-
PDF
- Split View
-
Views
-
Cite
Cite
Yao Chen, Qiaofeng Yang, Sihong Sang, Zhaoyun Wei, Peng Wang, Rice Inositol Polyphosphate Kinase (OsIPK2) Directly Interacts with OsIAA11 to Regulate Lateral Root Formation, Plant and Cell Physiology, Volume 58, Issue 11, November 2017, Pages 1891–1900, https://doi.org/10.1093/pcp/pcx125
Close - Share Icon Share
Abstract
The plant hormone auxin controls many aspects of plant growth and development by promoting the degradation of Auxin/Indole-3-acetic acid (Aux/IAA) proteins. The domain II (DII) of Aux/IAA proteins is sufficient for eliciting the degradation by directly interacting with the auxin receptor F-box protein TIR1 to form a TIR1/AFBs-Aux/IAA complex in an auxin-dependent manner. However, the underlying mechanisms of fine-tuning Aux/IAA degradation by auxin stimuli remain to be elucidated. Here, we show that OsIPK2, a rice (Oryza sativa) inositol polyphosphate kinase, directly interacts with an Aux/IAA protein OsIAA11 to repress its degradation. In a rice protoplast transient expression system, the auxin-induced degradation of Myc-OsIAA11 fusion was delayed by co-expressed GFP-OsIPK2 proteins. Furthermore, expressing additional OsIPK2 or its N-terminal amino acid sequence enhanced the accumulation of OsIAA11 proteins in transgenic plants, which in turn caused defects in lateral root formation and auxin response. Taken together, we identify a novel co-factor of Aux/IAA in auxin signaling and demonstrate its role in regulating lateral root development.
Introduction
Auxin is a major plant hormone that controls plant growth and development by regulating the expression of hundreds of auxin-responsive genes. The cooperative actions of auxin receptor TIR1/AFBs F-box proteins, Aux/IAA transcriptional repressors and auxin response factors (ARFs) are required for auxin signal transduction. Aux/IAA negatively regulates auxin-mediated transcription by binding to the ARF transcriptional factor through a conserved domain III and IV (DIII, IV) (Guilfoyle et al. 1998). The EAR motif of Aux/IAA in domain I (DI) binds TOPLESS (TPL) co-repressor to execute its repressive activity (Tiwari et al. 2004, Szemenyei et al. 2008). During auxin-induced transcriptional regulation, auxin promotes the association of TIR1/AFB F-box protein and domain II (DII) to form a co-receptor complex, leading to the polyubiquitination and degradation of Aux/IAA repressors by 26S proteasome system (Gray et al. 2001, Ramos et al. 2001, Tan et al. 2007, Villalobos et al. 2012).
The DII of Aux/IAA contains a conserved sequence ‘GWPPV/I’ called degron that is necessary for the instability of this protein. Crystal structure of Arabidopsis TIR1-IAA7 reveals that the conserved central ‘GWPP’ directly binds to TIR1 pocket with the assistance of auxin (Tan et al. 2007). A direct cis/trans isomerization modification of degradation signal was reported to facilitate the TIR1-Aux/IAA interaction and the proteasomal degradation of Aux/IAA (Jing et al. 2015). The dominant gain-of-function mutations in the degron stabilize Aux/IAA repressors and result in pleiotropic abnormal phenotypes in higher plant Arabidopsis and rice (Uehara et al. 2008, Jun et al. 2011, Kitomi et al. 2012, Zhu et al. 2012). Although DII dominantly controls the rapid degradation, the Aux/IAA family proteins with conserved degron motif display diversified degradation rate (Dreher et al. 2006, Shimizu-Mitao and Kakimoto 2014). Except for the potential rate motifs flanking the degron, cofactors also contribute to the dynamic turnover of Aux/IAA (Moss et al. 2015). A RDV capsid protein P2 has been recently reported to bind DII of OsIAA10 and inhibit its degradation, which enhancing the viral infection (Jin et al. 2016). These results indicate that the auxin-mediated plant development can be tuned by altering the degradation rate of Aux/IAA. However, the mechanism on how plant regulates the stability of Aux/IAA proteins is still elusive.
Inositol polyphosphate kinase (IPK2), also called IPMK, plays an important role in inositol polyphosphate metabolism, converting inositol 1,4,5-triphosphate (IP3) into IP4, and IP4 to IP4. Mammalian IPMK also has non-catalytic signaling functions by directly interacting with multiple signaling factors, such as mTOR (mammalian target of rapamycin), AMPK, p53 and TRAF6 (Kim et al. 2011, Bang et al. 2012, Xu et al. 2013, Kim et al. 2017). In yeast, as an indispensable component of the ArgR-Mcm1 transcriptional complex, IPK2 activates genes responsive to arginine and phosphate disposition by the kinase-independent stabilization determinant for Mcm1 and ArgRI (El Bakkoury et al. 2000, El Alami et al. 2003, Bosch and Saiardi 2012). In higher plant Arabidopsis, there are two IPK2 genes, AtIPK2α and AtIPK2β (Stevenson-Paulik et al. 2002, Xia et al. 2003, Xu et al. 2005). Arabidopsis IPK2 plays important roles in diverse physiological processes, including root growth, axillary shoot branching, sexual reproduction and abiotic stress responses (Xu et al. 2005, Zhang et al. 2007, Yang et al. 2008, Zhan et al. 2015). As an auxin-responsive gene, over-expression of AtIPK2β results in an auxin-resistant phenotype. But the further mechanisms by which plant IPK2 participates in auxin signaling have not been revealed.
Rice inositol polyphosphate kinase gene (OsIPK2) has been previously isolated and identified as a candidate phytic acid biosynthetic gene (Suzuki et al. 2007). In this study, we report the role of OsIPK2 in auxin signaling and reveal the molecular mechanism underlying the degradation of OsIAA11, an Aux/IAA that is crucial for lateral root initiation. Our results indicate that OsIPK2 interacts with OsIAA11 to protect it from degradation and inhibit lateral root formation.
Results
OsIPK2 interacts with OsIAA11
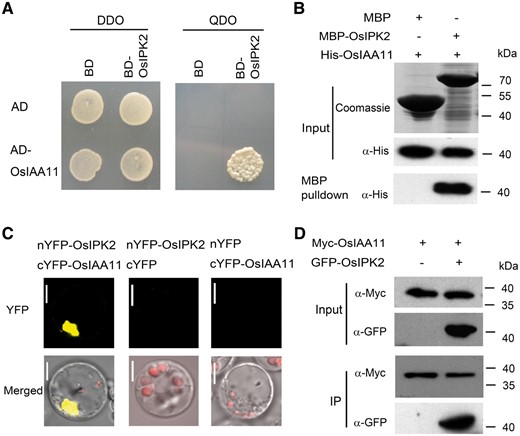
As a previous report showed that AtIPK2β is a nuclear protein and over-expression of AtIPK2β led to altered expression of auxin-related genes and an auxin-resistant phenotype, we considered the possibility that plant IPK2 physically interacts with auxin signaling components (Xia et al. 2003, Zhang et al. 2007). Yeast two-hybrid (Y2H) assay was performed to identify the potential OsIPK2-interactive proteins. The OsIPK2 protein was fused to the GAL4-DNA binding domain (BD) as bait. A series of rice Aux/IAAs were fused to the GAL4 activation domain (AD) and used as prey. Y2H results showed that OsIAA11 was a positive interactive protein of OsIPK2 (Fig. 1A). Because other members of the OsIAA11 clade did not have the capacity to bind OsIPK2 in yeast, it indicated the specificity of OsIAA11 protein for OsIPK2 (Supplementary Fig. S1). An in vitro pull-down experiment with recombinant His-tag fused OsIAA11 and MBP-tag fused OsIPK2 confirmed the Y2H observations, suggesting that OsIPK2 could directly interact with OsIAA11 (Fig. 1B). We also conducted bimolecular fluorescence complementation (BiFC) assays to verify the interaction. Yellow Fluorescent Protein (YFP) was split into N-terminal and C-terminal amino acid residues and fused with OsIPK2 and OsIAA11 respectively. Co-expression of the nYFP–OsIPK2 and cYFP–OsIAA11 fusion proteins in the rice protoplast produced an obvious fluorescence signal (Fig. 1C). As negative controls, co-expression of nYFP-OsIPK2/cYFP or nYFP/cYFP-OsIAA11 did not produce fluorescence signal in protoplasts. To further verify the interaction, we performed co-immunoprecipitation (Co-IP) assay using lysate of rice protoplasts co-expressing GFP-OsIPK2 and Myc-OsIAA11 fusion proteins. Co-IP result indicated that OsIPK2 could interact with OsIAA11 in vivo (Fig. 1D). Taken together, these observations demonstrated that OsIPK2 interacts with OsIAA11, suggesting that OsIPK2 is a potential regulator of auxin signaling.
OsIPK2 interacts with OsIAA11. (A) Yeast two-hybrid assays to detect the interaction between OsIPK2 and OsIAA11. OsIAA11 was used to fuse to GAL4 activation domain (AD) as prey and OsIPK2 was used to fuse to GAL4 DNA-binding domain (BD) as bait. The interactions were examined on the double dropout (DDO) medium (SD/-Leu/-Trp) and quaternary dropout (QDO) medium (SD/-Ade/-His/-Leu/-Trp). (B) In vitro pull-down assay of the OsIPK2-OsIAA11 interaction. Purified recombinant MBP-OsIPK2 or MBP proteins were used as bait to bind agarose beads and incubated with 1 μg His-OsIAA11 proteins. After wash, the input and pulled-down proteins were analyzed by immunoblotting using anti-His antibody. The MBP and MBP-OsIPK2 proteins were visualized on the Coomassie Blue-stained gel. (C) The bimolecular fluorescence complementation (BiFC) assays indicate OsIPK2 interacts with OsIAA11 in rice. nYFP-OsIPK2 and cYFP-OsIAA11 were co-expressed in rice protoplasts prepared from 12-day-old etiolated rice leaf. Scale bar = 5 μm. (D) OsIPK2 interacts with OsIAA11, as indicated by Co-immunoprecipitation (Co-IP) assays. DNAs of P35s-Myc-OsIAA11-Nos or together with P35s-GFP-OsIPK2-Nos were used to transfect rice protoplasts. Total proteins from protoplast extracts expressing GFP-OsIPK2/Myc-OsIAA11 or Myc-OsIAA11 alone were immunoprecipitated with an anti-GFP antibody.
Domain II is necessary for the OsIPK2-OsIAA11 interaction
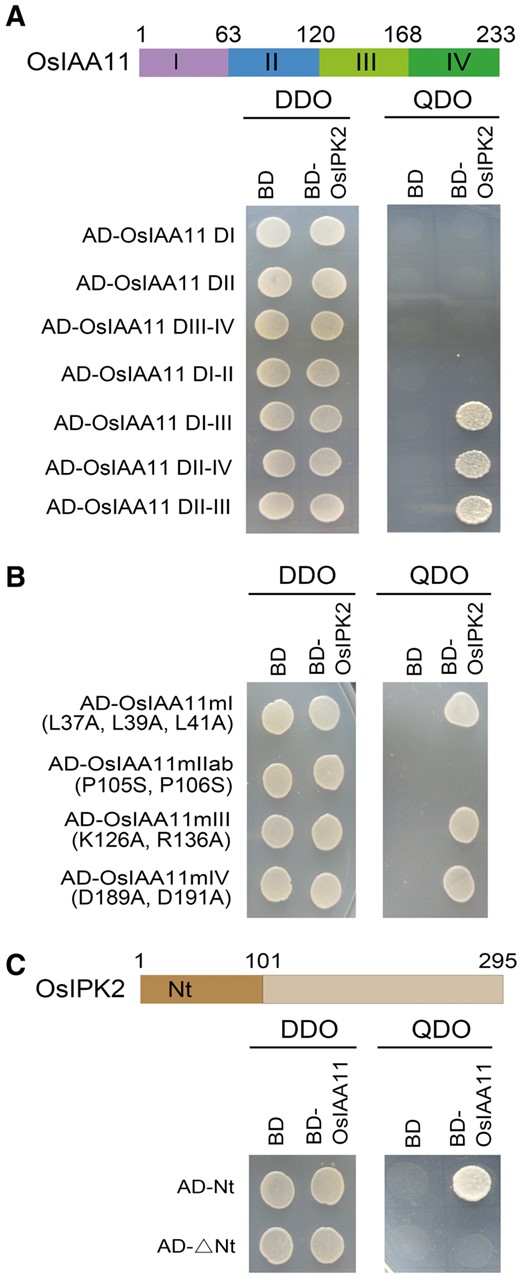
As a classic Aux/IAA protein, OsIAA11 has four highly conserved domains (DI, II, III, IV) . To determine which domain of OsIAA11 that mediates the interaction with OsIPK2, we fused seven OsIAA11 truncated variants to the AD domain of pGADT7 vector. Y2H assays showed that the DII-III region of OsIAA11 is required for the interaction with OsIPK2 (Fig. 2A).
DII is necessary for the OsIPK2-OsIAA11 interaction. (A) Determination of the functional domain of OsIAA11 for the interaction with OsIPK2 by yeast two-hybrid (Y2H) assays. The OsIAA11 protein contains four conserved domains, DI, DII, DIII and DIV. The truncations of OsIAA11 were used to fuse to GAL4 activation domain (AD) and OsIPK2 was used to fuse to GAL4 DNA-binding domain (BD). The interactions were examined on the DDO medium (SD/-Leu/-Trp) and QDO medium (SD/-Ade/-His/-Leu/-Trp). DII-DIII region of OsIAA11 is required for the interaction with OsIPK2 in yeast. (B) Determination of the amino acids of OsIAA11 that involved in the interaction with OsIPK2 by Y2H assays. The effects of point mutations in the core motifs of OsIAA11 were tested for interaction with OsIPK2. OsIAA11mI (L37A, L39A, L41A), OsIAA11mIIab (P105S, P106S), OsIAA11mIII (K126A, R136A) and OsIAA11mIV (D189A, D191A) were used as prey. OsIPK2 was used as bait. The interactions were examined on the DDO medium and QDO medium. Stabilized mutations in DII of OsIAA11 weakened the OsIPK2-OsIAA11 interaction. (C) Determination of the functional domain of OsIPK2 for the interaction with OsIAA11. OsIPK2 protein was splitted into the N-terminal 1-101 and the C-terminal fragments. The OsIPK2 truncations were used to fuse to GAL4 activation domain (AD) and OsIAA11 was used to fused to GAL4 DNA-binding domain (BD). The interactions were examined on the DDO and QDO mediums. OsIPK2-1-101 peptide is required for the interaction.
To define the amino acids of OsIAA11 that were involved in the interaction, we introduced point mutations into the core motifs of OsIAA11 by site-directed mutagenesis (Fig. 2B). An OsIAA11, in which three leucines at the core of the EAR domain were changed to alanines, was named OsIAA11mI (L37A, L39A, L41A). We also constituted OsIAA11mIIab (P105S, P106S) in which two proline residues at the degron region were mutated to serine residues. Besides, two OsIAA11 variants in which the substitutions of two central amino acid residues on the basic or acidic face of DIII-IV of OsIAA11 were introduced and named as OsIAA11mIII (K126A, R136A) and OsIAA11mIV (D189A, D191A) respectively (Dinesh et al. 2015). Y2H assays showed that the mutations in DI, DIII and DIV of OsIAA11 had no visible effect on the OsIPK2-OsIAA11 interaction in yeast. However, the substitutions of central amino acid residues in the degron severely weakened the interaction. These results suggested that the degron of OsIAA11 is critical for the interaction with OsIPK2 and other amino acid sequence outside the degron motif may also contribute to the specific interaction.
The coding region of OsIPK2 contains a N-terminal domain (Nt) and a C-terminal catalytic domain harboring sequences of inositol polyphosphate binding site and ATP binding site that are conserved in plant IPK2 (Bhati et al. 2014). To determine which region that regulates the interaction with OsIAA11, we fused the two truncated variants of OsIPK2 to the AD domain of pGADT7 vector. The full-length of OsIAA11 were fused to the BD domain of pGBDT7 vector. Y2H assays showed that the N-terminal amino acid sequence 1-101 of OsIPK2 itself could interact with OsIAA11 (Fig. 2C).
OsIPK2 can stabilize OsIAA11
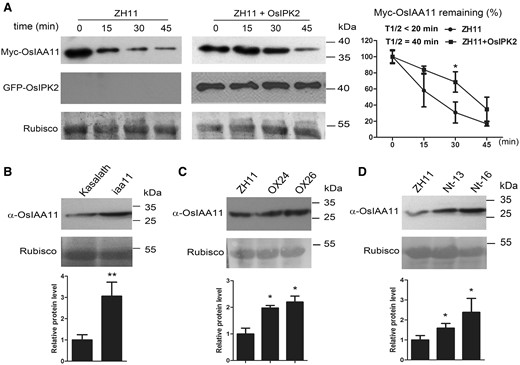
DII is a determinant of Aux/IAA stability and auxin response. We next developed a rice protoplast system to determine the biological significance of the physical interaction (Fig. 3A). Equal amounts of protoplasts prepared from the japonica rice ‘Zhonghua11’ (ZH11) transfected with the effector DNA of P35s-Myc-OsIAA11-Nos were used to express Myc-OsIAA11 fusion protein. Cycloheximide (CHX) was added to inhibit the newly synthesized OsIAA11 fusion protein. After 50 µM NAA incubation, total proteins were extracted at indicated time points. Western blot assays showed that the transiently expressed Myc-tagged OsIAA11 gradually degraded in rice protoplasts with auxin induction, while the co-expressed GFP-tagged OsIPK2 inhibited the degradation of Myc-tagged OsIAA11 proteins. The relative levels of remaining Myc-OsIAA11 were analyzed, showing that the half-life of OsIAA11 in rice cells was prolonged to two-fold of the control level by OsIPK2. The protein degradation assays indicated that OsIPK2 plays a negative role in the proteolysis of OsIAA11.
OsIPK2 affects the accumulation of OsIAA11 proteins in rice. (A) OsIPK2 inhibited the auxin-induced degradation of OsIAA11 in rice protoplasts. Protoplasts were prepared from 12-day-old etiolated wild type rice seedlings. ZH11, wild type. Equal amounts of protoplasts were transfected with 10 μg of P35s-Myc-OsIAA11-Nos DNA or together with 5 μg P35s-GFP-OsIPK2-Nos DNA for transiently expression. After 12 h, 20 μM cycloheximide (CHX) was added to inhibit the newly OsIAA11 fusion protein synthetic. 50 μM NAA was added to trigger the degradation of Myc-OsIAA11. Samples were cultured in 26°C/Dark and harvested at the indicated time points. Protein extracts were analyzed by immunoblotting using anti-Myc antibody. The PVDF membranes were stained with Ponceau to visualize the large subunit of Rubisco as loading control of total proteins. Quantitative analysis of the relative Myc-OsIAA11 level is presented on the right of the blots. The average values were obtained from three independent experiments. Error bar = SD. Asterisks indicate significant differences by Student’s t-test (*P < 0.05). (B, C, D) Immunoblotting analysis of the accumulation of OsIAA11 proteins in the wild-type (Kasalath or ZH11), iaa11 mutant (B), 35S:OsIPK2-GFP (C) and 35S:OsIPK2 Nt-GFP transgenic plants (D). The total proteins were extracted and the abundance of OsIAA11 proteins was detected by western blotting using anti-OsIAA11 antibody. The PVDF membranes were stained with Ponceau to visualize the large subunit of Rubisco as loading control of total proteins. Quantitative analysis of the relative OsIAA11 level is presented below the blots. The average values were obtained from three independent experiments. Error bars represent the SD. Asterisks indicate significant differences by Student’s t-test (*P < 0.05, **P < 0.01).
An OsIAA11 monoclonal antibody was used to detect the protein level of OsIAA11 in rice (Fig. 3B). Western blot assays showed that the point mutation in the degron of OsIAA11 (P106L) led to enhanced accumulation of OsIAA11 proteins in iaa11 mutant rice, which is consistent with previous report (Jing et al. 2015). To further explore the importance of OsIPK2 or OsIPK2-1-101 peptide for auxin signaling, we developed rice transformants constitutively expressing OsIPK2-GFP or OsIPK2 Nt-GFP fusion protein by Agrobacterium-mediated transformation (Supplementary Fig. S2). Homozygous T2 seeds of transgenic lines were used for further analysis. The relative OsIAA11 protein levels in 35S:OsIPK2-GFP lines were enhanced compared with wild type (WT) (Fig. 3C). Moreover, the accumulation of OsIAA11 proteins in transgenic plants expressing OsIPK2 Nt-GFP fusion protein was also increased compared with WT, indicating that OsIPK2-1-101 peptide could stabilize OsIAA11 in rice (Fig. 3D). Taken together, these observations indicate that OsIPK2 acts as a co-factor to protect OsIAA11 from degradation.
OsIPK2 inhibits lateral root formation and auxin response
Previous study showed that the mRNA level of OsIAA11 did not respond to exogenous auxin induction (Jain et al. 2006). To investigate whether auxin affects OsIPK2 transcription, we also determined the expression of OsIPK2 under the treatment of IAA (Supplementary Fig. S3A). A time-course analysis showed that the OsIPK2 transcripts were reduced 50% within 30 min of IAA application and reached the bottom after 3 h treatment, indicating that OsIPK2 was negatively regulated by auxin. Quantitative RT-PCR assays also showed that the expression of OsIAA11 was significantly up-regulated in iaa11 mutant, 35S:OsIPK2-GFP and 35S:OsIPK2 Nt-GFP transgenic lines compared with WT (Kasalath or ZH11), indicating that the transcription of OsIAA11 is correlated with its stability in rice (Supplementary Fig. S3B).
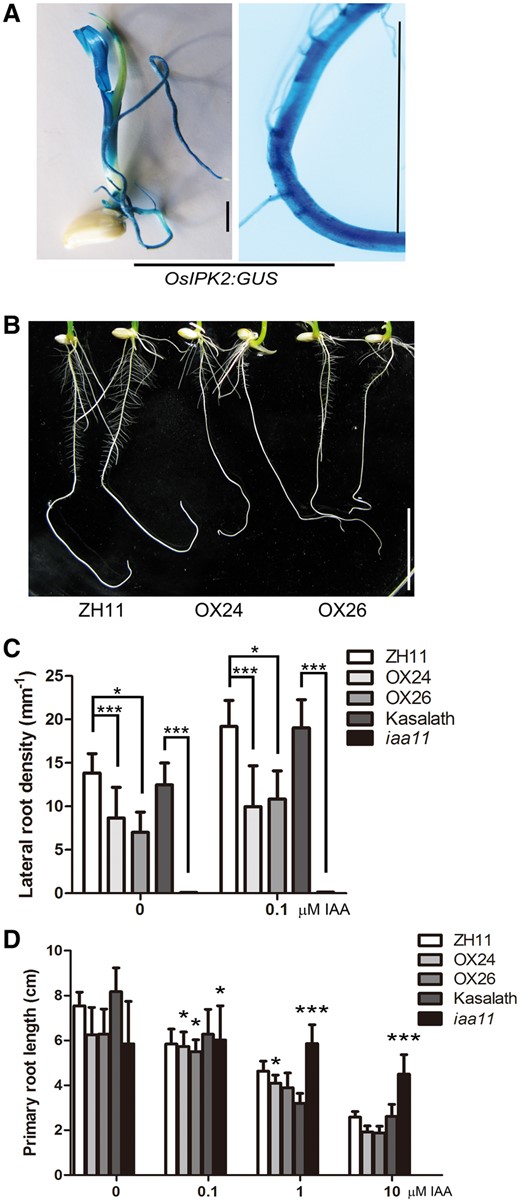
Functional interaction of OsIPK2 and OsIAA11 implies co-localization. OsIAA11 is mainly expressed in root, shoot base and etiolated shoot (Zhu et al. 2012). Previous quantitative RT-PCR analysis showed that OsIPK2 was expressed in the root, shoot, flower, anther and embryo (Suzuki et al. 2007). The staining assays of ProOsIPK2:GUS (β-glucuronidase) transgenic plants revealed that OsIPK2 was highly expressed in the primary root of 5-day-old rice seedlings, especially in the stele and lateral root initiation sites (Fig. 4A). The overlapped expression pattern of OsIPK2 and OsIAA11 in root implies the related physiological function of OsIPK2-OsIAA11 interaction.
35S:OsIPK2-GFP transgenic plant seedlings show defects in lateral root formation. (A) GUS staining assays of ProOsIPK2: GUS transgenic plants. GUS signals in 5-day-old rice seedling (left) and root (right). Scale bar = 5 mm. (B) Roots of ZH11 (left, wild type) and 35S:OsIPK2-GFP transgenic plant seedlings (right, OX24 and OX26) grown on 1/2 MS medium for 7 days. Scale bar = 2 cm. (C) The lateral root densities of WT (ZH11, Kasalath), 35S:OsIPK2-GFP transgenic lines and iaa11 mutant seedlings. Seeds were surface-sterilized and planted on MS medium for 2-day germination, then transferred onto medium without or with 0.1 µM IAA for another 5 days. Lateral root density is the number of lateral roots per linear mm primary root. There are significant differences between transgenic lines, iaa11 mutant and WT seedlings by Student's t-test (*P < 0.05, ***P < 0.001). (D) The response of primary root elongation of WT (ZH11, Kasalath), OX lines and iaa11 mutant to exogenous IAA (0–10 μM). The treatment × genotype interaction effect (two-way ANOVA) is indicated (*P < 0.05, ***P < 0.001). All means were shown with SD (n>15). These experiment were repeated three times with similar results.
The stability of Aux/IAA is essential for root development and auxin response. Complete stabilization of OsIAA11 impaired the initiation of lateral root primordia and retarded plant growth in iaa11 mutant rice (Zhu et al. 2012) (Supplementary Fig. S4). To determine the physiological function of OsIPK2, we further analyzed the root phenotypes of OsIPK2 over-expression transgenic plants. After grown on 1/2 Murashige-Skoog (MS) medium for 7 days, the primary roots of wild-type (ZH11) seedlings produced plenty of lateral roots, while the lateral root numbers of OX24 and OX26 lines were dramatically reduced compared with the WT (Fig. 4B). The lateral root densities of over-expression transgenic lines were also less than that of WT, suggesting that OsIPK2 altered auxin sensitivity of transgenic plants (Fig. 4C). To clarify this, we examined the effects of IAA on the root growth of WT and transgenic plants in Fig. 4D. Seeds were germinated on the IAA-free medium for two days, and then transferred to IAA-containing medium for another five days. Two-way Anova analysis was performed for comparison of the auxin response between WT and transgenic lines. Although the primary roots of OsIPK2 over-expression transgenic lines were shorter than WT at normal condition, over-expression plants were more resistant to exogenous IAA in the root growth inhibition assay. Besides, our observation also showed that 0.1 μM IAA reduced the primary root length of WT and transgenic plants to a similar extent and induced more lateral root formation. However, the lateral root defect of iaa11 mutant could not be reversed. These data suggested that the auxin sensitivities of OX24 and OX26 roots were attenuated but less inhibited than iaa11 mutant. Moreover, the aerial parts of over-expression plants also displayed a dwarf phenotype with decreased cell size in the shoot base, indicating that OsIPK2 suppressed the cell expansion in transgenic rice (Supplementary Fig. S5). Taken together, OsIPK2 over-expression plants displayed a phenotype similar to iaa11 mutant, but to a lesser extent.
Data above indicate that over-expression of OsIPK2 inhibits lateral root formation and auxin response. To determine the effects of OsIPK2 knockdown on rice development, transgenic rice plants expressing 230 bp double-strand RNA of OsIPK2 gene (191 bp–420bp) were produced (Supplementary Fig. S6A, B). Compared with the WT, qRT-PCR assays showed that the OsIPK2 transcripts in RNAi transgenic plants were reduced 30%∼50%, while the expression of OsIAA11 was reduced 50%∼70% in RNAi lines (Supplementary Fig. S6C). The relative protein level of OsIAA11 showed no significant difference between WT and RNAi lines (Supplementary Fig. S6D). Combined with these observations, phenotypic analysis showed that 10-day-old OsIPK2 RNAi transgenic plant seedlings failed to produce more lateral roots compared with WT plants (Supplementary Fig. S6E). The result is consistent with previous report that the OsIAA11 RNAi transgenic rice has similar number of lateral roots with WT (Jing et al. 2015). Because of the high redundancy among members of Aux/IAA family, loss of single Aux/IAA function may cause subtle phenotype (Overvoorde et al. 2005). Furthermore, the roots of 10-day-old RNAi plants exhibited hypersensitivity to increasing concentrations of exogenous IAA, indicating that the auxin response in RNAi plants was enhanced by down-regulation of OsIPK2 (Supplementary Fig. S6F). Taken together, these data suggested that OsIPK2 is involved in auxin response and plays a negative role in lateral root formation.
N-terminal region of OsIPK2 inhibits lateral root formation
The physiological function of OsIPK2 may depend on its interaction with OsIAA11 in rice. Because amino acids 1-101 of OsIPK2 could interact with OsIAA11 and enhance OsIAA11 level in rice, we explored whether OsIPK2 fragment 1-101 itself could function as the full-length OsIPK2 in plant growth and development. After growing on IAA-free medium for seven days, the lateral roots which emerged from the primary roots of 35S:OsIPK2 Nt-GFP seedlings were significantly reduced compared with WT, indicating that auxin response was altered in transgenic plants (Fig. 5A, B). Moreover, the expression of Nt-GFP fusion protein in transgenic rice caused shorter primary roots compared with the WT. The root growth inhibition assays showed that 35S:OsIPK2 Nt-GFP seedlings had reduced sensitivity to IAA but more lateral roots (Fig. 5C). Taken together, our data suggested that the amino terminus of OsIPK2, which is responsible for physical interaction with OsIAA11, inhibited lateral root formation and auxin response.
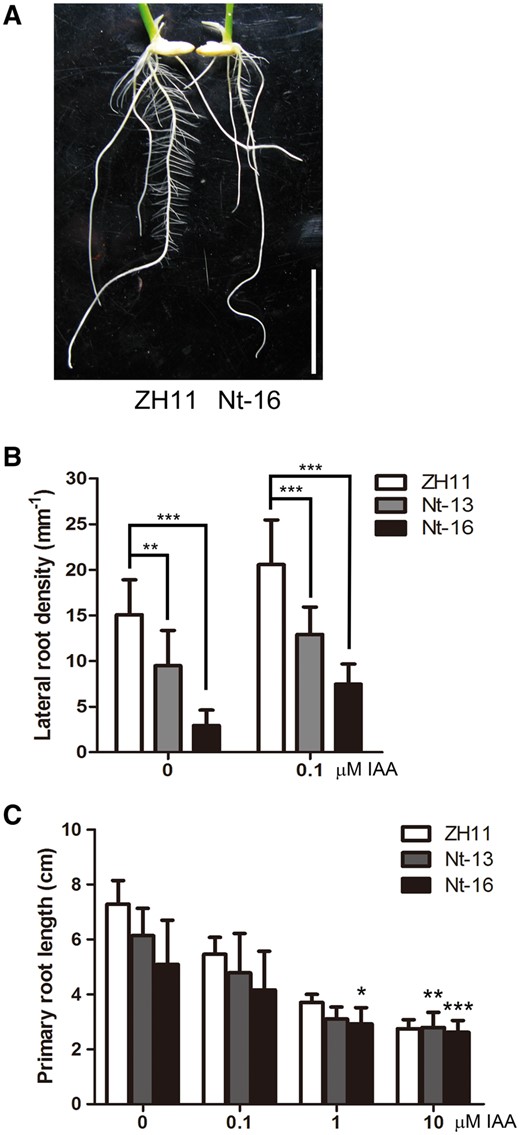
Root phenotypes of ZH11 and 35S:OsIPK2 Nt-GFP transgenic seedlings. (A) Roots of 7-day-old ZH11 (wild type) and 35S:OsIPK2 Nt-GFP transgenic plant seedlings. Scale bar = 2 cm. (B) The lateral root densities of ZH11 and 35 S:OsIPK2 Nt-GFP transgenic seedlings. Seeds were surface-sterilized and planted on MS medium for 2-day germination, then transferred onto medium without or with 0.1 μM) IAA for another 5 days. There are significant differences between transgenic lines and wild-type plants by Student’s t-test (**P < 0.01, ***P < 0.001). Error bars represent the SD (n > 20). (C) The response of primary root elongation of ZH11 and 35S:OsIPK2 Nt-GFP transgenic lines to exogenous IAA (0–10 μM). Root lengths of 7-day-old ZH11 and transgenic plants grown on 1/2 MS medium containing indicated concentration of IAA were measured. The treatment × genotype interaction effect (two-way ANOVA) is indicated (*P < 0.05, **P < 0.01, ***P < 0.001). All means are shown with SD (n > 15). These experiments were repeated three times with similar results.
Discussion
The degradation of Aux/IAA repressors is an essential step in auxin signaling. Plant has developed a SCF-ubiquitin-dependent system to degrade Aux/IAA in an auxin-dependent manner. Although Aux/IAA family proteins are highly redundant, they have different affinity to TIR1 and diversified degradation rate (Villalobos et al. 2012; Shimizu-Mitao and Kakimoto 2014). To provoke various auxin responses by the short-lived Aux/IAA throughout plant growth and development, some regulators have been found to add opportunity to regulate the degradation of Aux/IAA. For example, the cis/tans isomerization of degron motif catalyzed by LRT2/OsCYP2 intensifies the Aux/IAA-OsTIR1 interaction and promoted the degradation of OsIAA11 (Jing et al. 2015). In this study, we have linked rice inositol polyphosphate kinase (OsIPK2) to the auxin-mediated degradation of Aux/IAA by demonstrating that OsIPK2 is an OsIAA11-interacting protein (Fig. 1). Because of the importance of Aux/IAA repressors, OsIPK2 is a key regulator in auxin signaling.
As a cyclophilin-type peptidyl-prolyl cis/trans isomerase, OsCYP2 has been reported to facilitate the proteasomal degradation of multiple Aux/IAA proteins by catalyzing the conformation changes. However, OsIAA11 was the only Aux/IAA protein in this clade could bind OsIPK2 in yeast. Although the core amino acid sequences of the four domains are highly conserved among Aux/IAA family proteins, the overall identities of full-length Aux/IAA proteins range from 14% to 76% (Jain et al. 2006). The unique amino acid sequence in DII and DIII may lead to the special interaction between OsIPK2 and OsIAA11 (Fig. 2). Moreover, the mutations of conserved proline residues in the degron weakened the OsIPK2-OsIAA11 interaction, indicating that the DII of OsIAA11 can associate with multiple co-factors in auxin signaling. Our data presented in Fig. 3 showed that OsIPK2 plays an opposite role against the auxin signal in the proteolysis of OsIAA11. The sub-cellular interaction of OsIPK2-OsIAA11 occurs in the nucleus (Supplementary Fig. S7). OsIPK2 may protect OsIAA11 from degradation by interfering the interaction between OsTIR1 and OsIAA11. It suggested that the regulation of Aux/IAA degradation is more flexible and complex than we expected.
Phenotypic analysis of transgenic Arabidopsis expressing the degradation rate variants of IAA14 indicated that the timing of lateral root initiation and auxin sensitivity is plastic and can be tuned (Guseman et al. 2015). Over-accumulation of certain Aux/IAA or additional co-factor could interfere with the sequential degradation of co-expressed Aux/IAA proteins and proper auxin response (Pierre-Jerome et al. 2014). OsIAA11 is the prominent Aux/IAA gene functioning in rice lateral root development. Previous reports showed that the stabilized mutation in the degron of OsIAA11 strictly blocks lateral root initiation (Zhu et al. 2012). Loss-of-function mutation of CYP2 resulted in reduced degradation rate of OsIAA11 and defective development of lateral root, a phenotype which is similar to iaa11 mutant (Zhu et al. 2012; Kang et al. 2013; Jing et al. 2015). In this study, our results reveal a role of OsIPK2 in lateral root development. OsIPK2 over-expression transgenic plants exhibited reduced lateral root numbers and decreased sensitivity to IAA (Fig. 4). In contrast, RNAi plants had increased sensitivity to IAA (Supplementary Fig. S6). These data indicated that OsIPK2 is involved in auxin-mediated plant growth and development. Moreover, we also showed that the N-terminal region of OsIPK2 alone could bind OsIAA11, which led to increased OsIAA11 proteins and reduced lateral root density in transgenic rice (Fig. 5). These data suggested that the protein–protein binding function of OsIPK2 is important for the stability of OsIAA11 and lateral root development. Besides, IPK2 is also a rate-limiting enzyme for IP6 biosynthesis. As the major phosphorus storage form in plant seeds, IP6 also functions as a co-factor of TIR1 auxin receptor complex and plays important roles in plant development (Tan et al. 2007). Low phytic acid mutants (lpa), atipk1 and atmrp5 have been reported to exhibit an enhanced growth of lateral roots, suggesting that the inhibited IP6 biosynthesis may alter auxin signaling, which in turn affects lateral root formation (Kuo et al. 2014). It seems that the reduced lateral root formation in transgenic plants was not only caused by OsIPK2-induced stabilization of OsIAA11.
In addition, it is important to notice that over-expression of OsIPK2-GFP or OsIPK2 Nt-GFP also increases the transcription level of OsIAA11 mRNA at 1.5–3 fold, indicating the possibility that the enhanced transcription of OsIAA11 mRNA contributes to the accumulation of OsIAA11 proteins in over-expression lines (Supplementary Fig. S3B). As transcriptional repressor, Aux/IAA protein may regulate its own expression. Quantitative RT-PCR assays showed that OsIAA11 was significantly up-regulated in iaa11 gain-of-function mutant (Supplementary Fig. S3B). Because OsIPK2 is an OsIAA11-interactive protein and the DIII of OsIAA11 is also necessary for the interaction, it is not surprising that the expression of OsIAA11 in 35S:OsIPK2-GFP and 35S:OsIPK2 Nt-GFP transgenic lines was affected (Supplementary Fig. S3B). It seems that the transcriptional regulation is a more efficient way to affect the accumulation of OsIAA11. However, over-expression of wild-type Aux/IAA genes have subtle or no discernible phenotypic changes in previous reports, whereas expression of dominant mutants resulted in diverse auxin-related phenotypes (Sato and Yamamoto 2008). So OsIPK2-induced stabilization and transcription of OsIAA11 may both contribute to the accumulation of OsIAA11 and its function in rice development.
In conclusion, the functional interaction between OsIPK2 and OsIAA11 presented here provides a novel mechanism, showing that OsIPK2 participates in auxin-regulated rice development by stabilizing OsIAA11. Potential dynamic protein-protein interactions among the core auxin signaling components may contribute to the precise regulation of the turnover of OsIAA11 in response to a variety of cellular signals.
Materials and Methods
Materials and growth conditions
All transgenic plants in this study are in the rice variety Zhonghua11 (Oryza sativa L. ssp. japonica). iaa11 mutant rice kindly provided by Prof. Chuanzao Mao is in the rice variety Kasalath (Oryza sativa L. ssp. Indica). For phenotypic analysis, rice seeds were surface-sterilized and sown on 1/2 MS medium containing 0.8% agar and 3% (w/v) sucrose under a 14-h light/10-h dark photoperiod at 28°C/24°C. For breeding, rice plants were grown in field during summer or in a greenhouse during winter.
Constructs and rice transformation
To generate the ProOsIPK2:GUS construct, a 1.8-kb OsIPK2 (AK072296) promoter region was amplified from rice genomic DNA and inserted into the binary vector pCAMBIA1391Z (Cambia, Australia) at the PstI/BamHI sites to guide a GUS gene.
To construct P35s:OsIPK2-GFP and P35s:OsIPK2 Nt-GFP vectors, the full-length coding sequence or N-terminal sequence of OsIPK2 was amplified from genomic DNA. Then the DNA fragments were inserted into the NcoI/SpeI sites of pCAMBIA1302 (Cambia, Australia) under the control of a cauliflower mosaic virus (CaMV) 35S promoter.
To construct RNAi vector, the forward and reverse sequences of a 230-bp cDNA fragment were amplified and inserted into the SpeI/EcoRI and BamHI/NcoI sites of the intermediate vector pBJWI3 respectively (Cui et al. 2010). Then the whole fragment with the intron sequence was inserted into BamHI/SacI sites of the transformation vector PU1301 under control of a maize (Zea mays) ubiquitin promoter (Qiu et al. 2007). Primer sequences used for these constructs are provided in Supplementary Table S1.
Vectors were transformed into the callus of rice variety Zhonghua11 (Oryza sativa ssp. japonica) by the Agrobacterium (strain EHA105)-mediated transformation method (Hiei et al. 1994). Generally, the rice seeds were surface-sterilized and used to induce for embryonic calli on 2N6 medium. Transformed calli were selected on medium containing 25 mg/L hygromycin B, 0.625 mg/L 2,4-dichlorophenoxy and 400 ppm ampicillin. Homozygous T2 transgenic plants were used for further analysis.
Yeast two-hybrid assay
Yeast two-hybrid experiments were performed according to the manufacturer’s instructions (Clontech). To introducing point mutations into OsIAA11 (LOC_Os03g43400), PCR-based site-directed mutagenesis was performed using complementary primer pairs containing mutation sites of OsIAA11mI, OsIAA11mIIab, OsIAA11mIII, OsIAA11mIV. Primer sequences used for Y2H constructs are provided in Supplementary Table S1. Each bait/prey pair was co-transformed into the yeast strain AH109 (Clontech) and cultured on SD medium minus Leu and Trp (SD/-L/-T) at 30°C for 3 days. Interactions were tested on SD medium minus Leu, Trp, His and Ade (SD/-L/-T/-H/-M).
Pull-down assay
The coding region of OsIAA11 was amplified and cloned into the NcoI/BamHI sites of pET32a vector. The coding sequence of OsIPK2 was amplified and cloned into the EcoRI/SalI sites of pMAL-C2x vector. The recombinant proteins were expressed in BL21(ED3) induced by IPTG at 22°C. Primer sequences used here are provided in Supplementary Table S1.
For in vitro MBP-pulldown assay, 1 μg His-OsIAA11 proteins were incubated with 2 μg MBP-OsIPK2 fusion or MBP protein conjugated agarose beads (Cell Signaling Technology) and cultured at 4°C for 4 h, then washed five times with 500 μl wash buffer (50 mM Tris-HCl pH = 7.4, 400 mM NaCl, and 0.2% NP-40). Beads were re-suspended, de-naturated, separated on 10% SDS/PAGE and then analyzed by immunoblotting. The separated proteins were blotted onto the PVDF membrane and blocked with fat-free milk. The mouse polyclone anti-His antibody (Cell Signaling Technology) was used to detect His-OsIAA11 proteins. A 2000-fold dilution of secondary antibody conjugated with horseradish peroxidase (HRP) was then used. HRP signals were enhanced with a SuperSignal West Pico Chemiluminescent Substrate Kit (Thermo SCIENTIFIC).
Protoplasts isolation, transfection and BiFC assay
The protoplasts were isolated from 12-day-old etiolated rice seedlings as previously described with minor modifications (Zhang et al. 2011). The rice tissues were immersed into 10 ml enzyme solution (0.6 M mannitol, 10 mM MES, 1.5% cellulose R-10, 0.75% macerozyme R-10, 0.1% BSA, 1 mM CaCl2, 0.25 g/ml carbenicillin) and shaked at 80 rpm for 4 h. The mixture was filtered through two layers of 40 μm nylon meshes and washed twice with 10 ml W5 solution (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 5 mM glucose, 2 mM MES, pH to 5.8). The protoplasts were collected by centrifuging at 100 g for 2 min, and re-suspended with 1 ml MMG solution (0.6 M mannitol, 15 mM MgCl2, 4 mM MES).
For transiently expressing target proteins, the full-length CDS of OsIPK2 and OsIAA11 were amplified and fused to the carboxyl terminal of YFP variants by inserting into XbaI/KpnI sites of P35s-YFPn-Nos or P35s-YFPc-Nos vectors. Primers used in these experiments are listed in Supplementary Table S1. DNAs including the CaMV 35S promoter, coding sequence and a terminator used for protoplast transfection were amplified with primer pair 5′-GCACGACAGGTTTCCCGACT-3′/ 5′-GAAAGGGGGATGTGCTGCAA-3′ as previously described (Lu et al. 2013). For PEG-mediated transfection, 10–15 μg purified DNA fragments of various constructs, 100 μl prepared rice protoplast suspension liquid and 110 μl PEG solution (40% PEG4000, 0.6 M Mannitol, 100 mM CaCl2) were mixed thoroughly and cultured at 26°C for 10–15 min in dark. Then 440 μl W5 solution was added to stop the transfection. After being washed twice, the solution were re-suspended with WI solution (0.5 M Mannitol, 4 mM MES, 20 mM KCl) and cultured in the dark at 26°C for 8–12 h. Protoplasts were collected and checked for the presence of fluorescence with a confocal microscopy (Olympus FV 1000). To determine the sub-cellular interaction part, 1 μg/ml DAPI was added into the rice protoplast suspension liquid for the nuclear staining. After 10 min staining, the GFP and DAPI signals were observed and photographed using an Olympus IX71 microscope with an Olympus DP72 CCD camera.
Coimmunoprecipitation assay
For the construction of plasmids expressing Myc-OsIAA11 or GFP-OsIPK2 fusion protein, the coding region of OsIAA11 or OsIPK2 was inserted into the KpnI/BamHI sites of P35s-MCS-Nos vector, subsequently either the DNA sequence of Myc-tag or GFP-tag was fused to the amino terminal of target protein by inserting into the XbaI/KpnI sites. Primers used in these experiments are listed in Supplementary Table S1.
The co-immunoprecipitation assays were performed as described with some modification (Xu et al. 2014). The lysates of rice protoplasts expressing Myc-OsIAA11, or co-expressing Myc-OsIAA11 and GFP-OsIPK2 were extracted with the extraction buffer (1 × PBS pH = 7.4, Roche complete protease inhibitor cocktail, 1% Triton X-100). After centrifugation twice at 14,000 g for 15 min, the supernatant was collected. About 1–2 mg total proteins were used to incubate with 0.1% v/v anti-GFP antibody (Santa Cruz Biotechnology, Mouse) at 4°C for 4–8 h, then 5 μl Protein A magnetic beads (NEB) were added and incubated for another 2–4 h. After washed with 500 μl binding buffer (1 × PBS PH = 7.2, 0.1% Triton X-100) according to the procedure, the proteins binding to the magnetic beads were eluted by incubating at 70°C for 5 min and analyzed by Western blotting with anti-Myc antibody (Cell signaling Technology, Rabbit).
Protein degradation assay
The protein turnover assays were performed as described with some modifications (Maraschin et al. 2009). Equal amounts of protoplasts were transfected with 10 μg of P35s-Myc-OsIAA11-Nos DNA for transient expression and cultured in 26°C/Dark for 12 h. To analyze the effect of OsIPK2 on the degradation rate of OsIAA11, 5 μg DNA of P35s-GFP-OsIPK2-Nos was co-transfected with 10 μg DNA of P35s-Myc-OsIAA11-Nos into the rice protoplasts. The protoplast suspension liquid expressing Myc-tagged OsIAA11 or together with GFP-tagged OsIPK2 was incubated with 50 μM NAA and 20 μM cycloheximide (CHX), then immediately collected and resuspended by cold extraction buffer (PBS, 1 × Roche complete protease inhibitor cocktail, 1% Triton X-100) at the indicated time points. The total proteins were extracted by centrifugation at 14,000 g for 15 min and then analyzed by immunoblotting. The large subunit of Rubisco protein was used as loading control. Target bands were quantified with ImageJ software (http://rsb.info.nih.gov/ij/) and normalized against the Rubsico protein level. The mean values of three independent experiments were presented with statistical analysis of significant differences (Student’s t-test).
Antibody preparation and OsIAA11 protein level assay
The preparation of anti-OsIAA11 antibody was performed as previous described (Jing et al. 2015). The PET32a-OsIAA11 plasmid was used to produce recombinant protein tagged with 6×His. The purified 6×His-tagged OsIAA11 proteins were used to immunize mice. Total proteins were extracted from rice seedlings and grinded in liquid nitrogen, then were extracted with extraction buffer (50 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% Nonidet P-40 and 1 mM phenylmethylsulfonyl fluoride). After centrifugation twice at 14,000 g for 15 min, the supernatant was mixed with sample buffer and separated with 15% SDS-PAGE gel. The separated proteins were blotted onto the PVDF membrane and blocked with fat-free milk. The mouse polyclone anti-OsIAA11 antibody (1:5,000 dilution) was used to detect OsIAA11 proteins. A 2,000-fold dilution of secondary antibody conjugated with HRP was then used. HRP signals were enhanced with a SuperSignal West Pico Chemiluminescent Substrate Kit (Thermo SCIENTIFIC). The large subunit of Rubisco protein was used as loading control. Target bands were quantified with ImageJ software (http://rsb.info.nih.gov/ij/) and normalized against the Rubsico protein level.
Quantitative RT-PCR
For gene expression analysis, the total RNA was extracted from WT or transgenic plants with the TRIZOL reagent (Invitrogen). The DNA was removed from the RNA extractions with the DNase I (Fermentas). The first-strand cDNAs were synthesized with M-MLV Reverse Transcriptase reagent (Fermentas). qRT-PCR was carried out using SYBR Green Master Mix (TOYOBO) on a StepOnePlus real-time PCR System (Applied Biosystems) following the instructions. Primers were annealed at 56°C. Ub5 (AK061988) was used as an internal control. The primers used in these experiments are listed in Supplementary Table S1.
Analysis of GUS activity
Samples from five-day-old ProOsIPK2:GUS transgenic plant seedlings were used for histochemical staining. The transgenic plants were immersed into GUS staining solution (100 mM phosphate buffer, 10 mM EDTA, 1 mg/ml X-Gluc, 0.2% Triton X-100) and stained for GUS activity. After staining overnight, the samples were cleared in 50%∼90% (v/v) ethanol:water mixtures. Microscopic analysis was performed using an Olympus SZX16 microscope with bright-field optic. Images were photographed using a Canon PowerShot G10 camera (Fig. 4A, left) or Olympus DP72 CCD camera (Fig. 4A, right).
Root growth and auxin sensitivity
For root growth assays, two-day-old WT (ZH11 or Kasalath), transgenic lines and iaa11 mutant rice seedlings were transferred to 1/2 MS medium with indicated concentration of IAA (0, 0.1, 1, 10 μM) and incubated for additional 5–8 days. These seedlings were photographed with a Canon PowerShot G10 camera and processed using Adobe Photoshop (Adobe Systems). The root lengths of seedlings were measured and the numbers of lateral roots were calculated. To determine the lateral root densities, the mean values were presented with statistical analysis of significant differences (Student’s t-test). To determine how the root growth of WT (ZH11, Kasalath), transgenic lines and iaa11 mutant is affected by exogenous IAA, statistical analysis was carried out using GraphPad Prism (GraphPad Software, CA, USA). The mean values were presented with significant differences (two-way ANOVA).
Funding
This work was supported by grants from the National Natural Science Foundation of China (31170270).
Abbreviations
- AD
GAL4 activation domain
- ARF
auxin response factor
- Aux/IAA
Auxin/Indole-3-acetic acid
- BD
DNA-binding domain
- BiFC
bimolecular fluorescence complementation
- CaMV
cauliflower mosaic virus
- CHX
cycloheximide
- Co-IP
co-immunoprecipitation
- DI-IV
domain I-IV
- GUS
β-glucuronidase
- HRP
horseradish peroxidase
- IP3
inositol 1,4,5-triphosphate
- IPK2
inositol polyphosphate kinase
- MS
Murashige-Skoog
- mTOR
mammalian target of rapamycin
- Nt
N-terminal domain
- TPL
TOPLESS
- WT
wild type
- Y2H
yeast two-hybrid
- YFP
Yellow Fluorescent Protein
- ZH11
Zhonghua11
Acknowledgments
We sincerely thank Prof. Yunde Zhao (University of California, San Diego/Huazhong Agricultural University) and Prof. Huijun Xia (Wuhan University) for critical reading of the manuscript and valuable comments; Prof. Chuanzao Mao (Zhejiang University) for support of the iaa11 mutant seeds; Prof. Yongjun Lin (Huazhong Agricultural University) and Yue Li for rice transformation; Prof. Huijun Xia and State Key Laboratory of Hybrid Rice (Wuhan University) for reagents and analytical equipments.
Disclosures
The authors have no conflicts of interest to declare.