Volume 46, Issue D1
4 January 2018
Cover image

Cover image

Cover: The cover image shows cards representing entries in the ChannelsDB database. Each entry shows a channel, pore or tunnel important for the biological function of a protein, along with descriptors of its length and composition. For example, the biotransformation of codeine starts with its passage into the active site of cytochrome P450 2D6 via the substrate channel and, after the reaction on haem, morphine is released via a product channel.
For more information, see article by Pravda, L.et al. , pages D399–D405 in this issue.
For more information, see article by Pravda, L.
ISSN 0305-1048
EISSN 1362-4962
Issue navigation
Volume 46, Issue D1, 4 January 2018
Database issue
Editorial
The 2018 Nucleic Acids Research database issue and the online molecular biology database collection
Daniel J Rigden and Xosé M Fernández
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1–D7, https://doi.org/10.1093/nar/gkx1235
Database Resources
Database resources of the National Center for Biotechnology Information
NCBI Resource Coordinators
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D8–D13, https://doi.org/10.1093/nar/gkx1095
Database Resources of the BIG Data Center in 2018
BIG Data Center Members
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D14–D20, https://doi.org/10.1093/nar/gkx897
The European Bioinformatics Institute in 2017: data coordination and integration
Charles E Cook and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D21–D29, https://doi.org/10.1093/nar/gkx1154
Nucleic acid sequence, structure, and regulation
DNA Data Bank of Japan: 30th anniversary
Yuichi Kodama and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D30–D35, https://doi.org/10.1093/nar/gkx926
The European Nucleotide Archive in 2017
Nicole Silvester and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D36–D40, https://doi.org/10.1093/nar/gkx1125
GenBank
Dennis A Benson and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D41–D47, https://doi.org/10.1093/nar/gkx1094
The international nucleotide sequence database collaboration
Ilene Karsch-Mizrachi and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D48–D51, https://doi.org/10.1093/nar/gkx1097
3DIV: A 3D-genome Interaction Viewer and database
Dongchan Yang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D52–D57, https://doi.org/10.1093/nar/gkx1017
ASpedia: a comprehensive encyclopedia of human alternative splicing
Daejin Hyung and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D58–D63, https://doi.org/10.1093/nar/gkx1014
CirGRDB: a database for the genome-wide deciphering circadian genes and regulators
Xianfeng Li and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D64–D70, https://doi.org/10.1093/nar/gkx944
dbCoRC: a database of core transcriptional regulatory circuitries modeled by H3K27ac ChIP-seq signals
Moli Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D71–D77, https://doi.org/10.1093/nar/gkx796
DiseaseEnhancer: a resource of human disease-associated enhancer catalog
Guanxiong Zhang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D78–D84, https://doi.org/10.1093/nar/gkx920
dreamBase: DNA modification, RNA regulation and protein binding of expressed pseudogenes in human health and disease
Ling-Ling Zheng and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D85–D91, https://doi.org/10.1093/nar/gkx972
EpiDenovo: a platform for linking regulatory de novo mutations to developmental epigenetics and diseases
Fengbiao Mao and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D92–D99, https://doi.org/10.1093/nar/gkx918
EVLncRNAs: a manually curated database for long non-coding RNAs validated by low-throughput experiments
Bailing Zhou and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D100–D105, https://doi.org/10.1093/nar/gkx677
exoRBase: a database of circRNA, lncRNA and mRNA in human blood exosomes
Shengli Li and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D106–D112, https://doi.org/10.1093/nar/gkx891
HEDD: Human Enhancer Disease Database
Zhen Wang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D113–D120, https://doi.org/10.1093/nar/gkx988
ICG: a wiki-driven knowledgebase of internal control genes for RT-qPCR normalization
Jian Sang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D121–D126, https://doi.org/10.1093/nar/gkx875
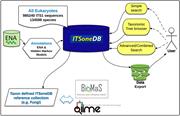
ITSoneDB: a comprehensive collection of eukaryotic ribosomal RNA Internal Transcribed Spacer 1 (ITS1) sequences
Monica Santamaria and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D127–D132, https://doi.org/10.1093/nar/gkx855
Lnc2Meth: a manually curated database of regulatory relationships between long non-coding RNAs and DNA methylation associated with human disease
Hui Zhi and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D133–D138, https://doi.org/10.1093/nar/gkx985
m6AVar: a database of functional variants involved in m6A modification
Yueyuan Zheng and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D139–D145, https://doi.org/10.1093/nar/gkx895
MeDReaders: a database for transcription factors that bind to methylated DNA
Guohua Wang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D146–D151, https://doi.org/10.1093/nar/gkx1096
MINTbase v2.0: a comprehensive database for tRNA-derived fragments that includes nuclear and mitochondrial fragments from all The Cancer Genome Atlas projects
Venetia Pliatsika and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D152–D159, https://doi.org/10.1093/nar/gkx1075
miRCarta: a central repository for collecting miRNA candidates
Christina Backes and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D160–D167, https://doi.org/10.1093/nar/gkx851
mirTrans: a resource of transcriptional regulation on microRNAs for human cell lines
Xu Hua and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D168–D174, https://doi.org/10.1093/nar/gkx996
MGA repository: a curated data resource for ChIP-seq and other genome annotated data
René Dréos and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D175–D180, https://doi.org/10.1093/nar/gkx995
MSDD: a manually curated database of experimentally supported associations among miRNAs, SNPs and human diseases
Ming Yue and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D181–D185, https://doi.org/10.1093/nar/gkx1035
OverGeneDB: a database of 5′ end protein coding overlapping genes in human and mouse genomes
Wojciech Rosikiewicz and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D186–D193, https://doi.org/10.1093/nar/gkx948
RISE: a database of RNA interactome from sequencing experiments
Jing Gong and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D194–D201, https://doi.org/10.1093/nar/gkx864
RNArchitecture: a database and a classification system of RNA families, with a focus on structural information
Pietro Boccaletto and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D202–D205, https://doi.org/10.1093/nar/gkx966
TranslatomeDB: a comprehensive database and cloud-based analysis platform for translatome sequencing data
Wanting Liu and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D206–D212, https://doi.org/10.1093/nar/gkx1034
APPRIS 2017: principal isoforms for multiple gene sets
Jose Manuel Rodriguez and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D213–D217, https://doi.org/10.1093/nar/gkx997
ARED-Plus: an updated and expanded database of AU-rich element-containing mRNAs and pre-mRNAs
Tala Bakheet and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D218–D220, https://doi.org/10.1093/nar/gkx975
Consensus coding sequence (CCDS) database: a standardized set of human and mouse protein-coding regions supported by expert curation
Shashikant Pujar and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D221–D228, https://doi.org/10.1093/nar/gkx1031
DBTSS/DBKERO for integrated analysis of transcriptional regulation
Ayako Suzuki and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D229–D238, https://doi.org/10.1093/nar/gkx1001
DIANA-TarBase v8: a decade-long collection of experimentally supported miRNA–gene interactions
Dimitra Karagkouni and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D239–D245, https://doi.org/10.1093/nar/gkx1141
Expression Atlas: gene and protein expression across multiple studies and organisms
Irene Papatheodorou and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D246–D251, https://doi.org/10.1093/nar/gkx1158
HOCOMOCO: towards a complete collection of transcription factor binding models for human and mouse via large-scale ChIP-Seq analysis
Ivan V Kulakovskiy and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D252–D259, https://doi.org/10.1093/nar/gkx1106
JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework
Aziz Khan and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D260–D266, https://doi.org/10.1093/nar/gkx1126
ReMap 2018: an updated atlas of regulatory regions from an integrative analysis of DNA-binding ChIP-seq experiments
Jeanne Chèneby and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D267–D275, https://doi.org/10.1093/nar/gkx1092
lncRNASNP2: an updated database of functional SNPs and mutations in human and mouse lncRNAs
Ya-Ru Miao and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D276–D280, https://doi.org/10.1093/nar/gkx1004
MeT-DB V2.0: elucidating context-specific functions of N6-methyl-adenosine methyltranscriptome
Hui Liu and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D281–D287, https://doi.org/10.1093/nar/gkx1080
MethBank 3.0: a database of DNA methylomes across a variety of species
Rujiao Li and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D288–D295, https://doi.org/10.1093/nar/gkx1139
miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions
Chih-Hung Chou and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D296–D302, https://doi.org/10.1093/nar/gkx1067
MODOMICS: a database of RNA modification pathways. 2017 update
Pietro Boccaletto and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D303–D307, https://doi.org/10.1093/nar/gkx1030
NONCODEV5: a comprehensive annotation database for long non-coding RNAs
ShuangSang Fang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D308–D314, https://doi.org/10.1093/nar/gkx1107
PolyA_DB 3 catalogs cleavage and polyadenylation sites identified by deep sequencing in multiple genomes
Ruijia Wang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D315–D319, https://doi.org/10.1093/nar/gkx1000
PRODORIC2: the bacterial gene regulation database in 2018
Denitsa Eckweiler and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D320–D326, https://doi.org/10.1093/nar/gkx1091
RMBase v2.0: deciphering the map of RNA modifications from epitranscriptome sequencing data
Jia-Jia Xuan and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D327–D334, https://doi.org/10.1093/nar/gkx934
Rfam 13.0: shifting to a genome-centric resource for non-coding RNA families
Ioanna Kalvari and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D335–D342, https://doi.org/10.1093/nar/gkx1038
TFClass: expanding the classification of human transcription factors to their mammalian orthologs
Edgar Wingender and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D343–D347, https://doi.org/10.1093/nar/gkx987
YEASTRACT: an upgraded database for the analysis of transcription regulatory networks in Saccharomyces cerevisiae
Miguel C Teixeira and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D348–D353, https://doi.org/10.1093/nar/gkx842
miRandola 2017: a curated knowledge base of non-invasive biomarkers
Francesco Russo and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D354–D359, https://doi.org/10.1093/nar/gkx854
mirDIP 4.1—integrative database of human microRNA target predictions
Tomas Tokar and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D360–D370, https://doi.org/10.1093/nar/gkx1144
MNDR v2.0: an updated resource of ncRNA–disease associations in mammals
Tianyu Cui and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D371–D374, https://doi.org/10.1093/nar/gkx1025
Updates to the RNA mapping database (RMDB), version 2
Joseph D Yesselman and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D375–D379, https://doi.org/10.1093/nar/gkx873
TRRUST v2: an expanded reference database of human and mouse transcriptional regulatory interactions
Heonjong Han and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D380–D386, https://doi.org/10.1093/nar/gkx1013
Protein sequence and structure, motifs, and domains
AmyPro: a database of proteins with validated amyloidogenic regions
Mihaly Varadi and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D387–D392, https://doi.org/10.1093/nar/gkx950
Anti-CRISPRdb: a comprehensive online resource for anti-CRISPR proteins
Chuan Dong and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D393–D398, https://doi.org/10.1093/nar/gkx835
ChannelsDB: database of biomacromolecular tunnels and pores
Lukáš Pravda and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D399–D405, https://doi.org/10.1093/nar/gkx868
STCRDab: the structural T-cell receptor database
Jinwoo Leem and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D406–D412, https://doi.org/10.1093/nar/gkx971
Target-Pathogen: a structural bioinformatic approach to prioritize drug targets in pathogens
Ezequiel J Sosa and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D413–D418, https://doi.org/10.1093/nar/gkx1015
VDJdb: a curated database of T-cell receptor sequences with known antigen specificity
Mikhail Shugay and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D419–D427, https://doi.org/10.1093/nar/gkx760
The eukaryotic linear motif resource – 2018 update
Marc Gouw and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D428–D434, https://doi.org/10.1093/nar/gkx1077
Gene3D: Extensive prediction of globular domains in proteins
Tony E Lewis and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D435–D439, https://doi.org/10.1093/nar/gkx1069
GPCRdb in 2018: adding GPCR structure models and ligands
Gáspár Pándy-Szekeres and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D440–D446, https://doi.org/10.1093/nar/gkx1109
iUUCD 2.0: an update with rich annotations for ubiquitin and ubiquitin-like conjugations
Jiaqi Zhou and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D447–D453, https://doi.org/10.1093/nar/gkx1041
KNOTTIN: the database of inhibitor cystine knot scaffold after 10 years, toward a systematic structure modeling
Guillaume Postic and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D454–D458, https://doi.org/10.1093/nar/gkx1084
MetalPDB in 2018: a database of metal sites in biological macromolecular structures
Valeria Putignano and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D459–D464, https://doi.org/10.1093/nar/gkx989
Minimotif Miner 4: a million peptide minimotifs and counting
Kenneth F Lyon and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D465–D470, https://doi.org/10.1093/nar/gkx1085
MobiDB 3.0: more annotations for intrinsic disorder, conformational diversity and interactions in proteins
Damiano Piovesan and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D471–D476, https://doi.org/10.1093/nar/gkx1071
The OMA orthology database in 2018: retrieving evolutionary relationships among all domains of life through richer web and programmatic interfaces
Adrian M Altenhoff and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D477–D485, https://doi.org/10.1093/nar/gkx1019
PDBe: towards reusable data delivery infrastructure at protein data bank in Europe
Saqib Mir and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D486–D492, https://doi.org/10.1093/nar/gkx1070
20 years of the SMART protein domain annotation resource
Ivica Letunic and Peer Bork
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D493–D496, https://doi.org/10.1093/nar/gkx922
An update on sORFs.org: a repository of small ORFs identified by ribosome profiling
Volodimir Olexiouk and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D497–D502, https://doi.org/10.1093/nar/gkx1130
NLSdb—major update for database of nuclear localization signals and nuclear export signals
Michael Bernhofer and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D503–D508, https://doi.org/10.1093/nar/gkx1021
Metabolic and signalling pathways, enzymes
ClusterCAD: a computational platform for type I modular polyketide synthase design
Clara H Eng and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D509–D515, https://doi.org/10.1093/nar/gkx893
dbCAN-seq: a database of carbohydrate-active enzyme (CAZyme) sequence and annotation
Le Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D516–D521, https://doi.org/10.1093/nar/gkx894
The DifferentialNet database of differential protein–protein interactions in human tissues
Omer Basha and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D522–D526, https://doi.org/10.1093/nar/gkx981
DISNOR: a disease network open resource
Prisca Lo Surdo and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D527–D534, https://doi.org/10.1093/nar/gkx876
fusionDB: assessing microbial diversity and environmental preferences via functional similarity networks
Chengsheng Zhu and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D535–D541, https://doi.org/10.1093/nar/gkx1060
iPTMnet: an integrated resource for protein post-translational modification network discovery
Hongzhan Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D542–D550, https://doi.org/10.1093/nar/gkx1104
jMorp: Japanese Multi Omics Reference Panel
Shu Tadaka and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D551–D557, https://doi.org/10.1093/nar/gkx978
Data Portal for the Library of Integrated Network-based Cellular Signatures (LINCS) program: integrated access to diverse large-scale cellular perturbation response data
Amar Koleti and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D558–D566, https://doi.org/10.1093/nar/gkx1063
Molecular Interaction Search Tool (MIST): an integrated resource for mining gene and protein interaction data
Yanhui Hu and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D567–D574, https://doi.org/10.1093/nar/gkx1116
PAMDB: a comprehensive Pseudomonas aeruginosa metabolome database
Weiliang Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D575–D580, https://doi.org/10.1093/nar/gkx1061
The online Tabloid Proteome: an annotated database of protein associations
Surya Gupta and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D581–D585, https://doi.org/10.1093/nar/gkx930
The TriForC database: a comprehensive up-to-date resource of plant triterpene biosynthesis
Karel Miettinen and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D586–D594, https://doi.org/10.1093/nar/gkx925
TCSBN: a database of tissue and cancer specific biological networks
Sunjae Lee and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D595–D600, https://doi.org/10.1093/nar/gkx994
FunCoup 4: new species, data, and visualization
Christoph Ogris and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D601–D607, https://doi.org/10.1093/nar/gkx1138
HMDB 4.0: the human metabolome database for 2018
David S Wishart and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D608–D617, https://doi.org/10.1093/nar/gkx1089
Mechanism and Catalytic Site Atlas (M-CSA): a database of enzyme reaction mechanisms and active sites
António J M Ribeiro and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D618–D623, https://doi.org/10.1093/nar/gkx1012
The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database
Neil D Rawlings and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D624–D632, https://doi.org/10.1093/nar/gkx1134
The MetaCyc database of metabolic pathways and enzymes
Ron Caspi and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D633–D639, https://doi.org/10.1093/nar/gkx935
MoonProt 2.0: an expansion and update of the moonlighting proteins database
Chang Chen and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D640–D644, https://doi.org/10.1093/nar/gkx1043
MultitaskProtDB-II: an update of a database of multitasking/moonlighting proteins
Luís Franco-Serrano and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D645–D648, https://doi.org/10.1093/nar/gkx1066
The Reactome Pathway Knowledgebase
Antonio Fabregat and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D649–D655, https://doi.org/10.1093/nar/gkx1132
SABIO-RK: an updated resource for manually curated biochemical reaction kinetics
Ulrike Wittig and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D656–D660, https://doi.org/10.1093/nar/gkx1065
WikiPathways: a multifaceted pathway database bridging metabolomics to other omics research
Denise N Slenter and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D661–D667, https://doi.org/10.1093/nar/gkx1064
PAGER 2.0: an update to the pathway, annotated-list and gene-signature electronic repository for Human Network Biology
Zongliang Yue and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D668–D676, https://doi.org/10.1093/nar/gkx1040
PULDB: the expanded database of Polysaccharide Utilization Loci
Nicolas Terrapon and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D677–D683, https://doi.org/10.1093/nar/gkx1022
Viruses, bacteria, protozoa and fungi
MicrobiomeDB: a systems biology platform for integrating, mining and analyzing microbiome experiments
Francislon S Oliveira and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D684–D691, https://doi.org/10.1093/nar/gkx1027
The MAR databases: development and implementation of databases specific for marine metagenomics
Terje Klemetsen and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D692–D699, https://doi.org/10.1093/nar/gkx1036
MVP: a microbe–phage interaction database
Na L Gao and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D700–D707, https://doi.org/10.1093/nar/gkx1124
Virus taxonomy: the database of the International Committee on Taxonomy of Viruses (ICTV)
Elliot J Lefkowitz and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D708–D717, https://doi.org/10.1093/nar/gkx932
ANISEED 2017: extending the integrated ascidian database to the exploration and evolutionary comparison of genome-scale datasets
Matija Brozovic and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D718–D725, https://doi.org/10.1093/nar/gkx1108
EBI Metagenomics in 2017: enriching the analysis of microbial communities, from sequence reads to assemblies
Alex L Mitchell and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D726–D735, https://doi.org/10.1093/nar/gkx967
Saccharomyces genome database informs human biology
Marek S Skrzypek and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D736–D742, https://doi.org/10.1093/nar/gkx1112
SubtiWiki in 2018: from genes and proteins to functional network annotation of the model organism Bacillus subtilis
Bingyao Zhu and Jörg Stülke
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D743–D748, https://doi.org/10.1093/nar/gkx908
TADB 2.0: an updated database of bacterial type II toxin–antitoxin loci
Yingzhou Xie and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D749–D753, https://doi.org/10.1093/nar/gkx1033
Human genome, model organisms, comparative genomics
Ensembl 2018
Daniel R Zerbino and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D754–D761, https://doi.org/10.1093/nar/gkx1098
The UCSC Genome Browser database: 2018 update
Jonathan Casper and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D762–D769, https://doi.org/10.1093/nar/gkx1020
IMOTA: an interactive multi-omics tissue atlas for the analysis of human miRNA–target interactions
Valeria Palmieri and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D770–D775, https://doi.org/10.1093/nar/gkx701
PICKLES: the database of pooled in-vitro CRISPR knockout library essentiality screens
Walter F Lenoir and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D776–D780, https://doi.org/10.1093/nar/gkx993
SCPortalen: human and mouse single-cell centric database
Imad Abugessaisa and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D781–D787, https://doi.org/10.1093/nar/gkx949
StemMapper: a curated gene expression database for stem cell lineage analysis
José P Pinto and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D788–D793, https://doi.org/10.1093/nar/gkx921
The Encyclopedia of DNA elements (ENCODE): data portal update
Carrie A Davis and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D794–D801, https://doi.org/10.1093/nar/gkx1081
Ensembl Genomes 2018: an integrated omics infrastructure for non-vertebrate species
Paul Julian Kersey and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D802–D808, https://doi.org/10.1093/nar/gkx1011
FlyAtlas 2: a new version of the Drosophila melanogaster expression atlas with RNA-Seq, miRNA-Seq and sex-specific data
David P Leader and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D809–D815, https://doi.org/10.1093/nar/gkx976
Genomicus 2018: karyotype evolutionary trees and on-the-fly synteny computing
Nga Thi Thuy Nguyen and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D816–D822, https://doi.org/10.1093/nar/gkx1003
GWIPS-viz: 2018 update
Audrey M Michel and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D823–D830, https://doi.org/10.1093/nar/gkx790
Expanded and updated data and a query pipeline for iBeetle-Base
Jürgen Dönitz and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D831–D835, https://doi.org/10.1093/nar/gkx984
Mouse Genome Database (MGD)-2018: knowledgebase for the laboratory mouse
Cynthia L Smith and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D836–D842, https://doi.org/10.1093/nar/gkx1006
Mouse Phenome Database: an integrative database and analysis suite for curated empirical phenotype data from laboratory mice
Molly A Bogue and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D843–D850, https://doi.org/10.1093/nar/gkx1082
RefSeq: an update on prokaryotic genome annotation and curation
Daniel H Haft and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D851–D860, https://doi.org/10.1093/nar/gkx1068
Xenbase: a genomic, epigenomic and transcriptomic model organism database
Kamran Karimi and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D861–D868, https://doi.org/10.1093/nar/gkx936
WormBase 2017: molting into a new stage
Raymond Y N Lee and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D869–D874, https://doi.org/10.1093/nar/gkx998
iSyTE 2.0: a database for expression-based gene discovery in the eye
Atul Kakrana and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D875–D885, https://doi.org/10.1093/nar/gkx837
Genomic variation, diseases, and drugs
AAgMarker 1.0: a resource of serological autoantigen biomarkers for clinical diagnosis and prognosis of various human diseases
Jianbo Pan and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D886–D893, https://doi.org/10.1093/nar/gkx770
aBiofilm: a resource of anti-biofilm agents and their potential implications in targeting antibiotic drug resistance
Akanksha Rajput and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D894–D900, https://doi.org/10.1093/nar/gkx1157
ActiveDriverDB: human disease mutations and genome variation in post-translational modification sites of proteins
Michal Krassowski and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D901–D910, https://doi.org/10.1093/nar/gkx973
ADReCS-Target: target profiles for aiding drug safety research and application
Li-Hong Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D911–D917, https://doi.org/10.1093/nar/gkx899
CR2Cancer: a database for chromatin regulators in human cancer
Beibei Ru and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D918–D924, https://doi.org/10.1093/nar/gkx877
CSCD: a database for cancer-specific circular RNAs
Siyu Xia and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D925–D929, https://doi.org/10.1093/nar/gkx863
ECOdrug: a database connecting drugs and conservation of their targets across species
Bas Verbruggen and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D930–D936, https://doi.org/10.1093/nar/gkx1024
eRAM: encyclopedia of rare disease annotations for precision medicine
Jinmeng Jia and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D937–D943, https://doi.org/10.1093/nar/gkx1062
Genome Variation Map: a data repository of genome variations in BIG Data Center
Shuhui Song and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D944–D949, https://doi.org/10.1093/nar/gkx986
HCMDB: the human cancer metastasis database
Guantao Zheng and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D950–D955, https://doi.org/10.1093/nar/gkx1008
LinkedOmics: analyzing multi-omics data within and across 32 cancer types
Suhas V Vasaikar and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D956–D963, https://doi.org/10.1093/nar/gkx1090
mSignatureDB: a database for deciphering mutational signatures in human cancers
Po-Jung Huang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D964–D970, https://doi.org/10.1093/nar/gkx1133
PancanQTL: systematic identification of cis-eQTLs and trans-eQTLs in 33 cancer types
Jing Gong and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D971–D976, https://doi.org/10.1093/nar/gkx861
PedAM: a database for Pediatric Disease Annotation and Medicine
Jinmeng Jia and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D977–D983, https://doi.org/10.1093/nar/gkx1049
PGG.Population: a database for understanding the genomic diversity and genetic ancestry of human populations
Chao Zhang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D984–D993, https://doi.org/10.1093/nar/gkx1032
PharmacoDB: an integrative database for mining in vitro anticancer drug screening studies
Petr Smirnov and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D994–D1002, https://doi.org/10.1093/nar/gkx911
PopHuman: the human population genomics browser
Sònia Casillas and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1003–D1010, https://doi.org/10.1093/nar/gkx943
SBCDDB: Sleeping Beauty Cancer Driver Database for gene discovery in mouse models of human cancers
Justin Y Newberg and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1011–D1017, https://doi.org/10.1093/nar/gkx956
SEECancer: a resource for somatic events in evolution of cancer genome
Hongyi Zhang and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1018–D1026, https://doi.org/10.1093/nar/gkx964
TC3A: The Cancer 3′ UTR Atlas
Xin Feng and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1027–D1030, https://doi.org/10.1093/nar/gkx892
TissGDB: tissue-specific gene database in cancer
Pora Kim and others
Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D1031–D1038, https://doi.org/10.1093/nar/gkx850
Advertisement
Advertisement