns |
| Soil moisture (Sm) | |||||||
| Sm Jn-HD | 0.69** | 0.18 ns | 0.39 ns | 0.22 ns | 0.46* | 0.58* | 0.40* |
| Sm HDb | 0.45* | 0.26 ns | 0.46* | 0.49* | 0.36 ns | 0.13 ns | 0.57* |
| Sm Jn-MD | 0.43* | 0.20 ns | 0.43* | 0.14 ns | 0.38 ns | 0.54* | 0.28 ns |
| Sm HD-MD | –0.27 ns | 0.14 ns | 0.34 ns | –0.08 ns | 0.02 ns | –0.15 ns | –0.08 ns |
| Temperature (T) | |||||||
| Tmean Em-HD | –0.46* | –0.19 ns | –0.37 ns | –0.40* | –0.13 ns | 0.13 ns | –0.43* |
| Tmean HD-MD | 0.26 ns | –0.16 ns | –0.02 ns | 0.20 ns | –0.01 ns | –0.30 ns | 0.24 ns |
| Tmax Em-HD | –0.43* | –0.55* | –0.68** | –0.62** | –0.22 ns | 0.56* | –0.62** |
| Tmax HD-MD | 0.17 ns | –0.32 ns | –0.21 ns | 0.10 ns | –0.07 ns | –0.22 ns | –0.22 ns |
Environmental factors measured for the main growth phases, where Em, emergence; Jn, jointing; HD, heading date; MD, maturity date.
Measured from 1 week before heading to 3 weeks after heading.
*,**, Significant at P 0.05 and P 0.01 levels, respectively; ns, not significant.
Correlations between the environmental factors and the phenotypic traits (heading date, HD; plant height, PH; ear peduncle length, PdL; number of grains m−2, Kpsm; thousand kernel weight, TKW; test weight, TW; grain yield, GY) across environments (15 envs. for HD, Kpsm, TKW, and GY; 13 envs. for PH and PdL, and 11 envs. for TW)
| Environmental factorsa | HD | PH | PdL | Kpsm | TKW | TW | GY |
| Water input (WI) | |||||||
| Wi Em-HD | 0.46* | –0.11 ns | –0.15 ns | –0.18 ns | 0.11 ns | –0.06 ns | –0.12 ns |
| Wi HD-MD | –0.23 ns | 0.41* | 0.59* | 0.12 ns | 0.55* | 0.10 ns | 0.32 ns |
| Soil moisture (Sm) | |||||||
| Sm Jn-HD | 0.69** | 0.18 ns | 0.39 ns | 0.22 ns | 0.46* | 0.58* | 0.40* |
| Sm HDb | 0.45* | 0.26 ns | 0.46* | 0.49* | 0.36 ns | 0.13 ns | 0.57* |
| Sm Jn-MD | 0.43* | 0.20 ns | 0.43* | 0.14 ns | 0.38 ns | 0.54* | 0.28 ns |
| Sm HD-MD | –0.27 ns | 0.14 ns | 0.34 ns | –0.08 ns | 0.02 ns | –0.15 ns | –0.08 ns |
| Temperature (T) | |||||||
| Tmean Em-HD | –0.46* | –0.19 ns | –0.37 ns | –0.40* | –0.13 ns | 0.13 ns | –0.43* |
| Tmean HD-MD | 0.26 ns | –0.16 ns | –0.02 ns | 0.20 ns | –0.01 ns | –0.30 ns | 0.24 ns |
| Tmax Em-HD | –0.43* | –0.55* | –0.68** | –0.62** | –0.22 ns | 0.56* | –0.62** |
| Tmax HD-MD | 0.17 ns | –0.32 ns | –0.21 ns | 0.10 ns | –0.07 ns | –0.22 ns | –0.22 ns |
| Environmental factorsa | HD | PH | PdL | Kpsm | TKW | TW | GY |
| Water input (WI) | |||||||
| Wi Em-HD | 0.46* | –0.11 ns | –0.15 ns | –0.18 ns | 0.11 ns | –0.06 ns | –0.12 ns |
| Wi HD-MD | –0.23 ns | 0.41* | 0.59* | 0.12 ns | 0.55* | 0.10 ns | 0.32 ns |
| Soil moisture (Sm) | |||||||
| Sm Jn-HD | 0.69** | 0.18 ns | 0.39 ns | 0.22 ns | 0.46* | 0.58* | 0.40* |
| Sm HDb | 0.45* | 0.26 ns | 0.46* | 0.49* | 0.36 ns | 0.13 ns | 0.57* |
| Sm Jn-MD | 0.43* | 0.20 ns | 0.43* | 0.14 ns | 0.38 ns | 0.54* | 0.28 ns |
| Sm HD-MD | –0.27 ns | 0.14 ns | 0.34 ns | –0.08 ns | 0.02 ns | –0.15 ns | –0.08 ns |
| Temperature (T) | |||||||
| Tmean Em-HD | –0.46* | –0.19 ns | –0.37 ns | –0.40* | –0.13 ns | 0.13 ns | –0.43* |
| Tmean HD-MD | 0.26 ns | –0.16 ns | –0.02 ns | 0.20 ns | –0.01 ns | –0.30 ns | 0.24 ns |
| Tmax Em-HD | –0.43* | –0.55* | –0.68** | –0.62** | –0.22 ns | 0.56* | –0.62** |
| Tmax HD-MD | 0.17 ns | –0.32 ns | –0.21 ns | 0.10 ns | –0.07 ns | –0.22 ns | –0.22 ns |
Environmental factors measured for the main growth phases, where Em, emergence; Jn, jointing; HD, heading date; MD, maturity date.
Measured from 1 week before heading to 3 weeks after heading.
*,**, Significant at P 0.05 and P 0.01 levels, respectively; ns, not significant.
A similar observation pertains to the correlations between GY and the mean and maximum temperatures (T). Considering the thermic range, significant negative correlations across envs. were observed with HD, PH, PdL, Kpsm, and GY (significant r values ranging from –0.40 to –0.68), particularly in the period from emergence to heading and, more particularly, from jointing to heading. Interestingly, TW was positively associated with Tmax from emergence to heading. All these data highlight the importance of the thermo-pluviometrical conditions from jointing to heading on GY (particularly in terms of fertility and, most probably, also of carbon reserve accumulation in the vegetative organs), compared with the conditions in the grain-filling phase, where it is known that the terminal stress can significantly reduce TKW and TW.
Correlations between envs. for each of the measured traits are summarized in Table 3. A relatively high frequency of significant correlations between envs. was observed for PH, PdL, and TKW, with 83, 85, and 87% of environment pairs showing highly significant correlations (P 0.01), respectively. As to GY, highly significant correlations were observed in only 24% of environment pairs, a finding in line with the higher functional complexity and the low heritability of this trait. A large portion of the non-significant pairwise correlations was due to Spn2-r05, i.e. the most stressed and lowest-yielding env.
Summary of correlations between different environments for each trait.
| Traits | Correlations | |||||
| Total | Not significant | Significant at P 0.05 | Significant at P 0.01 | |||
| (no.) | (no.) | (no.) | Range of r values | (no.) | Range of r values | |
| Heading date | 105 | 26 (25%) | 7 (7%) | from –0.19 to –0.17 | 72 (69%) | from –0.23 to 0.80 |
| Plant height | 78 | 12 (15%) | 1 (1%) | 0.17 | 65 (83%) | from 0.20 to 0.98 |
| Peduncle length | 78 | 12 (15%) | – | – | 66 (85%) | from 0.28 to 0.75 |
| Grain yield | 105 | 65 (62%) | 22 (21%) | from –0.16 to 0.18 | 18 (17%) | from 0.19 to 0.43 |
| Kernels m−2 | 105 | 26 (25%) | 3 (3%) | from 0.14 to 0.17 | 76 (72%) | from 0.19 to 0.70 |
| Thousand kernel weight | 105 | 13 (12%) | 1 (1%) | 0.15 | 91 (87%) | from 0.22 to 0.72 |
| Test weight | 55 | 23 (42%) | 4 (7%) | from 0.15 to 0.18 | 28 (51%) | from 0.19 to 0.67 |
| Traits | Correlations | |||||
| Total | Not significant | Significant at P 0.05 | Significant at P 0.01 | |||
| (no.) | (no.) | (no.) | Range of r values | (no.) | Range of r values | |
| Heading date | 105 | 26 (25%) | 7 (7%) | from –0.19 to –0.17 | 72 (69%) | from –0.23 to 0.80 |
| Plant height | 78 | 12 (15%) | 1 (1%) | 0.17 | 65 (83%) | from 0.20 to 0.98 |
| Peduncle length | 78 | 12 (15%) | – | – | 66 (85%) | from 0.28 to 0.75 |
| Grain yield | 105 | 65 (62%) | 22 (21%) | from –0.16 to 0.18 | 18 (17%) | from 0.19 to 0.43 |
| Kernels m−2 | 105 | 26 (25%) | 3 (3%) | from 0.14 to 0.17 | 76 (72%) | from 0.19 to 0.70 |
| Thousand kernel weight | 105 | 13 (12%) | 1 (1%) | 0.15 | 91 (87%) | from 0.22 to 0.72 |
| Test weight | 55 | 23 (42%) | 4 (7%) | from 0.15 to 0.18 | 28 (51%) | from 0.19 to 0.67 |
Summary of correlations between different environments for each trait.
| Traits | Correlations | |||||
| Total | Not significant | Significant at P 0.05 | Significant at P 0.01 | |||
| (no.) | (no.) | (no.) | Range of r values | (no.) | Range of r values | |
| Heading date | 105 | 26 (25%) | 7 (7%) | from –0.19 to –0.17 | 72 (69%) | from –0.23 to 0.80 |
| Plant height | 78 | 12 (15%) | 1 (1%) | 0.17 | 65 (83%) | from 0.20 to 0.98 |
| Peduncle length | 78 | 12 (15%) | – | – | 66 (85%) | from 0.28 to 0.75 |
| Grain yield | 105 | 65 (62%) | 22 (21%) | from –0.16 to 0.18 | 18 (17%) | from 0.19 to 0.43 |
| Kernels m−2 | 105 | 26 (25%) | 3 (3%) | from 0.14 to 0.17 | 76 (72%) | from 0.19 to 0.70 |
| Thousand kernel weight | 105 | 13 (12%) | 1 (1%) | 0.15 | 91 (87%) | from 0.22 to 0.72 |
| Test weight | 55 | 23 (42%) | 4 (7%) | from 0.15 to 0.18 | 28 (51%) | from 0.19 to 0.67 |
| Traits | Correlations | |||||
| Total | Not significant | Significant at P 0.05 | Significant at P 0.01 | |||
| (no.) | (no.) | (no.) | Range of r values | (no.) | Range of r values | |
| Heading date | 105 | 26 (25%) | 7 (7%) | from –0.19 to –0.17 | 72 (69%) | from –0.23 to 0.80 |
| Plant height | 78 | 12 (15%) | 1 (1%) | 0.17 | 65 (83%) | from 0.20 to 0.98 |
| Peduncle length | 78 | 12 (15%) | – | – | 66 (85%) | from 0.28 to 0.75 |
| Grain yield | 105 | 65 (62%) | 22 (21%) | from –0.16 to 0.18 | 18 (17%) | from 0.19 to 0.43 |
| Kernels m−2 | 105 | 26 (25%) | 3 (3%) | from 0.14 to 0.17 | 76 (72%) | from 0.19 to 0.70 |
| Thousand kernel weight | 105 | 13 (12%) | 1 (1%) | 0.15 | 91 (87%) | from 0.22 to 0.72 |
| Test weight | 55 | 23 (42%) | 4 (7%) | from 0.15 to 0.18 | 28 (51%) | from 0.19 to 0.67 |
As shown by correlations between traits in each env. (see Supplementary Table S3 at JXB online), GY was significantly influenced by HD, with two positive correlations in high-yielding envs. (Lbn-i05 and Tns-i05) and six negative r values (varying from –0.40 to –0.20) across low-, medium- and high-yielding envs. These results underline the positive role of ‘escape’ in determining yield exclusively under water-stressed conditions. PH and PdL were positively associated with GY (r values from 0.24 to 0.51 for PH and from 0.15 to 0.33 for PdL) only in medium- and high-yielding envs. As to yield components, GY was positively influenced by Kpsm (fertility component) across most envs. (highly significant positive correlations in 13 envs., with r values from 0.19 to 0.98), while TKW was positively correlated with GY particularly in medium- and low-productive envs. In fact, significant positive correlations were observed in six out of eight medium- and low-yielding envs. (r values from 0.14 to 0.78), while only one out of seven high-yielding envs. showed a significant correlation. Considering the correlations among the other traits, noticeable associations were observed between HD and Kpsm (10 envs. showed a significant correlation, positive in five high-yielding envs. and negative under one high- and four medium- or low-yielding envs.). The correlation between HD and TKW was significant in four high-yielding envs. only, and always negative in sign. HD and PdL were negatively correlated in seven envs. across all yielding levels. PdL and TW showed a highly significant and consistently positive correlation in eight envs. (r values from 0.15 to 0.42) across a broad range of GY (from 35.8 up to 67.3 q ha−1).
Polymorphism at SSR loci and at Vrn-A1, Ppd-A1, and RhT-B1 genes
The banding patterns of the 191 SSR loci showed a total of 970 alleles across the 189 elite durum wheat accessions (see Supplementry Table S4 at JXB online). On average, 5.1±2.2 (from 2 to 13) alleles per locus were observed including 2.5±0.8 (from 1 to 4) non-rare alleles per locus and 2.2±1.8 (from 0 to 9) rare alleles per locus. In the final data set used for diversity and association analyses, null alleles and residual heterogeneity were present at low frequency (4.3% and 2.6%, respectively). Twelve loci with only one non-rare allele were not considered for the association analysis.
As to polymorphism at the Vrn-A1 (vernalization response) gene, no allelic variation was detected in the promoter region using the ASO (Allelic Specific Oligonucleotide) assay developed by Yan et al. (2004) and only limited allelic variation was observed in the first intron of the gene; in this case, the vernalization-insensitive dominant Vrn-A1c allele (characterized by a 7222 bp deletion, Fu et al. 2005) was largely prevalent in the IDuWUE panel (allele present in 183 out of the 189 accessions; see Supplementary Table S5 at JXB online). Similar to Vrn-A1, the IDuWUE panel showed unbalanced allele frequencies at Rht-B1 (semi-dwarfing gene) with the majority of genotypes carrying the Rht-B1b allele for semi-dwarf phenotype (162 accessions over 189). More informative variation was observed at Ppd-A1 (photoperiod-responsive gene), where the three main alleles already described by Wilhelm et al. (2009) were represented with rather balanced frequencies in the germplasm: 34% of the accessions were characterized by the photoperiod-sensitive allele (intact promoter sequence, wild-type), while 16% and 50% of the accessions showed the two photoperiod-insensitive promoter deletions of 1027 and 1117 bp, respectively.
Genetic structure and linkage disequilibrium analysis
The genetic relationships among the accessions were investigated using both a genetic-similarity and a model-based Bayesian clustering method. Both methods pointed out that the minimum and optimal number of hypothetical well-distinct subgroups present in the IDuWUE panel was equal to five. A higher number (>5) of Bayesian clustering-based subgroups improved the overall fit to the model but led to a generalized decrease of the number of accessions assigned to a specific subgroup with high membership probability (>0.50), suggesting excessive partitioning of the diversity structure. Each subgroup was characterized by a distinct genetic origin tracing back to foundation accessions which were included in the IDuWUE panel. Supplemental materials report the UPGMA-based dendrogram of the accessions (see Supplementary Fig. S2 at JXB online) and the accessions included in each of the five subgroups together with their corresponding membership probability estimates (see Supplementary Table S1 at JXB online).
The effect of population structure on phenotypic traits was investigated by means of regression analysis (Table 4). Using the mean values across envs., a modest effect was observed for HD (R2=4.0%) and GY (R2=7.3%); a greater influence of population structure was detected for PH, PdL, Kpsm, TKW, and TW, with R2 values ranging from 16.3% to 30.6%. Similar results were also obtained when considering data of each env. (data not reported).
Relationships between the phenotypic traits and population structure computed using the mean values across environments
| Traits | Environments | Phenotype–population structure regression results | |
| (no.) | R2 (%) | P-value | |
| Heading date | 15a | 4.0 | 0.107 |
| Plant height | 13b | 19.1 | 0.001 |
| Peduncle length | 13b | 30.6 | 0.001 |
| Grain yield | 15a | 7.3 | 0.007 |
| Kernels m−2 | 15a | 27.7 | 0.001 |
| Thousand kernel weight | 15a | 28.9 | 0.001 |
| Test weight | 11c | 16.3 | 0.001 |
| Traits | Environments | Phenotype–population structure regression results | |
| (no.) | R2 (%) | P-value | |
| Heading date | 15a | 4.0 | 0.107 |
| Plant height | 13b | 19.1 | 0.001 |
| Peduncle length | 13b | 30.6 | 0.001 |
| Grain yield | 15a | 7.3 | 0.007 |
| Kernels m−2 | 15a | 27.7 | 0.001 |
| Thousand kernel weight | 15a | 28.9 | 0.001 |
| Test weight | 11c | 16.3 | 0.001 |
R2 and probability values of the phenotype–population structure association are from the multiple regression analysis.
Trait measured in the 15 environments listed in Table 1.
Trait not measured in Mrc-r04 and Mrc-i04.
Trait not measured in Mrc-r04, Mrc-i04, Syr-r05, and Syr-i05.
Relationships between the phenotypic traits and population structure computed using the mean values across environments
| Traits | Environments | Phenotype–population structure regression results | |
| (no.) | R2 (%) | P-value | |
| Heading date | 15a | 4.0 | 0.107 |
| Plant height | 13b | 19.1 | 0.001 |
| Peduncle length | 13b | 30.6 | 0.001 |
| Grain yield | 15a | 7.3 | 0.007 |
| Kernels m−2 | 15a | 27.7 | 0.001 |
| Thousand kernel weight | 15a | 28.9 | 0.001 |
| Test weight | 11c | 16.3 | 0.001 |
| Traits | Environments | Phenotype–population structure regression results | |
| (no.) | R2 (%) | P-value | |
| Heading date | 15a | 4.0 | 0.107 |
| Plant height | 13b | 19.1 | 0.001 |
| Peduncle length | 13b | 30.6 | 0.001 |
| Grain yield | 15a | 7.3 | 0.007 |
| Kernels m−2 | 15a | 27.7 | 0.001 |
| Thousand kernel weight | 15a | 28.9 | 0.001 |
| Test weight | 11c | 16.3 | 0.001 |
R2 and probability values of the phenotype–population structure association are from the multiple regression analysis.
Trait measured in the 15 environments listed in Table 1.
Trait not measured in Mrc-r04 and Mrc-i04.
Trait not measured in Mrc-r04, Mrc-i04, Syr-r05, and Syr-i05.
The LD pattern was assessed based on the 15 931 pairwise combinations of 179 SSR loci. LD was initially investigated for the set of 15 066 pairs of non-syntenic loci (i.e. loci located on different chrs.). Based on permutation analysis, the portion of SSR pairs that showed a significant LD was equal to 27.8% at P 0.01 and 15.1% at P 0.001; this LD was thus largely due to the presence of a residual population structure in the IDuWUE panel. As expected, pairs of physically linked (syntenic) loci (865 pairs in total) showed LD estimates largely influenced by their genetic linkage. In fact, 84% of the 126 pairs of loci with inter-marker distance comprised within 10 cM showed significant LD at P 0.01, with an average r2 and D' values equal to 0.15±0.12 and 0.54±0.24, respectively. A significant LD was also observed in 61% of the 149 pairs of markers positioned within 10–20 cM. Figure 1 shows the scatterplot of the distributions of r2 and D' values as a function of the genetic distance between the syntenic loci pairs. Due to the presence of significant LD effects caused by population structure, ad hoc critical thresholds for the LD estimates between syntenic loci were obtained from the distribution of the non-syntenic loci. According to Breseghello and Sorrells (2006), the 95th percentile of the LD estimate distributions (r2 and D' values equal to 0.06 and 0.48, respectively; see Fig. 1) was used as critical threshold to discriminate better the LD most likely due to genetic linkage. Significant LD, based on the transection point of the LOESS-curve with the critical LD threshold for r2 and D' values, can be estimated to extend on average up to ca. 10 cM. From a more applicative standpoint, when using molecular markers the detection power is reduced to a certain extent (i.e. quantified as a proportion of r2) as compared to testing based on the causal gene, hence leading to set reference values for the presence of LD higher than those herein identified based on the LD estimate distributions (Farnir et al., 2000; Rafalski and Morgante, 2004; Nordborg et al., 2005; Rostoks et al., 2006; McKay et al., 2007; Ersoz et al., 2008).
Scatterplots of the LD statistics (r2 and D') of linked (syntenic) marker pairs as a function of the genetic distance (cM) between markers. The locally weighted polynomial regression-based (LOESS) fitting curve and the LD critical threshold as estimated from the LD statistic distribution of the pairs of unlinked markers have been reported in each scatterplot. Distances (cM) between markers were obtained by combining two intra-specific durum RIL-based linkage maps (Maccaferri et al., 2008; Mantovani et al., 2008). LD thresholds were superimposed on the scatterplots. Thresholds discriminating between the LD due to population structure and to true linkage have been calculated based on the 95th percentile of the r2 and D' distributions from the unlinked marker-pairs and were equal to 0.06 (r2) and 0.48 (D'). The commonly used thresholds indicating appreciable LD higher than 0.10 and 0.20 for r2 and 0.60 for D' are also reported.
Setting an r2 LD threshold of 0.10, 112 pairs of markers (12.9% of the syntenic pairs of loci) showed r2 values above this threshold, with an average intermarker distance of 13.3 cM. Figure 1 shows that this average distance is largely influenced by a few marker pairs with an intermarker distance >20 cM, while the majority of marker pairs with r2 above the threshold was characterized by an intermarker distance varying from 0 cM and 10 cM. In addition, it should be noted that the LD threshold (r2=0.10) intersects the LOESS-curve at an intermarker distance equal to zero cM. When considering an r2 threshold of 0.20, only 42 marker pairs (4.8%) with an average intermarker distance of 6.1 cM, showed r2 values higher than the threshold. Strong evidence of LD (r2 values >0.20 and D' values >0.60) was mostly observed for linked marker pairs with distances <10 cM.
Marker–trait association
Marker–phenotype association analysis was based on the polymorphisms present at the three genes Vrn-A1, Ppd-A1, and Rht-B1 and at 179 SSR loci, each characterized by the presence of at least two non-rare alleles, i.e. with frequency >0.10, for a total of 472 non-rare alleles.
The allelic frequencies of the five subgroups at three major genes for wheat phenology and plant architecture (Vrn-A1, Ppd-A1, and Rht-B1) are reported in Supplementary Table S5 at JXB online. The majority of the accessions were vernalization-insensitive and semi-dwarf, as expected for elite durum wheat materials. The vernalization-sensitive allele vrn-A1 was present in only six accessions, all but one (Claudio) from ICARDA germplasm (subgroups S1 and S2). Interestingly, in subgroup S1, bred for the semi-arid areas, the majority of genotypes carried the wild-type Rht-B1a allele conferring the tall phenotype. The distribution of the three alleles at Ppd-A1 was not uniform among subgroups: the wild-type, photoperiod sensitive, allele was the only variant observed in subgroup S1, while the 1117 bp deletion prevailed in subgroup S5. Subgroups S2, S3, and S4 showed non-rare frequencies for at least two of the three variants.
Due to the limited frequency of the allelic variant putatively associated with the winter habit, the Vrn-A1-phenotype association test was not carried out. However, a slight delay in heading date was associated with the accessions not carrying the intron deletion in the promoter region (data not reported). The GLM allele–phenotype association test showed a significant association between Ppd-A1 and HD in 12 out of 15 envs., with eight envs. also showing a significant association at the experiment-wise level (Table 5). In these eight envs., high R2 marker values were observed (from 6.6% up to 32.7%); further, population structure effects were limited (R2 model values similar to R2 marker values). In most cases, the HD mean values associated with the three allelic variants differed significantly, with the wild type being the latest allele (as expected, based on Wilhelm et al., 2009) and with the two alleles carrying either the 1117 or 1027 bp deletion in the promoter region being characterized by 2.2 d and 2.8 d earlier heading, respectively, than the wild type (average across the envs. where the association was significant). The difference in HD between the two alleles carrying the promoter deletions, although small, was significant in seven envs., with a consistent direction across all envs. It is interesting to note that, as in the case of Vrn-A1, the allele conferring lateness through photoperiod-sensitivity (wild type) in the Mediterranean envs. was highly represented in the genotypes directly tracing to the native germplasm pool from North Africa and West Asia (subgroup S1; see Supplementary Table S5 at JXB online) while the photoperiod-insensitive alleles dominated in most of the modern germplasm (especially subgoups S4 and S5). Though limited to a few envs., Ppd-A1 also significantly affected GY (Table 5) and other morpho-physiological traits such as PH, PdL, and Kpsm (results not reported).
Phenotypic effects of the allelic variants at the Ppd-A1 locus on heading date (HD, d from the seedling emergence) and grain yield (GY, q ha−1).
| Traits | Environmentsa | ||||||||||||||
| High-yielding | Medium-yielding | Low-yielding | |||||||||||||
| Tns-i05 | Itl1-r04 | Spn1-r04 | Itl2-r04 | Syr-i05 | Itl2-r05 | Lbn-i05 | Syr-r05 | Tns-r05 | Lbn-i04 | Lbn-r05 | Mrc-i04 | Mrc-r04 | Lbn-r04 | Spn2-r05 | |
| HD (d) | |||||||||||||||
| Ppd-A1 wild type b | 100.2 | 175.6 | 121.7 | 113.5 | 111.5 | 145.7 | 107.5 | 138.5 | 100.1 | 144.4 | 108.0 | 112.7 | 110.6 | 143.4 | 121.0 |
| Ppd-A1 1117-bp deletionc | 99.9 | 173.8 | 121.2 | 111.5 | 110.7 | 143.7 | 106.9 | 136.5 | 99.6 | 141.9 | 107.4 | 108.8 | 103.3 | 141.1 | 121.2 |
| Ppd-A1 1027-bp deletiond | 99.8 | 173.4 | 120.3 | 110.7 | 110.1 | 144.3 | 107.2 | 137.1 | 97.9 | 142.1 | 107.7 | 107.6 | 101.0 | 140.6 | 121.1 |
| LSD (P 0.05)e | – | 0.8 | 0.8 | 0.7 | 0.6 | 0.7 | 0.6 | 0.9 | 0.9 | 0.8 | – | 0.9 | 1.8 | 0.8 | – |
| R2 model (%) | – | 25.6 | 22.1 | 24.5 | 19.3 | 7.8 | 7.6 | 15.9 | 18.3 | 25.8 | – | 35.4 | 34.8 | 22.6 | – |
| R2 marker (%) | – | 11.1 | 3.8 | 21.2 | 8.4 | 3.8 | 3.5 | 6.6 | 3.4 | 15.9 | – | 32.7 | 32.7 | 18.6 | – |
| – | |||||||||||||||
| GY (q ha-1) | |||||||||||||||
| Ppd-A1 wild type | 68.1 | 67.2 | 64.2 | 58.0 | 58.3 | 57.2 | 55.9 | 44.2 | 44.5 | 42.1 | 37.1 | 30.3 | 26.2 | 21.5 | 8.7 |
| Ppd-A1 1117-bp deletion | 68.8 | 68.8 | 64.7 | 58.0 | 58.4 | 57.7 | 55.9 | 44.9 | 45.8 | 42.1 | 36.6 | 34.0 | 27.8 | 22.4 | 9.2 |
| Ppd-A1 1027-bp deletion | 68.8 | 68.7 | 66.1 | 58.1 | 56.8 | 57.2 | 56.0 | 44.6 | 45.3 | 43.2 | 36.5 | 32.9 | 26.9 | 22.5 | 9.1 |
| LSD (P 0.05) | – | – | – | – | – | – | – | – | – | 0.5 | – | 1.5 | 0.9 | – | – |
| R2 model (%) | – | – | – | – | – | – | – | – | – | 17.6 | – | 19.6 | 19.3 | – | – |
| R2 marker (%) | – | – | – | – | – | – | – | – | – | 8.2 | – | 1.5 | 5.2 | – | – |
| Traits | Environmentsa | ||||||||||||||
| High-yielding | Medium-yielding | Low-yielding | |||||||||||||
| Tns-i05 | Itl1-r04 | Spn1-r04 | Itl2-r04 | Syr-i05 | Itl2-r05 | Lbn-i05 | Syr-r05 | Tns-r05 | Lbn-i04 | Lbn-r05 | Mrc-i04 | Mrc-r04 | Lbn-r04 | Spn2-r05 | |
| HD (d) | |||||||||||||||
| Ppd-A1 wild type b | 100.2 | 175.6 | 121.7 | 113.5 | 111.5 | 145.7 | 107.5 | 138.5 | 100.1 | 144.4 | 108.0 | 112.7 | 110.6 | 143.4 | 121.0 |
| Ppd-A1 1117-bp deletionc | 99.9 | 173.8 | 121.2 | 111.5 | 110.7 | 143.7 | 106.9 | 136.5 | 99.6 | 141.9 | 107.4 | 108.8 | 103.3 | 141.1 | 121.2 |
| Ppd-A1 1027-bp deletiond | 99.8 | 173.4 | 120.3 | 110.7 | 110.1 | 144.3 | 107.2 | 137.1 | 97.9 | 142.1 | 107.7 | 107.6 | 101.0 | 140.6 | 121.1 |
| LSD (P 0.05)e | – | 0.8 | 0.8 | 0.7 | 0.6 | 0.7 | 0.6 | 0.9 | 0.9 | 0.8 | – | 0.9 | 1.8 | 0.8 | – |
| R2 model (%) | – | 25.6 | 22.1 | 24.5 | 19.3 | 7.8 | 7.6 | 15.9 | 18.3 | 25.8 | – | 35.4 | 34.8 | 22.6 | – |
| R2 marker (%) | – | 11.1 | 3.8 | 21.2 | 8.4 | 3.8 | 3.5 | 6.6 | 3.4 | 15.9 | – | 32.7 | 32.7 | 18.6 | – |
| – | |||||||||||||||
| GY (q ha-1) | |||||||||||||||
| Ppd-A1 wild type | 68.1 | 67.2 | 64.2 | 58.0 | 58.3 | 57.2 | 55.9 | 44.2 | 44.5 | 42.1 | 37.1 | 30.3 | 26.2 | 21.5 | 8.7 |
| Ppd-A1 1117-bp deletion | 68.8 | 68.8 | 64.7 | 58.0 | 58.4 | 57.7 | 55.9 | 44.9 | 45.8 | 42.1 | 36.6 | 34.0 | 27.8 | 22.4 | 9.2 |
| Ppd-A1 1027-bp deletion | 68.8 | 68.7 | 66.1 | 58.1 | 56.8 | 57.2 | 56.0 | 44.6 | 45.3 | 43.2 | 36.5 | 32.9 | 26.9 | 22.5 | 9.1 |
| LSD (P 0.05) | – | – | – | – | – | – | – | – | – | 0.5 | – | 1.5 | 0.9 | – | – |
| R2 model (%) | – | – | – | – | – | – | – | – | – | 17.6 | – | 19.6 | 19.3 | – | – |
| R2 marker (%) | – | – | – | – | – | – | – | – | – | 8.2 | – | 1.5 | 5.2 | – | – |
Environments are listed in decreasing order based on the mean GY values (see Table 1). Details on the environment codes are provided in Materials and Methods.
Reference accessions for the Ppd-A1 intact promoter sequence (wild type): Capeiti 8, Cappelli, Creso, Lahn, Omrabi 3 and Valnova.
Reference accessions for the Ppd-A1 1117-bp deletion: Gallareta, Karim, Korifla, Quadalete and Svevo.
Reference accessions for the Ppd-A1 1027-bp deletion: Cham 1 and Kofa.
LSD is reported only for environments with significant differences among alleles.
For each environment, not underlined, underlined and bold values refer to alleles not significantly different, significantly different at P 0.05 and P 0.01 marker-wise level, respectively; bold underlined values refer to alleles significantly different at P 0.05 experiment-wise level.
Phenotypic effects of the allelic variants at the Ppd-A1 locus on heading date (HD, d from the seedling emergence) and grain yield (GY, q ha−1).
| Traits | Environmentsa | ||||||||||||||
| High-yielding | Medium-yielding | Low-yielding | |||||||||||||
| Tns-i05 | Itl1-r04 | Spn1-r04 | Itl2-r04 | Syr-i05 | Itl2-r05 | Lbn-i05 | Syr-r05 | Tns-r05 | Lbn-i04 | Lbn-r05 | Mrc-i04 | Mrc-r04 | Lbn-r04 | Spn2-r05 | |
| HD (d) | |||||||||||||||
| Ppd-A1 wild type b | 100.2 | 175.6 | 121.7 | 113.5 | 111.5 | 145.7 | 107.5 | 138.5 | 100.1 | 144.4 | 108.0 | 112.7 | 110.6 | 143.4 | 121.0 |
| Ppd-A1 1117-bp deletionc | 99.9 | 173.8 | 121.2 | 111.5 | 110.7 | 143.7 | 106.9 | 136.5 | 99.6 | 141.9 | 107.4 | 108.8 | 103.3 | 141.1 | 121.2 |
| Ppd-A1 1027-bp deletiond | 99.8 | 173.4 | 120.3 | 110.7 | 110.1 | 144.3 | 107.2 | 137.1 | 97.9 | 142.1 | 107.7 | 107.6 | 101.0 | 140.6 | 121.1 |
| LSD (P 0.05)e | – | 0.8 | 0.8 | 0.7 | 0.6 | 0.7 | 0.6 | 0.9 | 0.9 | 0.8 | – | 0.9 | 1.8 | 0.8 | – |
| R2 model (%) | – | 25.6 | 22.1 | 24.5 | 19.3 | 7.8 | 7.6 | 15.9 | 18.3 | 25.8 | – | 35.4 | 34.8 | 22.6 | – |
| R2 marker (%) | – | 11.1 | 3.8 | 21.2 | 8.4 | 3.8 | 3.5 | 6.6 | 3.4 | 15.9 | – | 32.7 | 32.7 | 18.6 | – |
| – | |||||||||||||||
| GY (q ha-1) | |||||||||||||||
| Ppd-A1 wild type | 68.1 | 67.2 | 64.2 | 58.0 | 58.3 | 57.2 | 55.9 | 44.2 | 44.5 | 42.1 | 37.1 | 30.3 | 26.2 | 21.5 | 8.7 |
| Ppd-A1 1117-bp deletion | 68.8 | 68.8 | 64.7 | 58.0 | 58.4 | 57.7 | 55.9 | 44.9 | 45.8 | 42.1 | 36.6 | 34.0 | 27.8 | 22.4 | 9.2 |
| Ppd-A1 1027-bp deletion | 68.8 | 68.7 | 66.1 | 58.1 | 56.8 | 57.2 | 56.0 | 44.6 | 45.3 | 43.2 | 36.5 | 32.9 | 26.9 | 22.5 | 9.1 |
| LSD (P 0.05) | – | – | – | – | – | – | – | – | – | 0.5 | – | 1.5 | 0.9 | – | – |
| R2 model (%) | – | – | – | – | – | – | – | – | – | 17.6 | – | 19.6 | 19.3 | – | – |
| R2 marker (%) | – | – | – | – | – | – | – | – | – | 8.2 | – | 1.5 | 5.2 | – | – |
| Traits | Environmentsa | ||||||||||||||
| High-yielding | Medium-yielding | Low-yielding | |||||||||||||
| Tns-i05 | Itl1-r04 | Spn1-r04 | Itl2-r04 | Syr-i05 | Itl2-r05 | Lbn-i05 | Syr-r05 | Tns-r05 | Lbn-i04 | Lbn-r05 | Mrc-i04 | Mrc-r04 | Lbn-r04 | Spn2-r05 | |
| HD (d) | |||||||||||||||
| Ppd-A1 wild type b | 100.2 | 175.6 | 121.7 | 113.5 | 111.5 | 145.7 | 107.5 | 138.5 | 100.1 | 144.4 | 108.0 | 112.7 | 110.6 | 143.4 | 121.0 |
| Ppd-A1 1117-bp deletionc | 99.9 | 173.8 | 121.2 | 111.5 | 110.7 | 143.7 | 106.9 | 136.5 | 99.6 | 141.9 | 107.4 | 108.8 | 103.3 | 141.1 | 121.2 |
| Ppd-A1 1027-bp deletiond | 99.8 | 173.4 | 120.3 | 110.7 | 110.1 | 144.3 | 107.2 | 137.1 | 97.9 | 142.1 | 107.7 | 107.6 | 101.0 | 140.6 | 121.1 |
| LSD (P 0.05)e | – | 0.8 | 0.8 | 0.7 | 0.6 | 0.7 | 0.6 | 0.9 | 0.9 | 0.8 | – | 0.9 | 1.8 | 0.8 | – |
| R2 model (%) | – | 25.6 | 22.1 | 24.5 | 19.3 | 7.8 | 7.6 | 15.9 | 18.3 | 25.8 | – | 35.4 | 34.8 | 22.6 | – |
| R2 marker (%) | – | 11.1 | 3.8 | 21.2 | 8.4 | 3.8 | 3.5 | 6.6 | 3.4 | 15.9 | – | 32.7 | 32.7 | 18.6 | – |
| – | |||||||||||||||
| GY (q ha-1) | |||||||||||||||
| Ppd-A1 wild type | 68.1 | 67.2 | 64.2 | 58.0 | 58.3 | 57.2 | 55.9 | 44.2 | 44.5 | 42.1 | 37.1 | 30.3 | 26.2 | 21.5 | 8.7 |
| Ppd-A1 1117-bp deletion | 68.8 | 68.8 | 64.7 | 58.0 | 58.4 | 57.7 | 55.9 | 44.9 | 45.8 | 42.1 | 36.6 | 34.0 | 27.8 | 22.4 | 9.2 |
| Ppd-A1 1027-bp deletion | 68.8 | 68.7 | 66.1 | 58.1 | 56.8 | 57.2 | 56.0 | 44.6 | 45.3 | 43.2 | 36.5 | 32.9 | 26.9 | 22.5 | 9.1 |
| LSD (P 0.05) | – | – | – | – | – | – | – | – | – | 0.5 | – | 1.5 | 0.9 | – | – |
| R2 model (%) | – | – | – | – | – | – | – | – | – | 17.6 | – | 19.6 | 19.3 | – | – |
| R2 marker (%) | – | – | – | – | – | – | – | – | – | 8.2 | – | 1.5 | 5.2 | – | – |
Environments are listed in decreasing order based on the mean GY values (see Table 1). Details on the environment codes are provided in Materials and Methods.
Reference accessions for the Ppd-A1 intact promoter sequence (wild type): Capeiti 8, Cappelli, Creso, Lahn, Omrabi 3 and Valnova.
Reference accessions for the Ppd-A1 1117-bp deletion: Gallareta, Karim, Korifla, Quadalete and Svevo.
Reference accessions for the Ppd-A1 1027-bp deletion: Cham 1 and Kofa.
LSD is reported only for environments with significant differences among alleles.
For each environment, not underlined, underlined and bold values refer to alleles not significantly different, significantly different at P 0.05 and P 0.01 marker-wise level, respectively; bold underlined values refer to alleles significantly different at P 0.05 experiment-wise level.
As to the average number of significant marker–phenotype associations per env. detected with the 182 loci considered here (Vrn-A1, Ppd-A1, Rht-B1, and 179 SSRs), such a number was high for TKW (18.7 markers env.−1), Kpsm (16.5), and HD (16.4), intermediate for PdL (12.4) and PH (10.7), and low for TW (9.6) and GY (9.6). Moreover, the number of significant associations per env. was stable across stress intensities as per GY levels (high-, medium-, and low-yielding envs.) for HD only (15.6, 17.6, and 16.3 significant associations per env., respectively) but not for the other traits. As an example, the average number of associations for PH was 15.3, 6.8, and 6.3 in the high-, medium- and low-yielding envs., respectively; a similar trend was also shown by PdL. Similar results pertain to yield components: for Kpsm, the average number of significant associations per env. was equal to 19.2, 16.0, and 11.0 in the high-, medium-, and low- yielding envs., respectively, while for TKW it dropped from 20.4 in the high-yielding envs. to 12.7 in the low-yielding ones. For GY, the number of significant associations per env. was also quite low (10.3) in the high-yielding envs. and slightly decreased in the low-yielding envs. (7.0).
A summary of the number of SSR loci that showed a significant association with the target traits is reported in Table 6. Considering the associations that were significant across multiple envs., it can be noted that (i) the total number of markers showing significant trait associations in two or more envs. was higher for HD, Kpsm, and TKW (61, 65, and 60 markers, respectively) than for PH and TW (37 and 45 markers, respectively) and (ii) the markers with significant and consistent effects in more than half of the envs. (from 8 to 13 envs.) were found in higher number for HD (seven markers), TKW (eight markers), and PdL (five markers), compared with TW (one marker) and, particularly, GY (no marker). In fact, marker–trait associations for GY were usually detected in one env. only (56 markers), while decreasing rapidly from two up to five envs. (from 20 to three markers, respectively) and with only one marker (Xbarc197 on chr. 5A) showing association to GY across a total of six envs. (ranging from low- to high-yielding). These results are not unexpected owing to the complexity of the genetic basis of GY and its high level of interaction with environmental conditions.
Summary of the significant marker-trait associations listed by trait and according to the number of environments in which the association was significant
| Trait | Number of environments | |||||||||||||
| a | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | |
| Heading date | 15 | 46 | 17 | 11 | 8 | 12 | 3 | 3 | 5 | 1 | – | – | 1 | – |
| Plant height | 13 | 39 | 7 | 9 | 4 | 7 | 3 | 3 | 3 | 1 | – | – | – | – |
| Peduncle length | 13 | 52 | 12 | 10 | 13 | 4 | 3 | 1 | 1 | 1 | 2 | 1 | – | – |
| Kernels per square meter | 15 | 40 | 15 | 19 | 16 | 4 | 4 | 3 | 3 | – | – | 1 | – | – |
| Thousand kernel weight | 15 | 30 | 13 | 11 | 10 | 8 | 6 | 4 | 2 | – | 5 | – | – | 1 |
| Test weight | 11 | 54 | 13 | 16 | 12 | 2 | 1 | – | 1 | – | – | – | – | – |
| Grain yield | 15 | 56 | 20 | 17 | 9 | 3 | 1 | – | – | – | – | – | – | – |
| Trait | Number of environments | |||||||||||||
| a | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | |
| Heading date | 15 | 46 | 17 | 11 | 8 | 12 | 3 | 3 | 5 | 1 | – | – | 1 | – |
| Plant height | 13 | 39 | 7 | 9 | 4 | 7 | 3 | 3 | 3 | 1 | – | – | – | – |
| Peduncle length | 13 | 52 | 12 | 10 | 13 | 4 | 3 | 1 | 1 | 1 | 2 | 1 | – | – |
| Kernels per square meter | 15 | 40 | 15 | 19 | 16 | 4 | 4 | 3 | 3 | – | – | 1 | – | – |
| Thousand kernel weight | 15 | 30 | 13 | 11 | 10 | 8 | 6 | 4 | 2 | – | 5 | – | – | 1 |
| Test weight | 11 | 54 | 13 | 16 | 12 | 2 | 1 | – | 1 | – | – | – | – | – |
| Grain yield | 15 | 56 | 20 | 17 | 9 | 3 | 1 | – | – | – | – | – | – | – |
The results are referred to significant marker–trait associations on the basis of consistent marker-wise tests (P 0.05) carried out with GLM, MLMLois and MLMSM.
Number of environments in which the trait was measured.
Summary of the significant marker-trait associations listed by trait and according to the number of environments in which the association was significant
| Trait | Number of environments | |||||||||||||
| a | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | |
| Heading date | 15 | 46 | 17 | 11 | 8 | 12 | 3 | 3 | 5 | 1 | – | – | 1 | – |
| Plant height | 13 | 39 | 7 | 9 | 4 | 7 | 3 | 3 | 3 | 1 | – | – | – | – |
| Peduncle length | 13 | 52 | 12 | 10 | 13 | 4 | 3 | 1 | 1 | 1 | 2 | 1 | – | – |
| Kernels per square meter | 15 | 40 | 15 | 19 | 16 | 4 | 4 | 3 | 3 | – | – | 1 | – | – |
| Thousand kernel weight | 15 | 30 | 13 | 11 | 10 | 8 | 6 | 4 | 2 | – | 5 | – | – | 1 |
| Test weight | 11 | 54 | 13 | 16 | 12 | 2 | 1 | – | 1 | – | – | – | – | – |
| Grain yield | 15 | 56 | 20 | 17 | 9 | 3 | 1 | – | – | – | – | – | – | – |
| Trait | Number of environments | |||||||||||||
| a | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | |
| Heading date | 15 | 46 | 17 | 11 | 8 | 12 | 3 | 3 | 5 | 1 | – | – | 1 | – |
| Plant height | 13 | 39 | 7 | 9 | 4 | 7 | 3 | 3 | 3 | 1 | – | – | – | – |
| Peduncle length | 13 | 52 | 12 | 10 | 13 | 4 | 3 | 1 | 1 | 1 | 2 | 1 | – | – |
| Kernels per square meter | 15 | 40 | 15 | 19 | 16 | 4 | 4 | 3 | 3 | – | – | 1 | – | – |
| Thousand kernel weight | 15 | 30 | 13 | 11 | 10 | 8 | 6 | 4 | 2 | – | 5 | – | – | 1 |
| Test weight | 11 | 54 | 13 | 16 | 12 | 2 | 1 | – | 1 | – | – | – | – | – |
| Grain yield | 15 | 56 | 20 | 17 | 9 | 3 | 1 | – | – | – | – | – | – | – |
The results are referred to significant marker–trait associations on the basis of consistent marker-wise tests (P 0.05) carried out with GLM, MLMLois and MLMSM.
Number of environments in which the trait was measured.
Figure 2 graphically reports the markers with significant marker-wise associations in at least two envs. In particular, Fig. 2 reports the location on the reference map of the markers with significant effects, together with the traits involved and the corresponding significance level for (i) each of the 15 envs., (ii) the mean values of the three groups of envs. based on GY (i.e. high-, medium-, and low-yielding envs.), and (iii) the mean values across all the 15 envs.
Detailed marker–trait association results for markers with significant marker-wise associations in at least two or more envs. The chromosome locations of the significant marker–trait associations have been reported on a map derived by combining two intra-specific durum linkage maps obtained from recombinant inbred populations (Maccaferri et al., 2008; Mantovani et al., 2008). Markers which have been used to profile the durum collection and to perform the marker–trait association tests are indicated in bold. The association results are reported for each of the 15 environments using their corresponding codes (see Materials and methods) and for the mean values of the three groups of environments based on their productivity level (high-, medium-, and low-yielding, respectively) and for the mean values across all experiments. Acronyms of the traits significantly associated have been reported to the right of the corresponding markers. White- and grey-filled circles indicate marker-wise significant and highly significant (P 0.05 and P 0.01, respectively) associations. Black-filled circles represent experiment-wise significant associations (Bonferroni's correction P 0.05). The marker–trait associations that showed consistent results across traits and/or envs. and that were further confirmed on the mean values of groups of envs. with similar GY have been highlighted in grey.
Features of marker–trait associations that were consistent across traits and/or envs. (highlighted in Fig. 2) and further confirmed by the mean values of the three groups of envs. have been reported in Table 7. Moreover, the corresponding allelic variant means are reported in Supplementary Table S6 at JXB online. Two to four non-rare allelic variants per SSR locus were informative for association analysis (Table 7). As expected with quantitative traits, R2 values of the significant marker–trait associations were rather low (<10%), while model R2 values were comprised between 4.1% and 44.0% depending on the marker, trait, and environment.
Detailed results for the most valuable marker-trait associations: chromosome (chr.), position (cM) on the linkage group and number of non-rare alleles
| Marker | Chr. position | Non-rare alleles | Trait | R2 range | |
| (cM) | (no.) | Model | Marker | ||
| Xwmc469 | 1AS 51 | 2 | TKW | 14.2–38.0 | 2.1–7.4 |
| TW | 13.2–32.6 | 2.0–4.6 | |||
| Xbarc213 | 1AL 106 | 2 | PdL | 4.1–17.5 | 2.5–3.6 |
| Kpsm | 12.3–42.1 | 1.7–4.1 | |||
| TKW | 18.6–37.6 | 2.0–6.8 | |||
| Xbarc287 | 1AL 126 | 2 | Kpsm | 9.4–25.3 | 1.9–5.9 |
| TKW | 9.6–29.4 | 2.1–3.9 | |||
| Xcfd65 | 1BS 36 | 2 | PdL | 19.7–43.1 | 1.9–4.3 |
| Kpsm | 20.8–22.9 | 1.9–4.8 | |||
| TKW | 22.0–23.6 | 2.4–2.8 | |||
| TW | 14.0–25.3 | 2.7–3.8 | |||
| Xcfa2086 | 1BL 8 | 2 | HD | 5.0–20.6 | 2.2–3.5 |
| PdL | 11.8–26.9 | 2.3–3.2 | |||
| GY | 14.1–15.5 | 4.9–5.6 | |||
| Xwmc419 | 1BL 59 | 3 | PH | 4.6–36.6 | 2.0–8.4 |
| PdL | 9.4–44.1 | 1.7–7.8 | |||
| Xgwm294 | 2AL 118 | 3 | HD | 6.5–21.0 | 3.9–4.5 |
| PH | 9.8–38.8 | 3.9–6.2 | |||
| PdL | 17.2–44.0 | 3.2–5.6 | |||
| Kpsm | 21.3–33.7 | 2.7–7.5 | |||
| TKW | 25.0–30.1 | 3.7–8.1 | |||
| TW | 7.8–25.6 | 2.8–3.3 | |||
| Xgwm410.1 | 2BS 56 | 3 | TKW | 15.2–26.7 | 3.2–5.9 |
| TW | 7.6–17.9 | 3.4–4.5 | |||
| GY | 14.5–19.2 | 4.3–5.9 | |||
| Xgwm47.1 | 2BL 117 | 3 | PH | 5.0–30.5 | 2.2–5.9 |
| PdL | 8.9–34.6 | 1.4–4.7 | |||
| Kpsm | 6.1–35.6 | 1.9–5.0 | |||
| TKW | 6.3–30.6 | 1.4–5.3 | |||
| Xcfa2234 | 3AS 52 | 2 | Kpsm | 16.7–34.6 | 2.3–3.0 |
| TKW | 7.1–30.4 | 1.7–3.9 | |||
| Xgwm493 | 3BS 11 | 2 | HD | 8.5–16.7 | 2.1–5.9 |
| PH | 7.3–33.4 | 1.7–3.1 | |||
| PdL | 12.6–37.6 | 1.9–4.0 | |||
| TW | 9.1–4.8 | 3.4–2.3 | |||
| GY | 12.2–9.4 | 2.1–3.4 | |||
| Xbarc193 | 4BS 0 | 3 | PH | 7.8–36.4 | 2.8–6.8 |
| PdL | 14.6–34.8 | 2.3–5.2 | |||
| Xgwm1278 | 4BS 13 | 2 | PH | 8.0–34.7 | 1.8–3.2 |
| PdL | 25.5–38.3 | 1.8–3.1 | |||
| Kpsm | 16.7–36.7 | 1.4–2.2 | |||
| TKW | 12.3–32.9 | 1.9–4.2 | |||
| GY | 7.4–15.6 | 1.2–3.0 | |||
| Xbarc303 | 5AS 21 | 2 | HD | 6.2–12.5 | 2.9–5.7 |
| PdL | 20.2–38.5 | 1.8–2.1 | |||
| Kpsm | 5.9–39.1 | 2.0–5.9 | |||
| TKW | 16.0–35.6 | 0.9–5.0 | |||
| TW | 12.9–28.7 | 2.2–3.2 | |||
| Xbarc197 | 5AL 73 | 2 | HD | 6.8–23.7 | 2.1–4.3 |
| Kpsm | 18.4–21.1 | 2.3–4.3 | |||
| GY | 9.5–20.3 | 2.5–4.2 | |||
| Xgwm1570 | 5ALa 15 | 2 | PdL | 8.7–34.4 | 2.0–3.6 |
| Kpsm | 22.6–38.7 | 1.8–5.5 | |||
| TKW | 9.7–33.8 | 2.2–6.9 | |||
| Xwmc256 | 6AL 66 | 2 | HD | 5.2– 22.5 | 2.2–4.4 |
| GY | 8.1–23.8 | 1.5–4.7 | |||
| Xdupw217 | 6BS 0 | 3 | PH | 6.6–32.7 | 1.3–5.6 |
| Kpsm | 6.1–40.0 | 1.5–4.6 | |||
| TKW | 6.7–36.6 | 1.6–5.2 | |||
| GY | 3.0–15.7 | 1.8–4.2 | |||
| Xwmc494 | 6BS 43 | 3 | TW | 3.6–28.2 | 1.5–7.4 |
| Xbarc14 | 6BS 57 | 2 | PH | 3.0–18.9 | 2.3–4.4 |
| Kpsm | 2.7–6.4 | 1.8–2.7 | |||
| TW | 19.6–20.1 | 2.8–3.1 | |||
| GY | 3.7–15.6 | 2.1–3.9 | |||
| Xcfa2257.1 | 7AL 145 | 2 | Kpsm | 18.4–37.4 | 1.5–3.6 |
| TKW | 10.8–33.6 | 1.9–4.6 | |||
| GY | 19.3–20.1 | 2.8–3.1 | |||
| Xgwm333 | 7BL 87 | 3 | HD | 6.0–24.0 | 1.5–3.1 |
| Kpsm | 5.1–38.0 | 1.8–6.1 | |||
| TKW | 10.3–34.3 | 1.8–3.6 | |||
| TW | 4.3–23.5 | 1.2–3.4 | |||
| Xgwm577 | 7BLa 0 | 3 | HD | 5.6–28.5 | 2.2–10.0 |
| PdL | 8.4–35.6 | 1.6–3.8 | |||
| TKW | 8.0–30.5 | 1.4–4.0 | |||
| Xwmc276 | 7BLa 5 | 4 | PH | 5.6–33.2 | 2.0–7.7 |
| PdL | 11.0–42.4 | 2.6–7.1 | |||
| Kpsm | 4.5–33.9 | 1.8–3.8 | |||
| TW | 3.7–23.6 | 1.1–5.6 | |||
| GY | 6.9–16.5 | 2.0–4.8 | |||
| Marker | Chr. position | Non-rare alleles | Trait | R2 range | |
| (cM) | (no.) | Model | Marker | ||
| Xwmc469 | 1AS 51 | 2 | TKW | 14.2–38.0 | 2.1–7.4 |
| TW | 13.2–32.6 | 2.0–4.6 | |||
| Xbarc213 | 1AL 106 | 2 | PdL | 4.1–17.5 | 2.5–3.6 |
| Kpsm | 12.3–42.1 | 1.7–4.1 | |||
| TKW | 18.6–37.6 | 2.0–6.8 | |||
| Xbarc287 | 1AL 126 | 2 | Kpsm | 9.4–25.3 | 1.9–5.9 |
| TKW | 9.6–29.4 | 2.1–3.9 | |||
| Xcfd65 | 1BS 36 | 2 | PdL | 19.7–43.1 | 1.9–4.3 |
| Kpsm | 20.8–22.9 | 1.9–4.8 | |||
| TKW | 22.0–23.6 | 2.4–2.8 | |||
| TW | 14.0–25.3 | 2.7–3.8 | |||
| Xcfa2086 | 1BL 8 | 2 | HD | 5.0–20.6 | 2.2–3.5 |
| PdL | 11.8–26.9 | 2.3–3.2 | |||
| GY | 14.1–15.5 | 4.9–5.6 | |||
| Xwmc419 | 1BL 59 | 3 | PH | 4.6–36.6 | 2.0–8.4 |
| PdL | 9.4–44.1 | 1.7–7.8 | |||
| Xgwm294 | 2AL 118 | 3 | HD | 6.5–21.0 | 3.9–4.5 |
| PH | 9.8–38.8 | 3.9–6.2 | |||
| PdL | 17.2–44.0 | 3.2–5.6 | |||
| Kpsm | 21.3–33.7 | 2.7–7.5 | |||
| TKW | 25.0–30.1 | 3.7–8.1 | |||
| TW | 7.8–25.6 | 2.8–3.3 | |||
| Xgwm410.1 | 2BS 56 | 3 | TKW | 15.2–26.7 | 3.2–5.9 |
| TW | 7.6–17.9 | 3.4–4.5 | |||
| GY | 14.5–19.2 | 4.3–5.9 | |||
| Xgwm47.1 | 2BL 117 | 3 | PH | 5.0–30.5 | 2.2–5.9 |
| PdL | 8.9–34.6 | 1.4–4.7 | |||
| Kpsm | 6.1–35.6 | 1.9–5.0 | |||
| TKW | 6.3–30.6 | 1.4–5.3 | |||
| Xcfa2234 | 3AS 52 | 2 | Kpsm | 16.7–34.6 | 2.3–3.0 |
| TKW | 7.1–30.4 | 1.7–3.9 | |||
| Xgwm493 | 3BS 11 | 2 | HD | 8.5–16.7 | 2.1–5.9 |
| PH | 7.3–33.4 | 1.7–3.1 | |||
| PdL | 12.6–37.6 | 1.9–4.0 | |||
| TW | 9.1–4.8 | 3.4–2.3 | |||
| GY | 12.2–9.4 | 2.1–3.4 | |||
| Xbarc193 | 4BS 0 | 3 | PH | 7.8–36.4 | 2.8–6.8 |
| PdL | 14.6–34.8 | 2.3–5.2 | |||
| Xgwm1278 | 4BS 13 | 2 | PH | 8.0–34.7 | 1.8–3.2 |
| PdL | 25.5–38.3 | 1.8–3.1 | |||
| Kpsm | 16.7–36.7 | 1.4–2.2 | |||
| TKW | 12.3–32.9 | 1.9–4.2 | |||
| GY | 7.4–15.6 | 1.2–3.0 | |||
| Xbarc303 | 5AS 21 | 2 | HD | 6.2–12.5 | 2.9–5.7 |
| PdL | 20.2–38.5 | 1.8–2.1 | |||
| Kpsm | 5.9–39.1 | 2.0–5.9 | |||
| TKW | 16.0–35.6 | 0.9–5.0 | |||
| TW | 12.9–28.7 | 2.2–3.2 | |||
| Xbarc197 | 5AL 73 | 2 | HD | 6.8–23.7 | 2.1–4.3 |
| Kpsm | 18.4–21.1 | 2.3–4.3 | |||
| GY | 9.5–20.3 | 2.5–4.2 | |||
| Xgwm1570 | 5ALa 15 | 2 | PdL | 8.7–34.4 | 2.0–3.6 |
| Kpsm | 22.6–38.7 | 1.8–5.5 | |||
| TKW | 9.7–33.8 | 2.2–6.9 | |||
| Xwmc256 | 6AL 66 | 2 | HD | 5.2– 22.5 | 2.2–4.4 |
| GY | 8.1–23.8 | 1.5–4.7 | |||
| Xdupw217 | 6BS 0 | 3 | PH | 6.6–32.7 | 1.3–5.6 |
| Kpsm | 6.1–40.0 | 1.5–4.6 | |||
| TKW | 6.7–36.6 | 1.6–5.2 | |||
| GY | 3.0–15.7 | 1.8–4.2 | |||
| Xwmc494 | 6BS 43 | 3 | TW | 3.6–28.2 | 1.5–7.4 |
| Xbarc14 | 6BS 57 | 2 | PH | 3.0–18.9 | 2.3–4.4 |
| Kpsm | 2.7–6.4 | 1.8–2.7 | |||
| TW | 19.6–20.1 | 2.8–3.1 | |||
| GY | 3.7–15.6 | 2.1–3.9 | |||
| Xcfa2257.1 | 7AL 145 | 2 | Kpsm | 18.4–37.4 | 1.5–3.6 |
| TKW | 10.8–33.6 | 1.9–4.6 | |||
| GY | 19.3–20.1 | 2.8–3.1 | |||
| Xgwm333 | 7BL 87 | 3 | HD | 6.0–24.0 | 1.5–3.1 |
| Kpsm | 5.1–38.0 | 1.8–6.1 | |||
| TKW | 10.3–34.3 | 1.8–3.6 | |||
| TW | 4.3–23.5 | 1.2–3.4 | |||
| Xgwm577 | 7BLa 0 | 3 | HD | 5.6–28.5 | 2.2–10.0 |
| PdL | 8.4–35.6 | 1.6–3.8 | |||
| TKW | 8.0–30.5 | 1.4–4.0 | |||
| Xwmc276 | 7BLa 5 | 4 | PH | 5.6–33.2 | 2.0–7.7 |
| PdL | 11.0–42.4 | 2.6–7.1 | |||
| Kpsm | 4.5–33.9 | 1.8–3.8 | |||
| TW | 3.7–23.6 | 1.1–5.6 | |||
| GY | 6.9–16.5 | 2.0–4.8 | |||
R2 ranges of the model (marker+Q matrix as covariate) and of the marker only as from the GLM analysis are also reported.
Second linkage group of the chromosome.
Detailed results for the most valuable marker-trait associations: chromosome (chr.), position (cM) on the linkage group and number of non-rare alleles
| Marker | Chr. position | Non-rare alleles | Trait | R2 range | |
| (cM) | (no.) | Model | Marker | ||
| Xwmc469 | 1AS 51 | 2 | TKW | 14.2–38.0 | 2.1–7.4 |
| TW | 13.2–32.6 | 2.0–4.6 | |||
| Xbarc213 | 1AL 106 | 2 | PdL | 4.1–17.5 | 2.5–3.6 |
| Kpsm | 12.3–42.1 | 1.7–4.1 | |||
| TKW | 18.6–37.6 | 2.0–6.8 | |||
| Xbarc287 | 1AL 126 | 2 | Kpsm | 9.4–25.3 | 1.9–5.9 |
| TKW | 9.6–29.4 | 2.1–3.9 | |||
| Xcfd65 | 1BS 36 | 2 | PdL | 19.7–43.1 | 1.9–4.3 |
| Kpsm | 20.8–22.9 | 1.9–4.8 | |||
| TKW | 22.0–23.6 | 2.4–2.8 | |||
| TW | 14.0–25.3 | 2.7–3.8 | |||
| Xcfa2086 | 1BL 8 | 2 | HD | 5.0–20.6 | 2.2–3.5 |
| PdL | 11.8–26.9 | 2.3–3.2 | |||
| GY | 14.1–15.5 | 4.9–5.6 | |||
| Xwmc419 | 1BL 59 | 3 | PH | 4.6–36.6 | 2.0–8.4 |
| PdL | 9.4–44.1 | 1.7–7.8 | |||
| Xgwm294 | 2AL 118 | 3 | HD | 6.5–21.0 | 3.9–4.5 |
| PH | 9.8–38.8 | 3.9–6.2 | |||
| PdL | 17.2–44.0 | 3.2–5.6 | |||
| Kpsm | 21.3–33.7 | 2.7–7.5 | |||
| TKW | 25.0–30.1 | 3.7–8.1 | |||
| TW | 7.8–25.6 | 2.8–3.3 | |||
| Xgwm410.1 | 2BS 56 | 3 | TKW | 15.2–26.7 | 3.2–5.9 |
| TW | 7.6–17.9 | 3.4–4.5 | |||
| GY | 14.5–19.2 | 4.3–5.9 | |||
| Xgwm47.1 | 2BL 117 | 3 | PH | 5.0–30.5 | 2.2–5.9 |
| PdL | 8.9–34.6 | 1.4–4.7 | |||
| Kpsm | 6.1–35.6 | 1.9–5.0 | |||
| TKW | 6.3–30.6 | 1.4–5.3 | |||
| Xcfa2234 | 3AS 52 | 2 | Kpsm | 16.7–34.6 | 2.3–3.0 |
| TKW | 7.1–30.4 | 1.7–3.9 | |||
| Xgwm493 | 3BS 11 | 2 | HD | 8.5–16.7 | 2.1–5.9 |
| PH | 7.3–33.4 | 1.7–3.1 | |||
| PdL | 12.6–37.6 | 1.9–4.0 | |||
| TW | 9.1–4.8 | 3.4–2.3 | |||
| GY | 12.2–9.4 | 2.1–3.4 | |||
| Xbarc193 | 4BS 0 | 3 | PH | 7.8–36.4 | 2.8–6.8 |
| PdL | 14.6–34.8 | 2.3–5.2 | |||
| Xgwm1278 | 4BS 13 | 2 | PH | 8.0–34.7 | 1.8–3.2 |
| PdL | 25.5–38.3 | 1.8–3.1 | |||
| Kpsm | 16.7–36.7 | 1.4–2.2 | |||
| TKW | 12.3–32.9 | 1.9–4.2 | |||
| GY | 7.4–15.6 | 1.2–3.0 | |||
| Xbarc303 | 5AS 21 | 2 | HD | 6.2–12.5 | 2.9–5.7 |
| PdL | 20.2–38.5 | 1.8–2.1 | |||
| Kpsm | 5.9–39.1 | 2.0–5.9 | |||
| TKW | 16.0–35.6 | 0.9–5.0 | |||
| TW | 12.9–28.7 | 2.2–3.2 | |||
| Xbarc197 | 5AL 73 | 2 | HD | 6.8–23.7 | 2.1–4.3 |
| Kpsm | 18.4–21.1 | 2.3–4.3 | |||
| GY | 9.5–20.3 | 2.5–4.2 | |||
| Xgwm1570 | 5ALa 15 | 2 | PdL | 8.7–34.4 | 2.0–3.6 |
| Kpsm | 22.6–38.7 | 1.8–5.5 | |||
| TKW | 9.7–33.8 | 2.2–6.9 | |||
| Xwmc256 | 6AL 66 | 2 | HD | 5.2– 22.5 | 2.2–4.4 |
| GY | 8.1–23.8 | 1.5–4.7 | |||
| Xdupw217 | 6BS 0 | 3 | PH | 6.6–32.7 | 1.3–5.6 |
| Kpsm | 6.1–40.0 | 1.5–4.6 | |||
| TKW | 6.7–36.6 | 1.6–5.2 | |||
| GY | 3.0–15.7 | 1.8–4.2 | |||
| Xwmc494 | 6BS 43 | 3 | TW | 3.6–28.2 | 1.5–7.4 |
| Xbarc14 | 6BS 57 | 2 | PH | 3.0–18.9 | 2.3–4.4 |
| Kpsm | 2.7–6.4 | 1.8–2.7 | |||
| TW | 19.6–20.1 | 2.8–3.1 | |||
| GY | 3.7–15.6 | 2.1–3.9 | |||
| Xcfa2257.1 | 7AL 145 | 2 | Kpsm | 18.4–37.4 | 1.5–3.6 |
| TKW | 10.8–33.6 | 1.9–4.6 | |||
| GY | 19.3–20.1 | 2.8–3.1 | |||
| Xgwm333 | 7BL 87 | 3 | HD | 6.0–24.0 | 1.5–3.1 |
| Kpsm | 5.1–38.0 | 1.8–6.1 | |||
| TKW | 10.3–34.3 | 1.8–3.6 | |||
| TW | 4.3–23.5 | 1.2–3.4 | |||
| Xgwm577 | 7BLa 0 | 3 | HD | 5.6–28.5 | 2.2–10.0 |
| PdL | 8.4–35.6 | 1.6–3.8 | |||
| TKW | 8.0–30.5 | 1.4–4.0 | |||
| Xwmc276 | 7BLa 5 | 4 | PH | 5.6–33.2 | 2.0–7.7 |
| PdL | 11.0–42.4 | 2.6–7.1 | |||
| Kpsm | 4.5–33.9 | 1.8–3.8 | |||
| TW | 3.7–23.6 | 1.1–5.6 | |||
| GY | 6.9–16.5 | 2.0–4.8 | |||
| Marker | Chr. position | Non-rare alleles | Trait | R2 range | |
| (cM) | (no.) | Model | Marker | ||
| Xwmc469 | 1AS 51 | 2 | TKW | 14.2–38.0 | 2.1–7.4 |
| TW | 13.2–32.6 | 2.0–4.6 | |||
| Xbarc213 | 1AL 106 | 2 | PdL | 4.1–17.5 | 2.5–3.6 |
| Kpsm | 12.3–42.1 | 1.7–4.1 | |||
| TKW | 18.6–37.6 | 2.0–6.8 | |||
| Xbarc287 | 1AL 126 | 2 | Kpsm | 9.4–25.3 | 1.9–5.9 |
| TKW | 9.6–29.4 | 2.1–3.9 | |||
| Xcfd65 | 1BS 36 | 2 | PdL | 19.7–43.1 | 1.9–4.3 |
| Kpsm | 20.8–22.9 | 1.9–4.8 | |||
| TKW | 22.0–23.6 | 2.4–2.8 | |||
| TW | 14.0–25.3 | 2.7–3.8 | |||
| Xcfa2086 | 1BL 8 | 2 | HD | 5.0–20.6 | 2.2–3.5 |
| PdL | 11.8–26.9 | 2.3–3.2 | |||
| GY | 14.1–15.5 | 4.9–5.6 | |||
| Xwmc419 | 1BL 59 | 3 | PH | 4.6–36.6 | 2.0–8.4 |
| PdL | 9.4–44.1 | 1.7–7.8 | |||
| Xgwm294 | 2AL 118 | 3 | HD | 6.5–21.0 | 3.9–4.5 |
| PH | 9.8–38.8 | 3.9–6.2 | |||
| PdL | 17.2–44.0 | 3.2–5.6 | |||
| Kpsm | 21.3–33.7 | 2.7–7.5 | |||
| TKW | 25.0–30.1 | 3.7–8.1 | |||
| TW | 7.8–25.6 | 2.8–3.3 | |||
| Xgwm410.1 | 2BS 56 | 3 | TKW | 15.2–26.7 | 3.2–5.9 |
| TW | 7.6–17.9 | 3.4–4.5 | |||
| GY | 14.5–19.2 | 4.3–5.9 | |||
| Xgwm47.1 | 2BL 117 | 3 | PH | 5.0–30.5 | 2.2–5.9 |
| PdL | 8.9–34.6 | 1.4–4.7 | |||
| Kpsm | 6.1–35.6 | 1.9–5.0 | |||
| TKW | 6.3–30.6 | 1.4–5.3 | |||
| Xcfa2234 | 3AS 52 | 2 | Kpsm | 16.7–34.6 | 2.3–3.0 |
| TKW | 7.1–30.4 | 1.7–3.9 | |||
| Xgwm493 | 3BS 11 | 2 | HD | 8.5–16.7 | 2.1–5.9 |
| PH | 7.3–33.4 | 1.7–3.1 | |||
| PdL | 12.6–37.6 | 1.9–4.0 | |||
| TW | 9.1–4.8 | 3.4–2.3 | |||
| GY | 12.2–9.4 | 2.1–3.4 | |||
| Xbarc193 | 4BS 0 | 3 | PH | 7.8–36.4 | 2.8–6.8 |
| PdL | 14.6–34.8 | 2.3–5.2 | |||
| Xgwm1278 | 4BS 13 | 2 | PH | 8.0–34.7 | 1.8–3.2 |
| PdL | 25.5–38.3 | 1.8–3.1 | |||
| Kpsm | 16.7–36.7 | 1.4–2.2 | |||
| TKW | 12.3–32.9 | 1.9–4.2 | |||
| GY | 7.4–15.6 | 1.2–3.0 | |||
| Xbarc303 | 5AS 21 | 2 | HD | 6.2–12.5 | 2.9–5.7 |
| PdL | 20.2–38.5 | 1.8–2.1 | |||
| Kpsm | 5.9–39.1 | 2.0–5.9 | |||
| TKW | 16.0–35.6 | 0.9–5.0 | |||
| TW | 12.9–28.7 | 2.2–3.2 | |||
| Xbarc197 | 5AL 73 | 2 | HD | 6.8–23.7 | 2.1–4.3 |
| Kpsm | 18.4–21.1 | 2.3–4.3 | |||
| GY | 9.5–20.3 | 2.5–4.2 | |||
| Xgwm1570 | 5ALa 15 | 2 | PdL | 8.7–34.4 | 2.0–3.6 |
| Kpsm | 22.6–38.7 | 1.8–5.5 | |||
| TKW | 9.7–33.8 | 2.2–6.9 | |||
| Xwmc256 | 6AL 66 | 2 | HD | 5.2– 22.5 | 2.2–4.4 |
| GY | 8.1–23.8 | 1.5–4.7 | |||
| Xdupw217 | 6BS 0 | 3 | PH | 6.6–32.7 | 1.3–5.6 |
| Kpsm | 6.1–40.0 | 1.5–4.6 | |||
| TKW | 6.7–36.6 | 1.6–5.2 | |||
| GY | 3.0–15.7 | 1.8–4.2 | |||
| Xwmc494 | 6BS 43 | 3 | TW | 3.6–28.2 | 1.5–7.4 |
| Xbarc14 | 6BS 57 | 2 | PH | 3.0–18.9 | 2.3–4.4 |
| Kpsm | 2.7–6.4 | 1.8–2.7 | |||
| TW | 19.6–20.1 | 2.8–3.1 | |||
| GY | 3.7–15.6 | 2.1–3.9 | |||
| Xcfa2257.1 | 7AL 145 | 2 | Kpsm | 18.4–37.4 | 1.5–3.6 |
| TKW | 10.8–33.6 | 1.9–4.6 | |||
| GY | 19.3–20.1 | 2.8–3.1 | |||
| Xgwm333 | 7BL 87 | 3 | HD | 6.0–24.0 | 1.5–3.1 |
| Kpsm | 5.1–38.0 | 1.8–6.1 | |||
| TKW | 10.3–34.3 | 1.8–3.6 | |||
| TW | 4.3–23.5 | 1.2–3.4 | |||
| Xgwm577 | 7BLa 0 | 3 | HD | 5.6–28.5 | 2.2–10.0 |
| PdL | 8.4–35.6 | 1.6–3.8 | |||
| TKW | 8.0–30.5 | 1.4–4.0 | |||
| Xwmc276 | 7BLa 5 | 4 | PH | 5.6–33.2 | 2.0–7.7 |
| PdL | 11.0–42.4 | 2.6–7.1 | |||
| Kpsm | 4.5–33.9 | 1.8–3.8 | |||
| TW | 3.7–23.6 | 1.1–5.6 | |||
| GY | 6.9–16.5 | 2.0–4.8 | |||
R2 ranges of the model (marker+Q matrix as covariate) and of the marker only as from the GLM analysis are also reported.
Second linkage group of the chromosome.
Most of the marker–trait associations which were consistent across environments were identified in the medium- and high-yielding envs.; conversely, the low-yielding envs. often showed association patterns different from each other and from the other environments (Fig. 2). In particular, the chr. regions tagged by Xcfd65 (chr. 1BS), Xwmc419 (1BL), Xgwm294 (2AL), Xcfa2234 (3AS), Xgwm493 (3BS), Xgwm1278 (4BS), Xbarc303 (5AS), Xbarc197 (5AL), Xgwm1570 (5AL), and Xwmc276 (7BL) showed marker–trait associations significant at experiment-wise level or at marker-wise (P 0.01) level across the medium- to high-productivity envs., but only at P 0.05 or mostly not significant in the low-yielding, highly stressed envs. Conversely, only a few regions (Xwmc469, chr. 1AS; Xbarc213 and Xbarc287, chr. 1AL; Ppd-A1, chr. 2AS, and Xwmc256, chr. 6AL) showed significant marker–trait associations across all environments, including those where drought and/or heat stress severely curtailed crop development and yield.
Some markers showed significant effects on several traits; as an example, Xcfd65 (1BS), Xcfa2086 (1BL), Xgwm294 (2AL), Xgwm493 (3BS), Xgwm1278 (4BS), Xbarc303 (5AS), Xbarc197 (5AL), Xbarc14 (6BS), and Xwmc276 (7BL) were associated with either HD or PH and PdL as well as with GY and/or GY components. In addition, Xbarc213, Xbarc287 (1AL), Xgwm410.1 (2BS), Xcfa2234 and Xwmc664 (3A centromeric), Xgwm1570 (5AL near Vrn-A1 locus), Xwmc273.2 and Xcfa2257.1 (7AL) were mostly associated with variation of the yield components (Kpsm and/or TKW), while showing only limited evidence of concomitant significant effects on HD and PH/PdL. Marker Xwmc419 (1BL) was strongly associated with PH and particularly PdL; however, no significant associations with GY and its components were observed. Although significant associations to TW across envs. were usually detected in parallel with TKW and PdL, two markers showed highly significant and consistent effects across envs. on TW only (Xwmc469 on chr. 1AS and Xwmc494 on chr. 6BS), indicating that the genetic basis of this trait is partially independent of TKW.
In some cases, multiple (two or three) markers positioned within a 10 cM interval showed significant associations with the same trait/s suggesting the presence of a few consistent long range-haplotypes, also called linkage blocks, as shown by the concurrence of the associations found in chr. 1AL (Xbarc213–Xcfa2147.2 interval; traits: PdL, Kpsm and TKW), 2AS (Xwmc177–Xgwm1198.3; HD, GY, and its components), 4BS (Xbarc193–Xgwm1278; PH and PdL), 5AS (Xbarc303–Xgwm293; Kpsm and TKW), and 7B (Xwmc276–Xwmc273.1;PH and GY).
The association between the allelic variants in the proximal region of chr. 5AS and TKW has been further investigated with five additional SSR markers near Xgwm293 and Xbarc303, allowing us to identify two common long-range haplotypes in the durum cultivated germplasm in the interval from Xgwm293 to Xbarc117 (ca. 5 cM in total; Fig. 3). In addition, the two haplotypes showed significantly different effects on Kpsm and TKW in the high- and medium-yielding envs. (experiment-wise significance). The significance of such associations peaked on Xbarc303 and Xbarc117 and quickly dropped when considering the adjacent Xwmc805 and Xbarc186 markers.
Marker–trait associations in the proximal region of chromosome 5AS associated with yield components (thousand kernel weight and number of kernels m−2). The association has been investigated in detail using seven polymorphic SSR markers spanning 10 cM in the target region. The marker order has been assumed as from the GrainGenes database (Ta-SSR-2004 and Ta-Synthetic/Opata-BARC-5A consensus maps). White- and grey-filled circles indicate marker-wise significant and highly significant (P 0.05 and P 0.01, respectively) associations. Black-filled circles represent experiment-wise significant associations (Bonferroni's correction P 0.05).
Comparative analysis with previous QTL studies in wheat
Concurrence between our AM results in durum wheat and those reported in previous wheat mapping studies was observed in several cases. As expected, the correspondence was higher when considering QTL studies in tetraploid wheat as compared to hexaploid wheat.
Notably, our results can be compared directly and more accurately with those obtained in a study conducted using 249 RILs derived from a biparental cross (Kofa×Svevo) that were tested in parallel under the same environmental conditions (Maccaferri et al., 2008) under which the AM panel was evaluated. The two elite cvs. Kofa and Svevo as well as other accessions sharing the same genetic backgrounds were included in our AM panel. The QTL analysis carried out on the Kofa×Svevo data for HD, PH, and GY allowed for the identification of 22 different QTLs (Maccaferri et al., 2008), 14 of which showed minor effects across no more than two to three envs. In particular, two major GY QTLs on chrs. 2BL and 3BS showed significant effects in seven and eight envs., with R2 values ranging from 4.8% to 18.1% and from 3.5% to 12.4%, respectively. In both cases, extensive overlap was observed between the LOD profiles of PdL (M Maccaferri, unpublished data), PH and GY, with only limited overlap with HD (Maccaferri et al., 2008). The importance of the distal regions of chr. 3BS (near Xgwm493) in the control of PH and GY in the Kofa×Svevo RILs (plus allele contributed by Kofa) was only partially confirmed in our AM study. In fact, significant effects associated to Xgwm493 were detected in eight and nine envs. for PH and PdL, respectively, but only in two envs. for GY (Fig. 2; see Supplementary Table S6 at JXB online). In keeping with the results reported by Maccaferri et al. (2008), the mean PH and PdL values of the group of accessions with the Kofa allele (see allele A4 at the Xgwm493 locus; in Supplementary Table S6 k at JXB online) were significantly higher than those of the Svevo allele group (allele A3) in most of the envs. Notably, this was not observed for GY. In addition, at the Xgwm493 locus, the Svevo allele was present in the IDuWUE panel with high frequency across subgroups, while the Kofa allele was rare (from one to three accessions) in three subgroups (S1, S4, and S5), rather balanced (18 versus 37) in the S2 subgroup, and prevalent only in the S3 subgroup (23 versus 5). Similar results were obtained for the loci on chr. 2BL near Xgwm846 and Xgwm1027. AM also confirmed the results of Maccaferri et al. (2008) in the Kofa×Svevo population for the QTL for HD found in the Ppd-A1 region (Xcfa2201–Xgwm1198 interval on chr. 2A) and the QTLs for PH identified in the proximal region of chr. 3AL (corresponding to Xgwm1159), limited to the more favourable envs., and on chr. 4BS near Rht-B1. Moreover, our AM study showed that four markers (Xbarc213, Xcfa2147.2, Xbarc287, and Xbarc158.2) spanning ca. 35 cM on the distal region of chr. 1AL were significantly associated with Kpsm and TKW (Fig. 2): at Xbarc213, the effects of the Kofa and Svevo alleles (both present as non-rare variants) on Kpsm and TKW are consistent with the results of the Kofa×Svevo QTL study (M Maccaferri, unpublished data). Although both AM and linkage mapping have clearly highlighted the effect of this region on Kpsm and TKW, the intermarker distances were too large to allow for the detection of shared, long-range haplotype patterns.
Contrary to what was observed in the Kofa×Svevo population, in some cases (e.g. the QTLs for HD on chrs. 2BL and 7BS and for PH and PdL on chr. 7AS) the single markers profiled in the germplasm failed to detect significant associations, most likely due to loss in statistical power caused by unbalanced allelic frequencies and/or a locally elevated LD-decay rate of the target regions in the IDuWUE panel.
As already underlined, AM showed a major effect for yield components (Kpsm and particularly TKW) in the proximal region of chr. 5AS near Xbarc303, with highly significant allelic differences between the two non-rare allelic variants per locus consistently observed across several environments (see Supplementary Table S6 at JXB online). Remarkably, this chr. region is coincident with the location of a major QTL for kernel weight and shape previously identified by Sorrells and co-workers in hexaploid wheat via linkage mapping and subsequently validated through AM (Breseghello and Sorrells, 2006).
In addition, concurrence between these AM results and the QTL locations reported by previous studies in durum and bread wheat was observed particularly for HD, PH, Kpsm, and TKW in the centromeric region of chrs. 1B and 4B and the proximal region of chr. 3AL (Börner et al., 2002; Campbell et al., 2003; Peng et al., 2003; Breseghello and Sorrells, 2006; M Maccaferri, unpublished data).
Discussion
The IDuWUE panel of durum wheat accessions used in this study has been assembled for AM purposes to complement the results obtained in biparental mapping studies. To the best of our knowledge, this is the only AM study in wheat (and other crops as well) that reports and compares results obtained in parallel to those of 15 field trials in which a biparental mapping population was tested under the same environmental conditions across a broad range of water availability. In particular, both the IDuWUE panel and the mapping population were evaluated for GY, yield components, and other morpho-physiological traits in 15 rainfed and irrigated field trials located in the Mediterranean Basin, thus allowing us to measure yield and other traits in a range of environments broadly different for the amount of water available to the crop and other growing conditions (e.g. temperature). Therefore, our study provides a meaningful comparison of the power and effectiveness of AM and biparental approaches in dissecting the genetic basis of adaptation and yield of durum wheat grown in stressed environments.
The 189 accessions of the IDuWUE panel were chosen from 330 elite cvs. with the aim of obtaining a panel of genotypes balanced in their genetic relationships and relatively uniform as to HD in order to minimize the effects that an excessive variability in phenology could have on GY under varying water regimes. The population size of the panel used in this study is larger than that of similar AM studies that have been carried out in wheat (Breseghello and Sorrells, 2006; Chao et al., 2007; Sanguineti et al., 2007; Somers et al., 2007). Further, the choice of a relatively high minimal threshold (equal to 18 counts, i.e. accessions) for rare alleles to be excluded in the association analysis allowed us to obtain estimates of the allelic effects not excessively biased by sampling variance.
Genetic diversity, population structure, and LD extent in the IDuWUE durum panel
The level of genetic diversity present in the IDuWUE panel (5.1 alleles per SSR locus) is comparable with that reported in the wheat elite germplasm collections that have been molecularly characterized up to now. In fact, Somers et al. (2007) found, on average, 6.9 and 5.6 alleles per SSR locus in their survey of 189 bread and 93 durum wheat accessions, respectively, while Chao et al. (2007) reported a mean of 7.2 alleles per SSR locus in a collection of 43 US bread wheat cvs.
Although the accessions included in the IDuWUE panel were pre-selected based on pedigree information, the partitioning of the global genetic diversity in subgroups remains a distinct feature of the IDuWUE panel, similar to what has been observed in other elite hexaploid and tetraploid wheat collections (Chao et al., 2007; Crossa et al., 2007; Somers et al., 2007; Sorrells and Yu, 2009). In fact, for the majority of crop species and, particularly for self-pollinating cereals, the presence of non-random ‘background’ co-ancestry among accessions, i.e. population structure and familial relationship, is a common feature of cvs. and advanced breeding materials (Flint-Garcia et al., 2005; Maccaferri et al., 2005; Camus-Kulandaivelu et al., 2006; Cockram et al., 2008). Since this greatly affects type I and II errors in marker–trait association tests, germplasm collection resources can be exploited for mapping only when the effects due to population structure are appropriately accounted for (Pritchard and Rosenberg, 1999; Breseghello and Sorrells, 2006; Yu et al., 2006; Malosetti et al., 2007).
As to the level of LD, our results are in keeping with those reported by Maccaferri et al. (2005), Breseghello and Sorrells (2006), Chao et al. (2007), and Somers et al. (2007). Collectively, these studies highlighted that LD in collections of elite wheat accessions extends from ca. 1 cM up to 10 cM. This indicates that the achievement of an adequate genome coverage for pursuing an exhaustive genome-wide association analysis in such collections may require up to a few thousand high-quality SSR markers, or thousands of biallelic Single Nucleotide Polymorphism (SNP) markers (Beló et al., 2008), the latter presently unavailable for wheat (Ganal and Röder, 2007). Alternatively, our results (see the association results for Ppd-A1 and linked SSRs) support the conclusions of Chao et al. (2007) that the optimal use of these collections is for a finer mapping and further dissection of QTLs identified in biparental populations via linkage mapping. A thorough characterization of target regions with markers at high density and further phenotyping allowed us to identify with higher precision the haplotype patterns of the accessions and to more accurately investigate the haplotype–phenotype associations (e.g. chr. 5AS near Xbarc303 and Xgwm293).
Genetic basis of the variation for the phenological and yield-related traits
Although the amount of genetic diversity present in most of the elite crop germplasm pools is usually assumed to be relatively limited, a wide variation for the recorded traits was observed in the IDuWUE panel, characterized by an extensive range of phenotypic values, the only exception being represented by HD as a result of the criterion adopted to assemble the panel. The wide range of continuous and normally distributed phenotypes observed in the panel highlights the quantitative nature of the investigated traits and suggests the substantial absence of functional polymorphism at major genes/QTLs, the only exceptions being, not unexpectedly, the Ppd-A1 locus which was associated with HD in 12 envs. and the chr. regions 5AS (Xbarc303) and 5AL (Xgwm1570, near Vrn-A1) associated with TKW in 11 and 13 envs., respectively.
In general, the observed complexity of the genetic control of the agronomic and adaptive traits considered here can be seen as a common feature of genetic studies on quantitative traits when dealing with broad germplasm collections instead of materials derived from bi-parental populations, as recently shown in maize by Buckler and coworkers with the nested-association mapping approach (McMullen et al., 2009). In fact, when dealing with a large pool of unrelated accessions, similar phenotypic values can result from the action of different physiological mechanisms and, consequently, different gene networks, hence reducing the occurrence of sizeable and stable marker–phenotype associations, a more likely possibility with a bi-parental mapping population. This circumstance is particularly relevant for physiologically complex traits such as yield, where similar phenotypic values can be attained via rather different morpho-physiological features (Araus et al., 2003; Reynolds and Tuberosa, 2008) and thus different underlining gene networks. Although this issue is also valid for identifying QTLs in a bi-parental population, its relevance increases rapidly as the variation in phenology and functional complexity of the tested materials escalate. Accordingly, the genetic analysis via AM of complex traits such as yield and yield components may quickly become untractable in collections that include landraces with a highly diverse genetic background and, most importantly, phenology. Although HD was purposely kept within a narrow range when assembling the IDuWUE panel, its correlation with fertility (Kpsm) and GY was often significant, pointing out that even the relatively small range in HD of this collection significantly influences adaptation to the stress conditions commonly present in Mediterranean envs.
In our study, TKW showed a primary relevance for GY as shown by the significant and positive correlations in the low- to medium-yielding envs. Similar considerations pertain to PdL, with the positive correlations with final GY possibly explained by indirect relationships (e.g. via plant water status that could affect peduncle elongation) as well as by relocation capacity as related to the amount of water-soluble carbohydrates stored in the peduncle.
Comparison of results from association and linkage mapping studies
Given the exploratory nature of this study, association results have been reported not only for markers with significant effects at the experiment-wise level, but also for markers showing significance at the marker-wise level, provided that the P 0.05 significance was attained in at least two envs. While increasing the chance of including false-positive associations, this approach highlights the presence of putative QTLs that could have been further masked by residual population structure effects and/or gene×environment and gene×gene interactions (Flint-Garcia et al., 2005). Nevertheless, many of the reported and discussed associations were observed across multiple environments and on the mean data across environments, a result that supports their validity.
Based on the average level of LD and the number and distribution of the SSRs herein considered, it is estimated that our study surveyed the effects of genetic variability present in ca. 50% of the durum wheat genome, a genome coverage rate comparable with those reported in similar AM studies in wheat (Chao et al., 2007; Somers et al., 2007). Notably, the number of accessions considered in our study is twice as large as that surveyed in Chao et al. (2007) and in Somers et al. (2007), thus contributing toward a more comprehensive genome-wide search of the A and B wheat genomes.
Switching from SSR profiling to high-throughput SNP-based genotyping platforms would allow for a genome-wide search with higher effectiveness and accuracy. To date, notwithstanding the ongoing efforts, SNP platforms are not yet available in wheat, mainly due to the allopolyploid and repetitive nature of its genome (Ganal and Röder, 2007).
Considering the marker–trait associations identified in this survey, it is important to underline that a higher consistency of associations across envs. was detected in field trials with high or medium yield. For the low-yielding envs., the patterns of marker–trait associations were highly scattered and inconsistent across envs., hence indicative of gene×environment interactions. Under these conditions, bi-parental mapping may provide a more effective way to dissect the genetic basis of adaptation to water-limited conditions.
The association results indicate that HD, PH, PdL, and TKW were less influenced by environmental conditions compared with GY. Likewise, marker–trait associations consistent across several (about four) envs. were detected for the former traits, but not for GY except for four chr. regions (including the Ppd-A1 locus and the regions tagged by Xgwm410.1 on chr. 2B, Xbarc197 on chr. 5A, and Xbarc14 on chr. 6B) where significant effects on GY were observed in five to six envs. and on the means across envs.
For most of the significant marker-trait associations, the marker R2 values varied between 1% and 5% of the total phenotypic variation (based on single env. tests) and only a few marker–trait combinations showed R2 values reaching 10%, the only exception being the Ppd-A1 allelic variants, with R2 values up to 33%. These results, coupled with the relatively high number of markers significantly associated with the investigated traits, reflect the polygenic nature of the investigated traits and the wider genetic variation present in germplasm collections compared with bi-parental mapping populations. Our association results, together with the comparative analysis, are in keeping with the general observation of Flint-Garcia et al. (2005), who defined a QTL characterized by a 10% R2 value as a ‘substantial QTL’ if detected in traditional biparental mapping populations, and as a ‘major QTL’ when detected in AM analysis.
When inspecting for the presence and distribution of the allelic variants at Ppd-A1, it appeared that substantial molecular variation associated with HD differences was present at this locus in the cultivated Mediterranean durums, which provides the basis for optimizing GY in different environments through the fine-tuning of flowering time and also the crop cycle length. Further, the association test fully validated the recently published observations of Wilhelm et al. (2009) on Ppd-A1 alleles.
Collectively, our results show that, notwithstanding the high selective pressure exherted by almost a century of modern breeding, a sizeable amount of allelic variation is still present in the elite durum germplasm. In this respect, Maccaferri et al. (2003) showed that the level of genetic diversity present in the modern germplasm has actually increased over time, even if a prevailing portion of the molecular variation traces back to a reduced number of foundation-genotypes.
As to the usefulness of AM in combination with the traditional linkage mapping approach, the case of the Ppd-A1 locus can be cited. The QTL analysis carried out by Maccaferri et al. (2008) in Kofa×Svevo led to the identification of a major QTL for HD in the chr. region surrounding Ppd-A1 and the AM analysis based on the sequence variation at the causative gene allowed for a more comprehensive and accurate evaluation of the effects of the three alleles prevalent in the IDuWUE panel. It is interesting to note that when considering the four SSR markers mapping within a 10 cM interval around Ppd-A1, a rather low marker–trait association was obtained (with the exception, to some extent, of Xcfa2201), even though the LD between Ppd-A1 and the markers tested was always high and highly significant. These results indicate that a more accurate estimate of the position and effect of a functional locus can be attained with the approach and materials reported here as long as a thorough haplotyping of the collection is carried out by saturating the target region with additional markers.
As a further example of complementarity between association and biparental mapping, our AM study highlighted the presence of a major QTL for TKW on chr. 5AS (near Xbarc303 and Xgwm293) that was previously evidenced for kernel size in bread wheat using both linkage and AM (Breseghello and Sorrells, 2006) as well as in the durum RIL population Colosseo×Lloyd (M Maccaferri, unpublished results).
In addition to the confirmation/validation of the presence of main effects at key chr. regions already identified with biparental mapping, the use of AM in a comprehensive germplasm collection such as the IDuWUE panel allowed us to detect significant effects at several chr. regions for which the presence of allelic variation for agronomic traits has not yet been reported in durum wheat. Linkage analysis in appropriately designed mapping populations will indicate to what extent these regions identified through AM might play a more crucial role in particular genetic backgrounds and might provide valuable targets for marker-assisted selection.
Conclusions
Our results indicate the suitability of using the IDuWUE panel to investigate via AM the genetic control in durum wheat of drought-adaptive traits and their role in determining yield across different soil-moisture regimes. As compared with envs. most severely affected by drought, the number and consistency of marker–trait associations across envs. were higher under more favourable soil moisture conditions. As to grain yield, significant marker effects were consistently detected in only a rather limited number of cases and with R2 values usually lower than 5%. Collectively, our results suggest that, as the level of moisture stress increases, the power to detect the relevant loci for grain yield via AM decreases, most likely because under such conditions similar grain yield values can be attained by different genotypes through different adaptive strategies and corresponding gene networks, thus undermining the occurrence of significant marker–trait associations. This notwithstanding, AM revealed the influence of several chr. regions on the variability of adaptive and drought-responsive traits such as heading date, plant height, ear peduncle length, grain volume weight, and yield component traits. Although a number of these regions have already been described in biparental populations, our study has contributed to assess their relevance in the cultivated germplasm of durum wheat and has highlighted the presence of novel QTLs, thus contributing valuable information for setting up new mapping and breeding populations.
Abbreviations
- AM
association mapping
- chr.
chromosome
- cv.
cultivar
- PdL
ear peduncle length
- Em
emergence
- env.
environment
- GLM
general linear model
- GY
grain yield
- HD
heading date
- Jn
jointing
- Kpsm
number of kernels per square meter
- LD
linkage disequilibrium
- MD
maturity date
- MLM
mixed linear model
- PH
plant height
- QTL
Quantitative Trait Locus
- RIL
Recombinant Inbred Line
- SNP
Single Nucleotide Polymorphism
- SSR
Simple Sequence Repeat
- Sm
soil moisture
- T
temperature
- TKW
thousand kernel weight
- TW
test weight
- Wi
water input
This research was supported by the European Union (IDuWUE project, INCO contract number ICA3-CT-2002-10 028 and TRITIMED Project, Sixth Framework Programme, Contract no. 509136) and the Italian Ministero Politiche Agricole Alimentari e Forestali (Progetto Internazionale Sequenziamento Genoma Frumenti: la mappa fisica del cromosoma 5). The Centre UdL-IRTA (part of the Centre CONSOLIDER INGENIO 2010 on Agrigenomics funded by the Spanish Ministry of Education and Science) and the Interdepartmental Centre for Biotechnology (University of Bologna, Italy) are gratefully acknowledged. The authors thank Sandra Stefanelli, Silvana Tamassia, Stefano Vecchi, and Marco Mantovani for technical assistance.



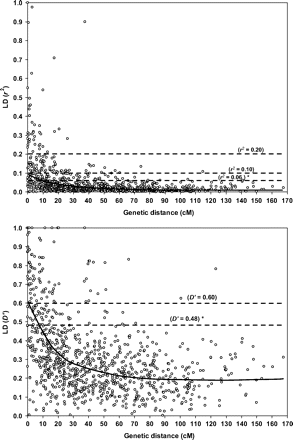
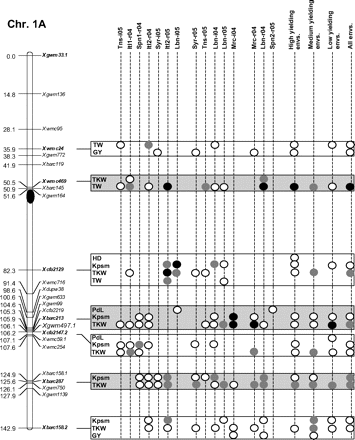
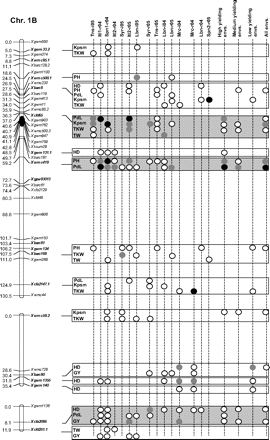
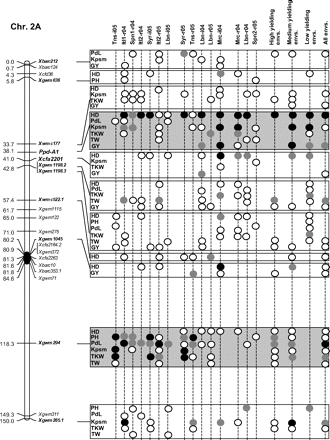
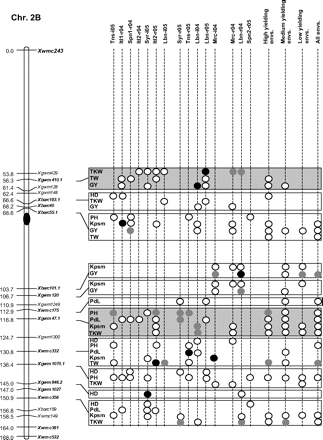
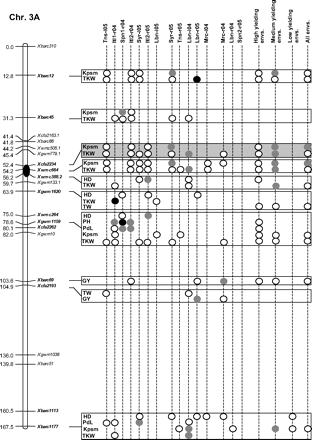
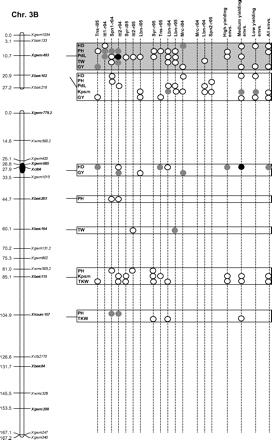


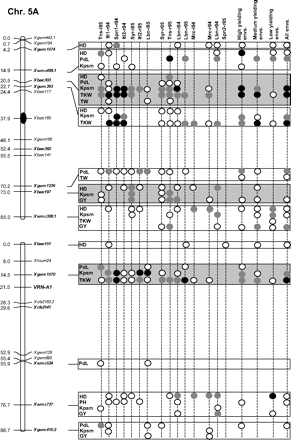

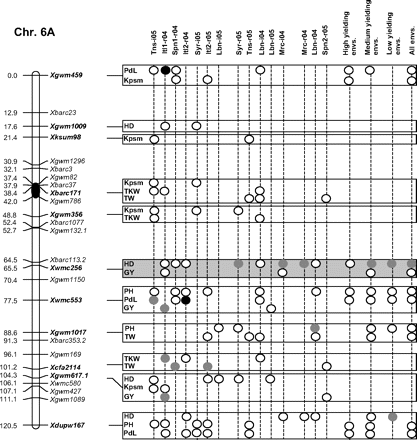
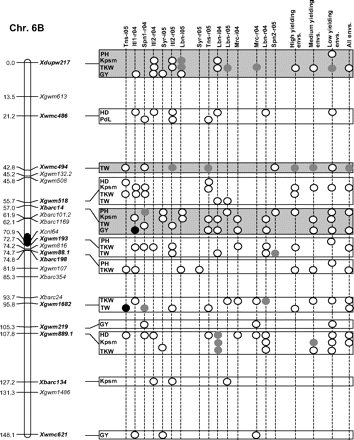
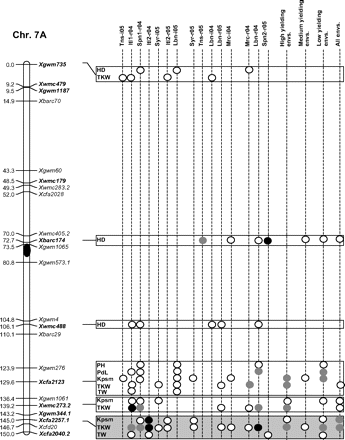

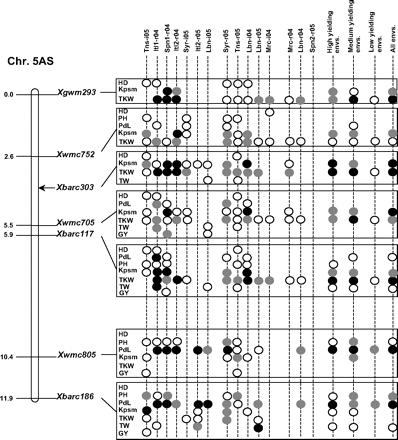

Comments