-
PDF
- Split View
-
Views
-
Cite
Cite
Gea Guerriero, Kjell Sergeant, Jean-Francois Hausman, Wood biosynthesis and typologies: a molecular rhapsody, Tree Physiology, Volume 34, Issue 8, August 2014, Pages 839–855, https://doi.org/10.1093/treephys/tpu031
Close - Share Icon Share
Abstract
Wood represents one of the most important renewable commodities for humanity and plays a crucial role in terrestrial ecosystem carbon-cycling. Wood formation is the result of a multitude of events that require the concerted action of endogenous and exogenous factors under the influence of photoperiod, for instance genes and plant growth regulators. Beyond providing mechanical support and being responsible for the increase in stem radial diameter, woody tissues constitute the vascular system of trees and are capable of reacting to environmental stimuli, and as such are therefore quite plastic and responsive. Despite the ecological and economic importance of wood, not all aspects of its formation have been unveiled. Many gaps in our knowledge are still present, which hinder the maximal exploitation of this precious bioresource. This review aims at surveying the current knowledge of wood formation and the available molecular data addressing the relationship between wood production and environmental factors, which have crucial influences on the rhythmic regulation of cambial activity and exert profound effects on tree stem growth, wood yield and properties. We will here go beyond wood sensu stricto, i.e., secondary xylem, and extend our survey to other tissues, namely vascular cambium, phloem and fibres. The purpose is to provide the reader with an overview of the complexity of the topic and to highlight the importance of progressing in the future towards an integrated knowledge on the subject.
Introduction
The term ‘wood’ is usually used to describe secondary xylem, i.e., a specific tissue of the plant vascular system composed of cellulose (50%), lignins, hemicellulose (25% each), proteins and phenolics (Déjardin et al. 2010). The ability to synthesize wood marked an important phase in the evolutionary history of Earth's ecosystems, as it allowed plants to conquer land and therefore to form the first stratified communities (i.e., forests) (Rothwell and Lev-Yadun 2005). The first vascular plants, which arose during the early Silurian, possessed only primary xylem, then later, in the Middle Devonian (∼370–380 million years ago), Progymnosperms appeared, displaying the first record of secondary xylem (Wight and Beck 1984, Rothwell and Lev-Yadun 2005), where wood played both a supportive and conductive role (Meyer-Berthaud 2000).
Wood is a natural, renewable resource that has undoubtedly played a crucial role in shaping human history and determining the rise of past great civilizations. Besides providing precious raw material, e.g., lumber for furniture or pulp for the paper industry, it is considered an environmentally friendly alternative to the use of fossil fuels (Plomion et al. 2001, Demura and Fukuda 2007), an inspiration for the development of biomimetic and eco-friendly materials (Ball 2005, Tampieri et al. 2009), and, last but not least, it is one of the major carbon (C)-reservoirs of terrestrial ecosystems.
Wood biosynthesis represents one of the most complex yet fascinating physiological processes, the complete understanding of which can unlock strategies for the full exploitation of woody biomass. Many aspects behind plant lignocellulosic biomass biosynthesis are indeed still not completely unveiled, and this constitutes a major hurdle towards a maximized use of this valuable natural resource. However, the completion of genome sequencing projects for several economically relevant tree species (e.g., poplar, grapevine, apple, peach, eucalyptus, sweet orange, Norway spruce; Tuskan et al. 2006, Jaillon et al. 2007, Velasco et al. 2010, Zhang et al. 2012, Nystedt et al. 2013, Verde et al. 2013, Xu et al. 2013, http://eucalyptusdb.bi.up.ac.za/) as well as the availability of advanced next-generation DNA sequencing (NGS) methods now allow tissue-specific gene profiling and comparative genomics/transcriptomics of wood formation between species (Li et al. 2010). This type of approach can be used to retrace the evolutionary history of wood formation, but likewise delivers important information with potential use in breeding programmes for the improvement of wood traits (Grattapaglia and Kirst 2008, Rengel et al. 2009, Dillon et al. 2010, Tian et al. 2012). The availability of high-throughput techniques applied at the systems level, together with a broad collection of wall mutants, will enable the acquisition of an integrated knowledge of one of the most intriguing, and economically and ecologically significant natural processes.
Biosynthesis of wood: behind the scenes
The formation of wood involves different steps: cell division, cell expansion, secondary wall synthesis and programmed cell death (PCD) (Plomion et al. 2001, Demura and Fukuda 2007, Déjardin et al. 2010).
Xylogenesis, i.e., cellular programmes culminating in PCD and in the formation of tracheids/vessels (Roberts and McCann 2000), is a complicated process involving a plethora of genes and their interaction with cellular signals (Demura et al. 2002, Pesquet et al. 2005, Endo et al. 2009, Yoshida et al. 2009, Sorce et al. 2013). Although many of the genes involved in xylogenesis have been identified and functionally characterized (e.g., Ko et al. 2007, Yamaguchi et al. 2008), our understanding of this pathway is still far from being complete. Moreover, the plasticity of plant metabolism, the redundancy of plant gene families (Hamberger and Hahlbrock 2004, Costa et al. 2005, Lu et al. 2006, Soltani et al. 2006, Huang et al. 2010) and the overlapping action of plant growth regulators, as well as the occurrence of alternative splicing (Bao et al. 2013) and post-translational events, add further complexity to the topic. In this section of the review, we will try to retrace the crucial steps that are involved in the development of a functional vascular tissue, from stem cell establishment to the determination of vascular cell fate.
Establishment of procambial precursor cells and vascular stem cell homeostasis
Plant meristems are tissues characterized by high mitotic activity and containing stem cells responsible for indeterminate growth. Apical meristems give rise to vascular meristematic tissue (the procambium), which will be responsible for the postembryonic development of primary xylem and phloem. Sandwiched between these two tissues, the procambium keeps its pluripotent nature and the maintenance of its meristematic activity is crucial for the correct development of vascular tissue (Miyashima et al. 2013).
In Arabidopsis shoots, the procambium separates the primary xylem from the primary phloematic poles in the vascular bundles, but later cambial activity extends to the interfascicular parenchymatic regions, which divide periclinally to form a continuous ring of pluripotent cells (the vascular cambium). Procambial cells can differentiate de novo from existing parenchyma cells and the parenchymatic cells committed to a procambial fate can be easily recognized by their elongated shape and the disposition in files (Schuetz et al. 2013).
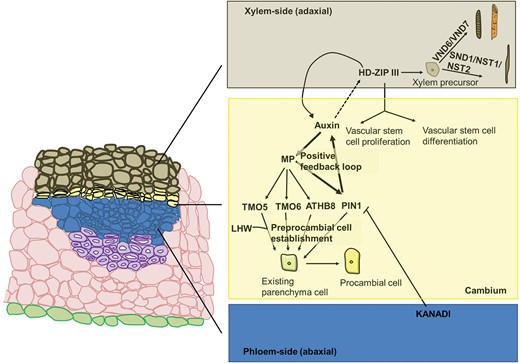
Auxin has been shown to play an essential role in the differentiation of procambial precursor cells (Scarpella et al. 2006, Ilegems et al. 2010): one of the earliest events is indeed the establishment of a self-reinforcing auxin flow. MONOPTEROS/AUXIN RESPONSE FACTOR 5 (MP/ARF5) is induced by auxin and in turn it induces the expression of the efflux-associated protein PINHEAD1 (PIN1) (Figure 1). Other targets of MP involved in procambial cell establishment are the bHLH transcription factor (TF) TARGET OF MP5 (TMO5), the Dof-type Zn-finger protein TMO6 (Schlereth et al. 2010) and the Class III homeodomain-leucine zipper protein (HD-ZIP III) ARABIDOPSIS THALIANA HOMEOBOX GENE 8 (ATHB8; Donner et al. 2009) (Figure 1). The promoter of ATHB8 contains an Auxin-Response Element (ARE) that is bound by MP. ATHB8 is involved in stabilizing the preprocambial fate in narrow cell files against perturbations in auxin concentration (Donner et al. 2009) (Figure 1).
Cartoon depicting a cross section of vascular bundles, showing the epidermis, parenchyma cells, schlerenchymatic phloem fibres, phloem, cambium and xylem. For the vascular tissue layers and the cambium, a description of the key events involved in their establishment is provided on the right. Starting from the cambium, auxin induces MP, whose targets are TMO5/TMO6, ATHB8 and PIN1. TMO5 (together with LHW, LONESOME HIGHWAY), TMO6, ATHB8 and PIN1 contribute to the differentiation of preprocambial cells through de novo differentiation of existing parenchyma cells. PIN1 is involved in reinforcing the canalization of auxin by creating a polar flow. HD-ZIP III expression on the xylematic side is activated by auxin (dotted arrow) and these genes participate in maintaining auxin flow canalization by positively acting on auxin transport genes. HD-ZIP III genes keep the balance between vascular cell stem division and differentiation and activate the pathway leading to the development of xylem precursors. The genes VND6 and VND7 activate the program specific for vessel formation, while SND1, NST1 and NST2 are responsible for fibre formation. KANADI genes, on the phloematic side, inhibit PIN1 activity.
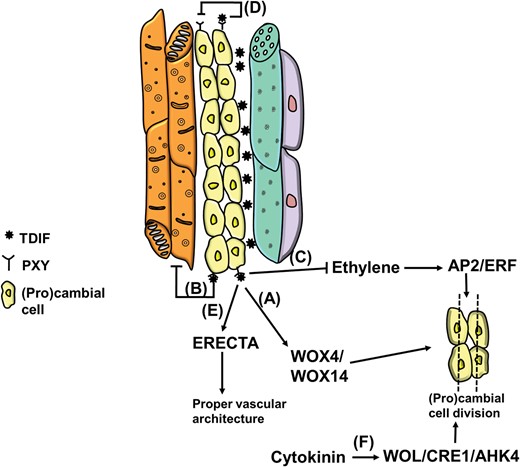
Signalling molecules and cell-to-cell communication are involved in procambial meristem homeostasis. Substantial progress in our understanding of vascular stem cell maintenance and ordered spatial organization of vascular tissues comes from studies that identified the small peptide Tracheary element Differentiation Inhibitory Factor (TDIF) (Ito et al. 2006, Hirakawa et al. 2010a) and its receptor TDIF RECEPTOR/PHLOEM INTERCALATED WITH XYLEM (TDR/PXY) (Fisher and Turner 2007, Hirakawa et al. 2010a). TDIF, which belongs to the family of CLAVATA3/ESR-RELATED (CLE) peptides that in Arabidopsis correspond to CLE41, CLE42 and CLE44 (Ito et al. 2006, Strabala et al. 2006, Oelkers et al. 2008, Etchells and Turner 2010), is secreted from the phloem and binds to the TDR/PXY receptor on the membrane of procambial cells: this signal inhibits the differentiation of procambial cells into xylem cells and favours instead their proliferation (Figure 2). The increased rate of procambial cell division is promoted by WOX4 (a WUSCHEL-related HOMEOBOX gene), which receives the signal via TDR/PXY (Hirakawa et al. 2010b). Very recently it was shown that another WOX gene, WOX14, acts together with WOX4 (Etchells et al. 2013).
Cartoon summarizing the key genes and events affecting (pro)cambial cell division. (A) WOX4/WOX14-dependent regulation of division; (B) TDIF/PXY-dependent inhibition of xylem differentiation; (C) alternative, PXY-independent pathway promoting (pro)cambial cell division via ethylene; (D) negative feedback regulation of TDIF on PXY (Etchells and Turner 2010); (E) PXY/ERECTA-mediated signalling for correct vascular organization (Etchells et al. 2013); (F) positive role of cytokinin on procambial cell proliferation (Hirakawa et al. 2011) via signalling through WOL/CRE1/AHK4 (Mähönen et al. 2000, Inoue et al. 2001, Suzuki et al. 2001).
The regulation of procambial cell homeostasis follows a signalling pathway that is similar to the one operating in the shoot and root apical meristems, and this shows that mechanisms operating in primary meristems are used in plants to control girth increase.
In the absence of TDR/PXY, plants are still able to promote procambial proliferation through a bypass pathway that is ethylene dependent (Barton 2012, Etchells et al. 2012). The ethylene signal is broadcast via TFs of the APETALA2/ETHYLENE RESPONSE FACTOR (AP2/ERF) family. In pxy mutants this route becomes active and the levels of the gene encoding ACS6 (an ethylene biosynthesizing enzyme) consequently increase (Barton 2012, Etchells et al. 2012). However, under normal conditions when TDR/PXY is expressed, the ethylene-dependent bypass pathway is turned off (Figure 2).
The TDIF-TDR/PXY system also regulates the orientation of cell divisions: the asymmetric distribution of TDIF, which is produced exclusively in the phloematic side, determines the correct longitudinal orientation of cell division planes in the procambium (Fisher and Turner 2007, Etchells and Turner 2010, Barton 2012). An alteration of TDIF distribution either on both sides or on the xylematic side leads to highly irregular cell division planes (Etchells and Turner 2010, Barton 2012).
It was demonstrated that procambial cell division and correct organization of vascular tissues are two genetically separable events (Etchells et al. 2013): on one hand, the PXY/TDIF-promoted signalling through WOX4/WOX14 regulates procambial cell division planes and rate of division; on the other, PXY regulates the correct architecture of the vascular tissue through the receptor kinase ERECTA (Etchells et al. 2013) (Figure 2).
Among plant growth regulators, cytokinin also plays a fundamental role in cell division and regeneration: an important gene is WOODEN LEG (WOL/CYTOKININ RESPONSE 1 CRE1/AHK4), which codes for a histidine kinase that functions as a sensor of vascular morphogenesis by regulating asymmetric cell divisions of the phloem and procambium (Mähönen et al. 2000, Inoue et al. 2001, Suzuki et al. 2001) (Figure 2). This receptor senses aminopurine and diphenylurea derivatives and broadcasts the signal across the membrane by starting a His → Asp phosphorelay (Yamada et al. 2001).
HD-ZIP III vs KANADI: the clash of the titans
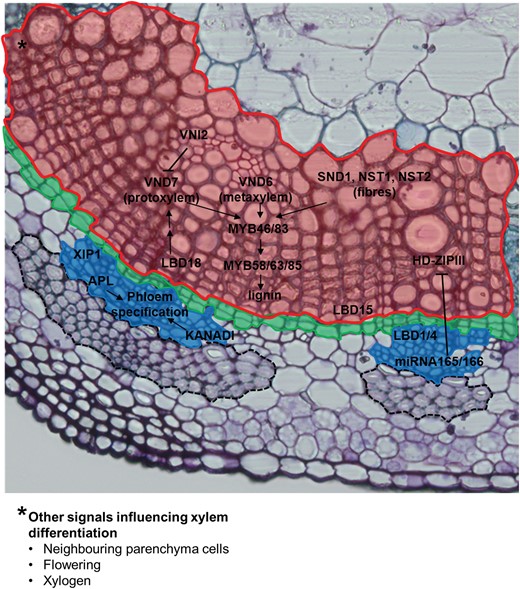
When plants undergo primary development, primary xylem and phloem formation is subject to a radial (or dorsiventral) specification (Emery et al. 2003, Ilegems et al. 2010): the procambium in the vascular bundles will give rise to xylem towards the internal adaxial side and phloem towards the external abaxial side. This specific location of vascular tissues is determined by the opposed activities of two gene families: HD-ZIP III, expressed in the procambium, cambium and developing xylem, and the GARP (a myb-like domain) family of TFs KANADI (KAN), expressed in the phloem (Ilegems et al. 2010) (Figure 3). These genes play an important role in plant development and organogenesis: they control the correct polarity and, consequently, the final shape of plant organs (Kerstetter et al. 2001, Prigge et al. 2005). Polarization along the growth axes of developing plant organs (such as leaves) determines the asymmetric distribution of cell types, which is crucial for the overall plant physiology: the correct dorsiventral polarity of cell types in plant leaves determines the presence of light-harvesting cells on the adaxial side and of cells specialized in gaseous exchange on the abaxial side (Moon and Hake 2011).
Summary of the key genes involved in xylem and phloem specification. Cambium and schlerenchymatic phloem fibres (dotted line) are also indicated. The figure also reports other signals involved in the correct development of xylem tissue, but not directly determining vascular cell identity (asterisk).
Five genes belonging to the HD-ZIP III group have been identified in Arabidopsis thaliana: PHABULOSA (PHB), PHAVOLUTA (PHV), REVOLUTA/INTERFASCICULAR FIBERLESS1 (REV/IFL1), CORONA/INCURVATA4 (CNA/ATHB15/ICU4) and ATHB8. The domain of activity of these genes is quite complex, as they have been shown to perform overlapping, distinct antagonistic roles that cannot be immediately deduced from their phylogenetic kinships (Prigge et al. 2005). Gain-of-function mutations in these genes lead to adaxialization of lateral organs and to amphivasal vascular architecture (i.e., xylem envelops phloem) (Emery et al. 2003). Interestingly, gain-of-function mutations of the HD-ZIP III genes map to a region that is the target of miRNA165/166 and that codes for the lipid/sterol binding domain START (Caño-Delgado et al. 2010, Zhang et al. 2011a). These mutations abolish recognition of the domain by miRNAs, which results in an increased stability of the HD-ZIP III transcripts.
Recently, in roots, a new pattern for xylem specification has been described, which involves the TFs SHORTROOT (SHR) and SCARECROW (SCR) (Carlsbecker et al. 2010): SHR is produced in the stele and transported into the endodermis where, together with SCR, it induces miRNA165/166. These move into the protoxylem where they degrade the cognate HD-ZIP III mRNAs. The gene-dose decrease in HD-ZIP III leads to protoxylem formation (Carlsbecker et al. 2010, Ohashi-Ito and Fukuda 2010).
Four KANADI genes have been identified, KAN1–4, and loss of function of these genes leads to amphivasal vascular tissue (Emery et al. 2003).
The study by Ilegems et al. (2010) has shed light on the interplay existing between KANADI, HD-ZIP III genes and auxin: more specifically, KANADI genes act to restrict auxin canalization by inhibiting the activity of PIN1, while HD-ZIP III genes promote axial elongation of cells and xylem differentiation. In particular, HD-ZIP III genes, which have been shown to interact with the auxin-response-related proteins DORNROESCHEN (DRN) and DRN-like (DRNL) (Chandler et al. 2007), might exert their action through auxin perception/signalling and balance the rate of stem cell division and differentiation (Ilegems et al. 2010).
KAN1 expression directed in procambial precursor cells was indeed shown to exert a negative action on procambium cell formation and division (Ilegems et al. 2010) via an auxin flow-related effect. Therefore, the antagonistic effect of these two classes of genes is not direct, but rather determined by their different control over auxin canalization (Ilegems et al. 2010) (Figure 1).
Cambial activity
The continuous vascular cambium ring ensures both tree girth increase and annual renewability of vascular tissues. It consists of two types of cells, cambial fusiform initials and ray initials. Fusiform initials are elongated pluripotent cells that give rise to, through position-dependent periclinal divisions in the tangential plane, the secondary phloem and xylem. Ray initials give rise to ray cells, which are important for nutrient translocation between the phloem and xylem (Plomion et al. 2001).
Cambial activity is characterized by a rhythmic activity: cells of the cambium can be either active or inactive. One of the hallmarks of cambial meristematic inactivity (i.e., dormancy) is the presence of a narrower cambial layer, together with a different vacuolar structure and cell wall thickening. A very recent study on Chinese fir (Cunninghamia lanceolata (Lamb.) Hook.) has provided a detailed cytological and molecular overview of the events associated with rhythmic cambial activity (Wang et al. 2013): reactivation of cambial activity is accompanied by the presence of a dense cytoplasm and many small vacuoles; active growth is characterized by several layers of thin-walled cambial cells containing a large vacuole and many organelles; during the transition to dormancy the layers decrease to three and the cells have a dense cytoplasm populated by a large vacuole and several smaller ones; dormant cambium has between three and four layers of cells with a very dense cytoplasm (Wang et al. 2013). These transitions in cambial stages are accompanied by modifications in the expression pattern of genes known to be involved in cambial homeostasis division. In particular, the study by Wang et al. (2013) showed that the expression of the Chinese fir orthologues of WOX, CLV and PIN1-like correlated positively with changes in cambial activity.
The regulation of cambial activity is additionally temperature dependent: it is known that an increase in temperature can favour the exit of cambial cells from the dormancy period (e.g., Gričar et al. 2006, Begum et al. 2007) and that spring frost can induce depolymerization of microtubules, thus negatively affecting tree growth (Begum et al. 2012a, 2013). Microtubules have indeed been proposed as cambial ‘sensors’ of temperature fluctuations and appear to control the dormancy period by showing higher stability in cold seasons (Begum et al. 2012b).
The vascular cambium, although characterized by different types of cells (phloem and xylem mother cells and initials), is difficult to differentiate anatomically. However, it is characterized by zones showing specific transcriptional fingerprints. An elegant study carried out on the aspen cambial zone has revealed the presence of different transcriptomes between cambial initials and mother cells (Schrader et al. 2004). Interestingly, the careful sampling strategy of this study enabled the identification of marker genes associated with the phloem/xylem side and the centre of the cambial zone. Flavonoid biosynthesis- and stress-related genes (which might be involved in protecting the cambium under stress conditions) are specific for the phloem side, and genes involved in cell division/expansion and protein turnover are specific for the xylem side (Schrader et al. 2004). Only six known genes related to metabolism, signal transduction and transcriptional regulation were differentially regulated in the central cambial region (Schrader et al. 2004).
The above-mentioned studies represent emblematic examples of the importance of molecular analyses on cambial tissues. Understanding more about the regulation of cambial activity might indeed disclose important knowledge that can be used for genetic engineering programmes aiming at increasing the woody biomass and boosting its qualities. Because of its location, susceptibility to temperature and structural variability, cambial tissue represents a challenging tissue to sample and to study. However, several studies have described successful protocols for the sampling of cambial tissue (scraping of the inner layer of fresh bark, or of the exposed surface of immature xylem, as reported in Durand et al. 2010 and Wang et al. 2013), and an experimental set-up mimicking the initiation and subsequent differentiation of cambial cells has even been devised. This technique, known as stem girdling, consists in peeling off a strip of bark from the tree: after girdling, new cambium and xylem can reform in sequence in the living tree. After girdling, the newly formed sieve elements were shown to appear before the regenerated cambium and both derive from differentiating xylem cells (Pang et al. 2008, Zhang et al. 2011b, Chen et al. 2013). This shows that bark regeneration can take place via cells transforming their fates directly into other cell types, without the activity of stem cells or the transition into a pluripotent state (Sena et al. 2009, Chen et al. 2013). The regeneration of secondary vascular tissue involves three main steps: callus formation and xylem dedifferentiation, sieve element formation, and cambium and phellogen regeneration (Chen et al. 2013). If tissue sampling is carried out during the different stages of secondary tissue reformation, ‘snapshots’ of the regenerative events can be obtained (e.g., Du et al. 2006).
The possibility of combining gene expression studies (through NGS and qPCR of a selected subgroup of genes) with proteomics and metabolomics will dramatically increase the depth of the information obtained. It is desirable in the future to progress towards an integrated knowledge of wood biosynthesis by providing analyses carried out at multiple analytical levels.
Xylem and phloem fate: two sides of the same coin
The xylogenic fate of differentiating cambial cells is under the control of master regulators, which are TFs regulating the morphological events accompanying secondary cell wall deposition. Transcription factors of the NAM/ATAF/CUC (NAC) domain family have been shown to regulate xylem and fibre differentiation: in particular, VND6-VND7 establish metaxylem and protoxylem formation (Kubo et al. 2005, Demura and Fukuda 2007), while SND1/NST3, NST1 and NST2 regulate fibre fate (Zhong et al. 2006, Demura and Fukuda 2007, Mitsuda et al. 2007) (Figures 1 and 3 and Table 1). Recently an interacting partner of VND7 was identified, VND-INTERACTING 2 (VNI2), which negatively regulates the activity of VND7 (Yamaguchi et al. 2010; Figure 3).
Summary of the key genes, with details concerning their function and gene family, determining phloem and xylem development.
| Function . | Gene family . | Gene name in A. thaliana . |
|---|---|---|
| Cambial organization/stem cell maintenance | LRR-RLK | PXY/TDR |
| CLE | TDIF | |
| WOX | WOX4/WOX14 | |
| Radial patterning (adaxial) | HD-ZIP III | ATHB8 |
| REV/IFL1 | ||
| PHB | ||
| PHV | ||
| CNA/ATHB15 | ||
| Radial patterning (abaxial) | KANADI | KAN1 |
| KAN2 | ||
| KAN3 | ||
| KAN4 | ||
| Specification of phloem tissue identity | MYB | APL |
| Specification of xylem tissue identity | NAC | VND6 |
| VND7 | ||
| SND1/NST1 | ||
| AGP/nsLTP | XYP1/XYP2 | |
| Activation of phloem differentiation | LBD | LBD1/LBD4 |
| Activation of xylem differentiation | LBD | LBD15/LBD18 |
| Function . | Gene family . | Gene name in A. thaliana . |
|---|---|---|
| Cambial organization/stem cell maintenance | LRR-RLK | PXY/TDR |
| CLE | TDIF | |
| WOX | WOX4/WOX14 | |
| Radial patterning (adaxial) | HD-ZIP III | ATHB8 |
| REV/IFL1 | ||
| PHB | ||
| PHV | ||
| CNA/ATHB15 | ||
| Radial patterning (abaxial) | KANADI | KAN1 |
| KAN2 | ||
| KAN3 | ||
| KAN4 | ||
| Specification of phloem tissue identity | MYB | APL |
| Specification of xylem tissue identity | NAC | VND6 |
| VND7 | ||
| SND1/NST1 | ||
| AGP/nsLTP | XYP1/XYP2 | |
| Activation of phloem differentiation | LBD | LBD1/LBD4 |
| Activation of xylem differentiation | LBD | LBD15/LBD18 |
Summary of the key genes, with details concerning their function and gene family, determining phloem and xylem development.
| Function . | Gene family . | Gene name in A. thaliana . |
|---|---|---|
| Cambial organization/stem cell maintenance | LRR-RLK | PXY/TDR |
| CLE | TDIF | |
| WOX | WOX4/WOX14 | |
| Radial patterning (adaxial) | HD-ZIP III | ATHB8 |
| REV/IFL1 | ||
| PHB | ||
| PHV | ||
| CNA/ATHB15 | ||
| Radial patterning (abaxial) | KANADI | KAN1 |
| KAN2 | ||
| KAN3 | ||
| KAN4 | ||
| Specification of phloem tissue identity | MYB | APL |
| Specification of xylem tissue identity | NAC | VND6 |
| VND7 | ||
| SND1/NST1 | ||
| AGP/nsLTP | XYP1/XYP2 | |
| Activation of phloem differentiation | LBD | LBD1/LBD4 |
| Activation of xylem differentiation | LBD | LBD15/LBD18 |
| Function . | Gene family . | Gene name in A. thaliana . |
|---|---|---|
| Cambial organization/stem cell maintenance | LRR-RLK | PXY/TDR |
| CLE | TDIF | |
| WOX | WOX4/WOX14 | |
| Radial patterning (adaxial) | HD-ZIP III | ATHB8 |
| REV/IFL1 | ||
| PHB | ||
| PHV | ||
| CNA/ATHB15 | ||
| Radial patterning (abaxial) | KANADI | KAN1 |
| KAN2 | ||
| KAN3 | ||
| KAN4 | ||
| Specification of phloem tissue identity | MYB | APL |
| Specification of xylem tissue identity | NAC | VND6 |
| VND7 | ||
| SND1/NST1 | ||
| AGP/nsLTP | XYP1/XYP2 | |
| Activation of phloem differentiation | LBD | LBD1/LBD4 |
| Activation of xylem differentiation | LBD | LBD15/LBD18 |
Transcription factors of the MYB family, MYB46 and MYB83, act downstream of the NAC regulators (Zhong and Ye 2007) (Figure 3 and Table 1). In turn, they activate MYBs (e.g., MYB58, MYB63, MYB85) regulating genes in the lignin pathway: the promoters of these genes possess AC-rich elements (H-boxes) and, by binding to these, MYBs regulate their xylem-specific expression (Rogers and Campbell 2004, Goicoechea et al. 2005, Demura and Fukuda 2007, Winzell et al. 2010).
Subtle molecular events also contribute to fine-tune the onset of secondary cell wall biogenesis. This was recently demonstrated in poplar: for the first time a xylem-specific splice variant of the SND1-A2 TF was shown to act as a dominant negative regulator repressing the activation of PtrMYB021, a master switch of the secondary wall deposition programme (Li et al. 2012).
Neighbouring parenchyma cells contribute to xylem lignification in Arabidopsis by nursing, supporting and fuelling the differentiating xylem cells with important precursors. A recent study showed that specific silencing of CINNAMOYL CoA-REDUCTASE1 (CCR1) in lignifying cells does not hinder lignin deposition in the xylem; however, it impairs lignification of extraxylary fibres, which show cell-autonomous lignin deposition (Smith et al. 2013).
The plant growth regulator brassinosteroid and xylogen, a secreted arabinogalactan/non-specific lipid-transfer protein, are also involved in the differentiation of xylem cells (Caño-Delgado et al. 2004, Motose et al. 2004, Yamamoto et al. 2007). Interestingly, in Arabidopsis a correlation between flowering and secondary growth of the hypocotyl and roots was found: it was proposed that girth increase could reinforce the plant in preparation for bolting (Sibout et al. 2008).
It is known that the MYB coiled-coil TF altered phloem development (APL) is necessary (but not sufficient) for the establishment of phloem identity in A. thaliana (Bonke et al. 2003), and that lateral organ boundary 1 (LBD1) is a positive regulator of phloem differentiation (Yordanov et al. 2010) (Figure 3 and Table 1). Altered phloem development promotes phloem differentiation and inhibits xylem formation. Indeed a recessive apl mutation triggers the formation of xylem in places where the phloem is normally present, whereas its ectopic expression inhibits the formation of xylem (Bonke et al. 2003). Members of the LBD family have recently been shown to play an essential role in correctly determining the boundaries between cambium–xylem and cambium–phloem: in particular, LBD1 and LBD4 are expressed at the boundary between phloem and cambium, while LBD15 and LBD18 are expressed at the boundary with xylem (Yordanov et al. 2010) (Figure 3). These genes promote xylem and phloem differentiation by activating as yet unknown genes and by confining the activity of meristem identity genes to the cambium. Moreover, LBD members (LBD18 and LBD30) have been shown to be expressed in immature tracheary elements (TEs) and LBD18 is involved in a positive feedback loop for VND7 expression (Soyano et al. 2008, Demura and Ye 2010) (Figure 3).
Markers associated with phloem precursor cells that enable the spatial and temporal analysis of vascular tissue differentiation have been identified in A. thaliana (Bauby et al. 2007). Five marker genes characterizing different stages in phloem development have been found by gene trapping in Arabidopsis: PD1–PD5, with PD1–PD4 being expressed in the protophloem and PD5 in the metaphloem as well (Bauby et al. 2007). The annotation of these markers showed that PD1 is a glycosylphosphatidylinositol-anchored protein, PD3 a phosphatidylinositol phosphate-4,5-kinase, PD4 a transcriptional activator and PD5 a glycine-rich protein (Bauby et al. 2007).
The receptor-like kinase (RLK) XYLEM INTERMIXED WITH PHLOEM 1 (XIP1) prevents ectopic lignification in phloem cells and therefore contributes to control the correct organization/morphology of conductive elements (Bryan et al. 2012).
Beyond transcriptional regulation, post-translational events are also crucial for determining correct secondary growth. As an example, proteases, which have so far been given less attention, play a key role in regulating vascular tissue development: they are for instance involved in the processing of the above-mentioned CLE peptides and therefore represent important targets for further functional studies (Petzold et al. 2012).
Classic and emerging models to study secondary growth
Although trees constitute the optimal system to understand the process of wood biosynthesis and the best models for field testing (Plomion et al. 2001), milestone data concerning xylogenesis have been and are still being obtained through the study of non-woody organisms (Plomion et al. 2001, Zhang et al. 2011a, 2011b). Examples are the xylogenic Zinnia elegans Jacq. mesophyll cell suspension line (Fukuda and Komamine 1980), now considered a ‘classic’ in the study of xylogenesis and TE formation, and the tobacco BY-2 cell suspension line overexpressing VND7 under the control of oestrogen- or glucocorticoid-inducible systems (Yamaguchi et al. 2008, 2010). More cell lines are being developed for the study of xylogenesis in other organisms (e.g., Ogita et al. 2012), which will contribute to foster knowledge of xylogenesis in economically relevant species.
Rather unexpectedly, also the small annual weed A. thaliana is very useful for the study of wood formation (e.g., Strabala and Macmillan 2013): the hypocotyl of this model organism undergoes secondary growth (Chaffey et al. 2002, Nieminen et al. 2004, Ragni and Hardtke 2013). A strong body of evidence in the literature points towards the conservation of cellular, metabolic and biochemical pathways across the plant kingdom (Plomion et al. 2001). Many of the genetic mechanisms involved in secondary growth also regulate the shoot apical meristem and, most interestingly, the woody habit likely resulted from the modification of genetic programmes already present in herbaceous progenitors (Groover 2005). Therefore, a similar regulatory mechanism governing secondary development is shared between herbaceous and woody organisms (Zhang et al. 2011a, 2011b).
This means that if a common regulatory mechanism controlling cambium and secondary growth is shared among herbaceous and woody plants, initial research can be performed on the experimentally more amenable herbaceous plants and the analysis can be later extended to woody species (Groover 2005).
Auxin signalling, the role of microtubules and transcriptional wiring (Oh et al. 2003) are elements regulating wood formation that have been extensively studied in A. thaliana. It has even been shown that the deposition of reaction wood (RW) can be induced under certain conditions in this weed (Wyatt et al. 2010). The advantages of using herbaceous plants and in particular A. thaliana are evident: shorter growth times, the possibility of obtaining knock-out or overexpressing lines relatively quickly and availability of a broad collection of wall mutants.
Additional important information can also be obtained by studying a specific group of plants that show a natural heterogeneity in lignification patterns, i.e., fibre crops (Neutelings 2011, Guerriero et al. 2013). Their stems constitute an excellent system to study secondary growth, as they show different cell wall metabolic stages both radially and axially. The presence of a specific point along the axis of the stem, i.e., the ‘snap point’ (Gorshkova et al. 2003), marks an easily recognizable region below which secondary cell wall thickening takes place. Additionally, fibre crops are characterized by the occurrence of extraxylary structures that derive from the (pro)cambium, the so-called bast fibres, characterized by the presence of a gelatinous layer (G-layer) similar to the one observed in tension wood (TW).
Although TW and bast fibres have different origins, a set of genes, among which fasciclin-like arabinogalactan proteins (FLAs) are particularly represented (e.g., Kaku et al. 2009), was shown to be enriched in the two tissues (Hobson et al. 2010). The study of fibre crops can potentially allow further progress in elucidating common mechanisms operating during the formation of bast fibres and RW of trees.
The stem of most fibre crops also shows a specific tissular organization, with woody tissues being confined to the core and bast fibres/phloematic tissues located in the cortex (Guerriero et al. 2013). The possibility of peeling off the outer tissue containing phloem, bast fìbres and cambium from the inner woody core with libriform cells enables the molecular study of xylary and extraxylary structures of isogenic tissues exposed to 100% identical growth conditions (Guerriero et al. 2013).
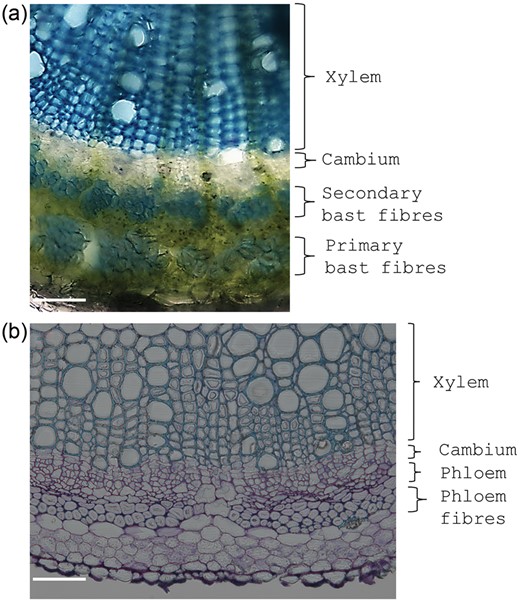
It has furthermore been reported that hypocotyls of flax show a parallelism with the sequential stages of stem differentiation in the upper, middle and bottom regions (Roach and Deyholos 2008) and that they display a phase of secondary cell wall biosynthesis, which could make these hypocotyls interesting, practical systems in which to study secondary growth. We have observed the same behaviour in Cannabis sativa L. hypocotyls, where at later stages of development even secondary bast fibres begin to appear (Figure 4a). The advantage of using fibre crops to study secondary growth is further supported by the availability of genomes for some species (van Bakel et al. 2011, Wang et al. 2012) and by the possibility of studying the orthologues of known genes from model plants (such as A. thaliana) involved in vascular tissue development (Table 2).
Cannabis sativa putative orthologues of A. thaliana genes involved in vascular tissue development. Details of locus id, sequence id (from the Medicinal Plants Genomic Resource, http://medicinalplantgenomics.msu.edu/) and e-values are indicated.
| Gene A. thaliana locus id . | Putative C. sativa orthologue sequence id . | e-value . |
|---|---|---|
| AT1G07900 LBD1 | csa_locus_28967_iso_1_len_702_ver_2 | 6.3 × 10−56 |
| AT1G31320 LBD4 | csa_locus_50726_iso_1_len_941_ver_2 | 6.7 × 10−68 |
| AT2G40470 LBD15 | csa_locus_33561_iso_1_len_416_ver_2 | 7.9 × 10−58 |
| AT2G45420 LBD18 | csa_locus_5911_iso_1_len_834_ver_2 | 1.1 × 10−49 |
| AT5G61480 PXY/TDR | csa_locus_15819_iso_2_len_3586_ver_2 | 0 |
| AT1G79430 APL | csa_locus_6872_iso_2_len_1251_ver_2 | 1.1 × 10−85 |
| AT5G64080 XYP1 | csa_locus_7253_iso_1_len_1173_ver_2 | 9 × 10−41 |
| AT5G52380 VND6 | csa_locus_14303_iso_1_len_1052_ver_2 | 8.1 × 10−88 |
| AT1G71930 VND7 | csa_locus_13095_iso_2_len_1074_ver_2 | 4.4 × 10−72 |
| AT2G46770 NST1 | csa_locus_18617_iso_1_len_1319_ver_2 | 8.8 × 10−103 |
| AT1G32770 NST3/SND1 | csa_locus_13044_iso_1_len_1547_ver_2 | 3.5 × 10−92 |
| Gene A. thaliana locus id . | Putative C. sativa orthologue sequence id . | e-value . |
|---|---|---|
| AT1G07900 LBD1 | csa_locus_28967_iso_1_len_702_ver_2 | 6.3 × 10−56 |
| AT1G31320 LBD4 | csa_locus_50726_iso_1_len_941_ver_2 | 6.7 × 10−68 |
| AT2G40470 LBD15 | csa_locus_33561_iso_1_len_416_ver_2 | 7.9 × 10−58 |
| AT2G45420 LBD18 | csa_locus_5911_iso_1_len_834_ver_2 | 1.1 × 10−49 |
| AT5G61480 PXY/TDR | csa_locus_15819_iso_2_len_3586_ver_2 | 0 |
| AT1G79430 APL | csa_locus_6872_iso_2_len_1251_ver_2 | 1.1 × 10−85 |
| AT5G64080 XYP1 | csa_locus_7253_iso_1_len_1173_ver_2 | 9 × 10−41 |
| AT5G52380 VND6 | csa_locus_14303_iso_1_len_1052_ver_2 | 8.1 × 10−88 |
| AT1G71930 VND7 | csa_locus_13095_iso_2_len_1074_ver_2 | 4.4 × 10−72 |
| AT2G46770 NST1 | csa_locus_18617_iso_1_len_1319_ver_2 | 8.8 × 10−103 |
| AT1G32770 NST3/SND1 | csa_locus_13044_iso_1_len_1547_ver_2 | 3.5 × 10−92 |
Cannabis sativa putative orthologues of A. thaliana genes involved in vascular tissue development. Details of locus id, sequence id (from the Medicinal Plants Genomic Resource, http://medicinalplantgenomics.msu.edu/) and e-values are indicated.
| Gene A. thaliana locus id . | Putative C. sativa orthologue sequence id . | e-value . |
|---|---|---|
| AT1G07900 LBD1 | csa_locus_28967_iso_1_len_702_ver_2 | 6.3 × 10−56 |
| AT1G31320 LBD4 | csa_locus_50726_iso_1_len_941_ver_2 | 6.7 × 10−68 |
| AT2G40470 LBD15 | csa_locus_33561_iso_1_len_416_ver_2 | 7.9 × 10−58 |
| AT2G45420 LBD18 | csa_locus_5911_iso_1_len_834_ver_2 | 1.1 × 10−49 |
| AT5G61480 PXY/TDR | csa_locus_15819_iso_2_len_3586_ver_2 | 0 |
| AT1G79430 APL | csa_locus_6872_iso_2_len_1251_ver_2 | 1.1 × 10−85 |
| AT5G64080 XYP1 | csa_locus_7253_iso_1_len_1173_ver_2 | 9 × 10−41 |
| AT5G52380 VND6 | csa_locus_14303_iso_1_len_1052_ver_2 | 8.1 × 10−88 |
| AT1G71930 VND7 | csa_locus_13095_iso_2_len_1074_ver_2 | 4.4 × 10−72 |
| AT2G46770 NST1 | csa_locus_18617_iso_1_len_1319_ver_2 | 8.8 × 10−103 |
| AT1G32770 NST3/SND1 | csa_locus_13044_iso_1_len_1547_ver_2 | 3.5 × 10−92 |
| Gene A. thaliana locus id . | Putative C. sativa orthologue sequence id . | e-value . |
|---|---|---|
| AT1G07900 LBD1 | csa_locus_28967_iso_1_len_702_ver_2 | 6.3 × 10−56 |
| AT1G31320 LBD4 | csa_locus_50726_iso_1_len_941_ver_2 | 6.7 × 10−68 |
| AT2G40470 LBD15 | csa_locus_33561_iso_1_len_416_ver_2 | 7.9 × 10−58 |
| AT2G45420 LBD18 | csa_locus_5911_iso_1_len_834_ver_2 | 1.1 × 10−49 |
| AT5G61480 PXY/TDR | csa_locus_15819_iso_2_len_3586_ver_2 | 0 |
| AT1G79430 APL | csa_locus_6872_iso_2_len_1251_ver_2 | 1.1 × 10−85 |
| AT5G64080 XYP1 | csa_locus_7253_iso_1_len_1173_ver_2 | 9 × 10−41 |
| AT5G52380 VND6 | csa_locus_14303_iso_1_len_1052_ver_2 | 8.1 × 10−88 |
| AT1G71930 VND7 | csa_locus_13095_iso_2_len_1074_ver_2 | 4.4 × 10−72 |
| AT2G46770 NST1 | csa_locus_18617_iso_1_len_1319_ver_2 | 8.8 × 10−103 |
| AT1G32770 NST3/SND1 | csa_locus_13044_iso_1_len_1547_ver_2 | 3.5 × 10−92 |
(a) Stem cross section of the hemp hypocotyl (21 days after sowing) showing the presence of primary and secondary bast fibres, cambium and xylem. (b) Stem cross section of the alfalfa stem showing the presence of phloem bundles, phloem tissue, cambium layer and xylem. Scale bars refer to 100 µm.
Fibre crop hypocotyls can also be quite useful for studies concerning secondary growth and abiotic stress: data on the effects of heavy metals (e.g., cadmium, Cd) on hypocotyl cell wall structure and anatomical organization are already available in the literature (Citterio et al. 2003, Douchiche et al. 2007, 2010, 2011).
A further useful system to study secondary wall formation and lignification is the herbaceous crop alfalfa (Medicago sativa L.). Progress in cell wall lignification can be observed along its stem, with clear bundles of primary vascular tissues in younger internodes (Figure 4b) which become progressively thicker and more lignified at the bottom of the stem. Alfalfa was shown to be a valuable model for the study of wall deposition in dicots (Tesfaye et al. 2009), and although its genome has not yet been sequenced, the availability of the genome of the closely related barrel medic (Medicago truncatula Gaertn.) can be used to address molecular studies on this plant (Yang et al. 2010).
It is therefore clear that beyond A. thaliana and xylogenic cell lines, great potential for further molecular investigations on secondary growth is held by non-woody models, such as fibre crops and M. sativa.
Wood typologies: nature vs nurture
Wood is a fascinating biomaterial that displays a natural variability. Its 3D structure differs not only among species (Déjardin et al. 2010), but can also be influenced by other factors, such as tree developmental stage, position along the stem and seasonality. In this section, the different wood typologies will be discussed, i.e., those variations in wood anatomy/structure that are both natural and specifically triggered by exogenous events (such as, for instance, environmental cues). Although many other exogenous aspects, such as soil composition and fertility, regulate plant metabolism and, ultimately, wood formation, they will not be discussed in the present review. Studies in the literature have already extensively dealt with this topic (e.g., Dubouzet et al. 2013).
The impact that environmental cues have on forests is a factor of the utmost importance not only for ecological reasons, but also from an economic point of view. Environmental constraints trigger consequences for forest productivity, by affecting critical parameters such as tree growth, wood properties and density (Bouriaud et al. 2005). Wood density is an important feature determining overall wood quality and performance and it is highly sensitive to fluctuations in the climate. It is hence clear how intimate the relationship between environment and wood formation is and how environmental cues can induce the formation of wood anomalies, and affect density and therefore quality.
Natural wood typologies
An emblematic example of natural group-specific wood typology is the distinction between soft- and hardwood. They differ in their microscopic anatomical appearance, with gymnosperm softwood being more compact and lacking the porous aspect of angiosperm hardwood. This disparity is reflected by differences in the xylem transcriptome between gymnosperms and angiosperms. Conifers share a highly conserved xylem transcriptome with gymnosperm-specific genes, while angiosperms display a largely diversified xylem transcriptome, which is the result of a greater susceptibility to evolutionary forces (Li et al. 2010).
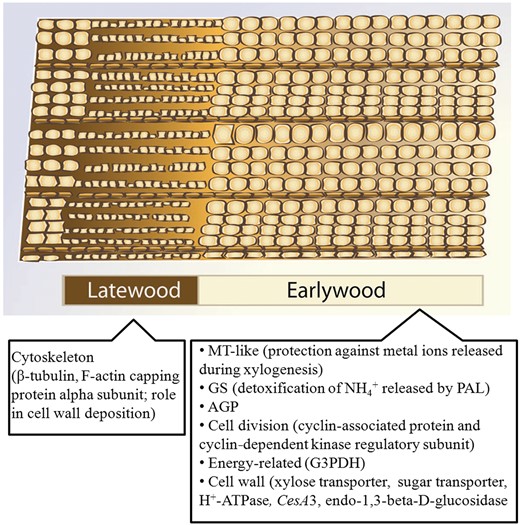
The natural wood variability can be clearly observed in the annual growth pattern of wood rings, which is the direct outcome of seasonal oscillations in cambial activity. Obvious anatomical differences between the wood deposited in spring (earlywood/springwood; EW) and autumn (latewood/autumn- or summerwood; LW) can be distinguished (Figure 5): LW appears as a denser layer with thicker and smaller fibres, while EW is broader with wider tracheids/vessel elements and with a more ‘relaxed’ texture.
Cartoon representing the differences between LW and EW in successive growth rings, with details on the most representative upregulated genes (Paiva et al. 2008a). MT, metallothionein; GS, glutamine synthetase; AGP, arabinogalactan protein; G3PDH, glyceraldehyde-3-phosphate dehydrogenase.
The different characteristics of EW and LW are one of the key factors determining intra-ring variation, i.e., variation in wood density, a critical parameter defining the overall quality of wood.
The majority of studies on gene expression in EW and LW come from conifers (Egertsdotter et al. 2004, Paiva et al. 2008a, Li et al. 2009, 2010, 2011). In particular, the work by Paiva et al. (2008a) resulted in a comprehensive view on the transcriptome changes associated with EW/LW deposition. Cell-division-related (e.g., cyclin-associated proteins, cyclin-dependent kinase) and energy-related genes (cytochrome c1, naphthoate synthase, glyceraldehyde-3-phosphate dehydrogenase, fructose-bisphosphate aldolase) genes together with genes associated with sugar transport, intracellular trafficking and cell wall biosynthesis (e.g., cellulose synthase 3, CesA3) were strongly upregulated in EW (Figure 5). This comprehensive analysis shows the rhythmic cadence of gene expression that regulates secondary growth in trees and provides evidence for a higher cellular (and consequently metabolic) activity during spring.
Differences in wood typology can also be observed in young vs mature trees (Déjardin et al. 2010) as well as in wood from the upper and lower parts of a tree stem (crown- and basewood, CrW/BW; Li et al. 2011). Molecular and phenotypic profiling of BW and CrW in maritime pine has shown that the cellulose content at the base of the stem was higher than at the top and that this was reflected by the preferential expression of CesAs at the base of the tree (Paiva et al. 2008b). Proteomic analysis showed that enzymes involved in methionine metabolism were highly upregulated at the base, which might indicate an increased requirement of methyl-transfer reactions necessary for monolignol biosynthesis (Paiva et al. 2008b). This explains why more lignin is usually present at the base of the stem, a finding in agreement with the thicker cell walls of BW tracheids (Paiva et al. 2008b).
Interestingly, in pine BW, stress-related genes were also upregulated, in particular low-molecular-weight heat-shock proteins, which might be involved in lengthening the wall thickening phase and delaying PCD (Paiva et al. 2008b).
In CrW, the higher rate of cell division and expansion results in the preferential abundance of cytoskeleton-related proteins, of genes belonging to the ‘protein synthesis’ group and of GA/abscisic acid (ABA) responsive ones (Paiva et al. 2008b).
A gradient in wood density, wall thickness, microfibril angle (MFA) and gene expression along the Pinus radiata D. Don stem has also been described (Cato et al. 2006). Interestingly, protein kinases, together with WD40 repeat proteins, three CesAs, a sucrose synthase (SuSy) and lignin-related genes were upregulated at the base (Cato et al. 2006). These data are quite interesting, considering the role that these genes play in cell wall deposition (Brill et al. 2011, Guerriero et al. 2012).
Juvenile wood (JW, sometimes referred to as ‘corewood’), which constitutes an inner cylinder running along the stem of a tree (Pikk and Kask 2004) closest to the pith, is characterized by a lower density and higher MFA, which confers a certain elasticity to the stem, an important feature in case of bending due to wind (Déjardin et al. 2010). Mature wood (MW, sometimes referred to as ‘outerwood’) shows a lower MFA which, consequently, determines a stiffer, stronger, mechanically superior wood with respect to JW and is thus desirable for use in the construction industry.
Transcriptome profiling of xylem tissue in P. radiata JW and MW has identified >140 genes responding to wood maturation, 34% of which are involved in cell wall formation (Li et al. 2011). In MW the majority of upregulated genes were indeed involved in secondary cell wall deposition and cytoskeleton-related processes, while in JW the greatest proportion of genes was involved in the pectin pathway, hormone signalling, water transport, stress response, cell division/differentiation and wall expansion (Li et al. 2011). The upregulation of genes involved in hormone signalling, water transport and stress response indicates that young trees are subjected to greater environmental stress (e.g., higher water competition). As the tree gets older, the pathways controlling the synthesis of cellulose and lignin, as well as the orientation of the cellulosic microfibrils, are preferentially upregulated to allow adaptation to the tensile stress generated by the increased tree weight (Li et al. 2011). This same work, which also studied seasonal wood formation in young and mature trees, revealed that the greatest changes in expression were observed during spring with respect to autumn.
Reaction wood
Trees respond to mechanical stress (e.g., wind and gravitational stimuli) by synthesizing the so-called RW, which has the purpose of re-orienting the tree axis towards an upright position. Reaction wood comprises TW, typical of angiosperms, and compression wood (CW), typical of conifers. Tension wood forms on the upper side of the stem that has been displaced, while CW forms on the lower side. Tension wood and CW are very different, both in morphology and in composition: the former is characterized by few small vessels and its fibres possess the typical G-layer composed almost exclusively of pure cellulose with a low MFA. Compression wood, on the other hand, is composed of thick-walled tracheids with a higher lignin content (increased p-hydroxyphenylpropane units) and higher MFA (Hellgren et al. 2004).
The formation of RW comprises a re-organization of the wall biosynthetic gene repertoire and the synthesis of specific metabolites (Andersson-Gunnerås et al. 2006). A detailed molecular analysis of TW formation in poplar has identified differential expression of carbohydrate- and wall-related genes. Fasciclin-like arabinogalactan proteins in particular showed a consistent increase in expression with respect to normal wood, while lignin biosynthetic genes were downregulated. CesAs also showed variation in gene expression; in particular PttCesA1 decreased, while PttCesA3-2 and PttCesA8-2 were slightly upregulated (Andersson-Gunnerås et al. 2006). SuSy genes were upregulated in TW, a finding that points towards an increased C-shunting towards cellulose production. Interestingly, the increased expression of fructokinase during TW formation yields an excess of fructose-6-P that could be shunted through the pentose phosphate and shikimic acid pathway towards the biosynthesis of N-acetylglucosamine (for glycosylation of arabinogalactan proteins, AGPs) or sucrose regeneration, instead of lignin biosynthesis (Andersson-Gunnerås et al. 2006).
The formation of TW is under the control of TFs and phytohormones: PttMYB21a was shown to be upregulated in poplar TW, together with three LIM genes. As expected, genes related to ethylene and auxin signalling were differentially expressed in TW (Andersson-Gunnerås et al. 2006).
In Populus deltoides W. Bartram ex Marshall, it was demonstrated that the G-layer does contain lignin with a predominance of syringyl units (Joseleau et al. 2004); therefore, TW is partially lignified. However, as Andersson-Gunnerås et al. (2006) pointed out, the glycosylation of monolignols was inhibited during TW formation; moreover laccases and peroxidases, involved in the polymerization step of monolignols, were downregulated (Pilate et al. 2004).
There also appears to be a stereoselective regulation of lignin polymerization during TW formation (Kwon 2008). In yellow poplar, Akiyama et al. (2003) reported a higher abundance of the erythro over threo forms of β-O-4-linked lignin.
A study performed on Eucalyptus showed a complex response of the lignin biosynthetic genes to tension stress (Paux et al. 2005). During the early response to gravistimulus, the expression of the genes involved in monolignol biosynthesis was downregulated (as opposed to a secondary cell wall CesA, which was upregulated), but then the expression levels reached values similar to the control condition; the lignin biosynthetic genes related to methylation reactions were instead downregulated at later stages of gravistimulus, and therefore it was proposed that the reduction in lignin content (more specifically of G-units) observed in TW is most likely due to the regulation of genes encoding enzymes involved in methylation steps (Paux et al. 2005).
The study of TW formation has important outcomes for the development of strategies aimed at increasing the saccharification potential of woody biomass: its high cellulose and low lignin content are desirable features for the production of biofuels. Indeed many studies have recently addressed the enzymatic hydrolysis of wood produced in trees subjected to gravistimulation (e.g., Brereton et al. 2011). Moreover, the response to RW induction is a trait that has been shown to characterize the variability in wall glucan accessibility of different willow genotypes (and possibly other angiosperms) (Brereton et al. 2011).
As seen for TW, the formation of CW also requires modified expression of genes involved in wall biosynthesis: in particular, the lignin pathway was shown to be induced (in particular monolignol biosynthetic genes), together with an increase in cellulose biosynthesis (Villalobos et al. 2012). Expression of both CesA and SuSy increased and this can be linked to the deposition of a thicker S2 layer that is typically found in the tracheids of CW (Villalobos et al. 2012). Cytoskeleton-associated genes (namely tubulins and microtubule-associated proteins) were also upregulated: this might be linked to secondary cell wall biosynthesis and changes in MFA orientation (Villalobos et al. 2012).
Genes involved in the methyl cycle (notably SAM) were upregulated in CW, as well as glutamine synthetase (GS), which is responsible for the assimilation of ammonia released by PAL during lignification. Genes involved in lignan metabolism were also shown to be upregulated and this is a very interesting finding, as lignans are related to defence/heartwood formation and might therefore help the tree cope with the oxidative stress triggered by lignin polymerization during CW formation (Villalobos et al. 2012).
Impact of salinity and drought on wood formation
In the current global climate change scenario, salinity and drought certainly represent relevant threatening factors. Salinity triggers changes at different levels: on the cambial region, on the xylem elements and, as a consequence of changes in density, on the mechanical properties of wood (Lautner 2013). When exposed to salinity, cambial cells appear disorganized, form a thinner layer and display a highly vacuolated cytoplasm; xylem vessels have a smaller lumen and typically increase in number (e.g., Escalante-Pérez et al. 2009, Lautner 2013). The increase in vessel number is the result of the ‘hydraulic safety principle’ (Janz et al. 2012), according to which cavitation episodes pose less dramatic problems if the number of conduits is higher.
Several studies have undertaken the analysis of the response to salinity in trees. It is not possible here to go through all the data available in the literature; however, we would like to draw the reader's attention to a recent study that describes the presence of a particular type of wood, ‘pressure wood’, triggered upon salt stress in poplar (Janz et al. 2012). This wood is the result of downregulation in expression of a set of genes that is instead upregulated during TW formation, namely those encoding FLAs and COBRA-like proteins, xyloglucan endotransglycosylase, pectin methylesterase, expansin and xylosidase (Janz et al. 2012). Besides modification in cell-wall-related gene expression, the study also showed the upregulation of genes involved in osmotic stress and antioxidative defence, which could point towards a role in maintaining cell homeostasis under stress (Janz et al. 2012).
The impact of drought on tree growth and the plasticity of wood formation are well known phenomena; it was shown that trees adjust the organization of vascular tissues in response to drought during early and late summer, by modulating xylem cell number, shape and size, and that seasonality has an influence on the response, with early summer being more critical (Arend and Fromm 2007). This might be due to the different susceptibility of cambial cells to drought during early and late summer, which could in part be mediated by ABA (Arend and Fromm 2007). A different strategy is adopted by Scots pine: these trees respond to drought by building larger water conducting cells with thinner walls, despite the higher vulnerability to cavitation of such vessels (Eilmann et al. 2011).
Further insights into the response of trees to drought come from proteomic data which reported that in poplar subjected to drought stress, the cambial proteome reacted later than the leaf proteome, and that it showed variations in bark storage and cytoskeleton-associated proteins (actin and tubulin) (Durand et al. 2011).
Anthropogenic effects on wood formation
The speed of global urbanization and industrialization is an ever-increasing source of concern because of the impact on agriculture and forest productivity. Heavy metal stress, CO2 and ozone represent three of the most important anthropogenic factors influencing plant production. Heavy metals interfere with lignification by enhancing it (Moura et al. 2010), as recently shown in poplar exposed to Cd (Elobeid et al. 2012). Metals alter xylem hydraulic conductivity, as for instance recently reported in red maple seedlings (de Silva et al. 2012). A proteomic study carried out on leaf and cambial tissues of hybrid poplar subjected to acute metal stress showed a reduction in the abundance of cell-wall-related proteins (Durand et al. 2010). The abundance of rhamnose metabolism-related proteins decreased upon Cd stress, similarly to what was observed for two enzymes involved in the phenylpropanoid pathway, phenylcoumaran benzylic ether reductase and phenylpropanoid cinnamoyl-CoA reductase (Durand et al. 2010).
The analysis of the response triggered in trees by elevated CO2 concentrations is very important, given the alarming rise in its concentrations registered in the atmosphere. Wood density was shown to increase in response to elevated CO2 concentrations in cottonwood and a trend towards decreased fibre length and diameter was experimentally observed (Druart et al. 2006). Trees respond to CO2 by regulating the activity of genes. In particular, the study by Druart et al. (2006) indicated a different regulation in cottonwood source and sink tissues in response to increasing CO2 concentration. While in the leaves cell growth and flavonol metabolism were promoted, they were repressed in stems, where instead genes linked to lignin biosynthesis were upregulated. Transcripts responsible for monolignol biosynthesis, lignin polymerization and ethylene biosynthesis were indeed induced (Druart et al. 2006). The increase in the expression of those genes was positively correlated with increasing CO2 concentrations, while, interestingly, genes involved in cell wall formation and cell growth were downregulated (Druart et al. 2006). The repression of genes involved in wall formation correlates well with the observed tendency towards shorter fibres and the upregulation of lignin-related genes supports the higher wood density observed in the stems of cottonwood subjected to increased CO2 concentration. From the above-mentioned data, it is possible to confirm the influence that CO2 exerts on cell wall and ultimately wood biosynthesis. What emerges from these analyses is that the response of trees to CO2 is heterogeneous and follows a tissue-specific response.
Ozone has been shown to interfere with both lignin and cellulose biosynthesis (Richet et al. 2011): it is known that it triggers the formation of condensed lignins in poplar leaves and that these ‘stress lignins’ might account for tolerance, by creating a ‘barrier’ and an antioxidant effect to counteract reactive oxygen species (Cabané et al. 2004, Renaut et al. 2009).
In artificially tilted hybrid poplar, ozone triggers the formation of fewer but broader vessels in the stems; interestingly these anatomical differences are accompanied by a decrease in enzyme activities and the expression of genes linked to cellulose and lignin metabolism (Richet et al. 2011). A study focused on the action of elevated CO2 and ozone levels in poplar showed that both triggered a relative increased lignification of cell walls, but through different mechanisms. While the first treatment increased the overall C supply, the second caused a concomitant reduction in lignin, cellulose and hemicellulose, with lignin however being less affected and compensating for the decrease in the other cell wall components (Richet et al. 2012).
Conclusions
Wood biosynthesis represents one of the most relevant processes for plant development and for industrial applications. Acquiring an integrated view of this process will enable a thorough understanding of C flow dynamics within forest ecosystems. Moreover, the targeted study of the effects that exogenous stimuli exert on cambial activity and consequently on wood deposition can inspire not only more efficient strategies for boosting woody biomass yield, but also the design of engineering approaches aimed at reducing the lignin content without dramatically affecting the vascular tissue of the plant and its hydraulic properties.
Conflict of interest
None declared.
Funding
Partial financial support was obtained through the FNR Project CANCAN C13/SR/5774202.
Acknowledgments
The conception of this review is linked to activities conducted within the COST FP1105 and COST FP1106 actions. The authors apologize for not having mentioned all the relevant literature available in the field, because of space limitations. They wish to thank Laurent Solinhac and Célia Bayard for the microscopy images, and Dr Fuad Bahram for drawing Figure 5.