-
PDF
- Split View
-
Views
-
Cite
Cite
Bezawit A Woldemeskel, Caroline C Garliss, Joel N Blankson, mRNA Vaccine-Elicited Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2)–Specific T Cells Persist at 6 Months and Recognize the Delta Variant, Clinical Infectious Diseases, Volume 75, Issue 1, 1 July 2022, Pages e898–e901, https://doi.org/10.1093/cid/ciab915
Close - Share Icon Share
Abstract
Little is known about the decay kinetics of coronavirus disease 2019 vaccine–elicited severe acute respiratory syndrome coronavirus 2–specific T cells. In this study we show a modest decline in the frequency of these T cells at 6 months and demonstrate robust expansion in response to antigen and recognition of spike peptides from the Delta variant.
Recent studies have shown a decline in severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)–specific neutralizing antibody titer 6 months after receipt of mRNA vaccines [1]. In contrast, little is known about the rate of decay of vaccine-elicited T cells. T cells protect against reinfection in macaques [2] and have been shown to recognize the Alpha and Beta variants as effectively as the vaccine strain [3–5], thereby possibly providing protection against severe disease. Given the important role of these cells, it will be important to determine how long they persist after vaccination. In this study, we compared the frequency of SARS-CoV-2–specific T-cell responses at 2 weeks and at 6 months postvaccination and measured the response of T cells to Delta variant spike peptides at 6 months. Our results have implications for vaccine boosting strategies.
METHODS
We obtained blood from 21 study participants (13 men, 8 women). For 15 of these participants (10 men, 5 women), we obtained blood prior to vaccination and postvaccination at 7–14 days and 6 months after the second dose of the vaccine. Fourteen of these participants received the Pfizer-BioNTech (BNT162b2) vaccine and 1 received the Moderna (mRNA-1273) vaccine. The median age of the participants was 41 years (range: 23 to 56 years) (Supplementary Table 1). Informed consent was obtained from all study participants. Peripheral blood mononuclear cells (PBMCs) were isolated from whole blood using Ficoll centrifugation. Plasma was used to screen for natural infection with SARS-CoV-2 by testing for antibodies to the nucleocapsid protein with the Bio-Rad Platelia SARS-CoV-2 Total Antibody assay (Marnes-la-Coquette, France). We determined cellular immunity to the SARS-CoV-2 spike protein by performing an interferon-γ (IFN-γ) Elispot assay with unfractionated PBMCs that were stimulated with a pool of overlapping SARS-CoV-2 spike peptides (BEI, Manassas, VA), as previously described [3]. The assay was also performed with CD8+ T-cell–depleted PBMCs to determine the relative contribution of CD4+ T cells and CD8+ T cells to the cellular immune response. To compare recognition of spike proteins from the vaccine strain and the Delta variant, we stimulated PBMCs with overlapping spike peptide pools from both viruses at a concentration of 1 μg/mL (JPT, Berlin, Germany). To determine how effectively SARS-CoV-2–specific T cells proliferate in response to the vaccine strain spike peptide, PBMCs were stimulated with overlapping peptides or media alone in standard media with 10 units/mL of interleukin 2 (IL-2) for 10–12 days and then washed and rested overnight in media alone for 24 hours. The cells were then stimulated for 12 hours in the presence of protein transport inhibitors (GolgiPlug, 1 μg/mL; GolgiStop, 0.7 μg/mL) and antibodies against CD28 and CD49d (all from BD Biosciences, Franklin Lakes, NJ) and then stained with annexin V (BD Biosciences) and antibodies to CD3 and CD4 (Biolegend) and then fixed and permeabilized and stained with antibodies to IFN-γ and tumor necrosis factor α (TNF-α; both from BD Biosciences). Flow cytometry was performed on a BD FACS LSRFortessa flow cytometer, and data were analyzed using FlowJo software, version 10. Data on a minimum of 100000 events in the lymphocyte gate were collected and analyzed.
All statistical analyses were performed using GraphPad Prism Software (version 9.2.0). For experiments requiring comparisons between 2 groups, a 2-tailed paired Student’s t test was used to determine significance. For experiments requiring multiple comparisons, a Friedman test with Dunn’s multiple-comparison test was used. A P value of less than .05 was considered statistically significant.
RESULTS
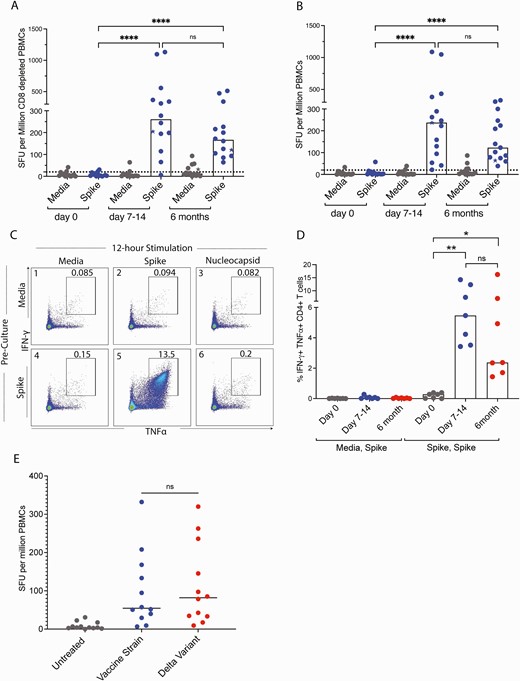
We previously determined that the frequency of T cells that recognized spike peptides was extremely low prior to vaccination, with a median of 2.7 spot forming units (SFU) per million PBMCs [3]. This frequency had increased to a median of 237 SFU per million PBMCs at day 7 to day 14 postvaccination [3]. At 6 months, the SFU had decreased to 122 SFU per million PBMCs, a number that was still significantly higher than the prevaccination level (day 0) (Figure 1A and Supplementary Table 2). In order to determine whether these responding cells were predominantly CD4+ or CD8+ T cells, we depleted CD8+ T cells from PBMCs and repeated the Elispot assay. As reported earlier, depletion of CD8+ T cells resulted in higher responses, suggesting that the responses were predominantly driven by CD4+ T cells. There was a median of 260 SFU per million cells at 7 to 14 days postvaccination [3], and at 6 months the response had declined to 166 SFU per million cells (Figure 1B and Supplementary Table 3). This number was significantly higher than the prevaccination level and not statistically different from the frequency present at days 7 to 14. We asked whether the cells present at the 6-month time point were capable of proliferating in response to stimulation with the spike peptides. To do this, we cultured PBMCs with spike peptides or media alone for a 10- to 12-day period to allow for the cells to proliferate, followed by restimulation with the peptide for 12 hours in order to induce cytokine expression. mRNA for the nucleocapsid gene is not included in the vaccines; therefore, we used a pool of peptides from this protein as a specificity control. As shown in Figure 1C, there was a low frequency of CD4+ T cells that co-expressed TNF-α and IFN-γ in response to spike peptides after the cells had been cultured with media alone (median of 0.01%). In contrast, after stimulation with spike peptides for 10 days, there was a marked increase in the frequency of spike-specific CD4+ T cells (median of 2.37%). There was no significant difference in the frequency of expanded cells that were detected at days 7–14 and 6 months postvaccination (Figure 1D). Finally, we asked whether the spike-specific cells present at 6 months postvaccination would also recognize spike proteins from the Delta variant virus, which is currently the predominant circulating virus. As shown in Figure 1E, there was no significant difference in the frequency of T cells that recognized spike peptide pools from the vaccine strain versus the Delta variant (median of 54.6 SFU vs 82 SFU per million PBMCs, respectively).
The number of SFU per million cells generated in response to stimulation with spike peptides is shown at different time points for unfractionated PBMCs (A) and CD8-depleted PBMCs (B). Values from study participants who received the BNT162b2 vaccine are shown as circles and values from the study participant who received the mRNA-1273 are shown as a star. Horizontal bars represent the median value. (C) Flow cytometry plot showing co-expression of TNF-α and IFN-γ of CD4+ T cells after 10–12-day preculture with either media alone or with SARS-CoV-2 spike peptides followed by 12-hour stimulation with media alone, spike peptides, or nucleocapsid peptides. (D) The percentage of cells co-expressing both cytokines in response to media preculture followed by spike peptide stimulation (media, spike) versus spike peptide preculture and spike peptide stimulation (spike, spike) at different time points. (E) The number of SFU generated in response to stimulation with spike peptides from the vaccine strain or Delta variant viruses. 0.1234 (ns), 0.0332 (∗), 0.0021 (∗∗), 0.0002 (∗∗∗), <0.0001 (∗∗∗∗). Abbreviations: IFN-γ, interferon-γ; ns, nonsignificant; PBMC, peripheral blood mononuclear cell; SARS-CoV-2, severe acute respiratory syndrome coronavirus 2; SFU, spot forming units; TNFα, tumor necrosis factor α.
DISCUSSION
Recent studies have shown a decline in the titer of neutralizing antibodies 6 months after vaccination [1]. Furthermore, the Delta variant, which is the predominant variant of concern currently in circulation, is not fully neutralized by antibodies generated by current mRNA vaccines [6, 7]. These 2 factors may partially explain breakthrough infections of vaccinated individuals, especially given the correlation of breakthrough infections with lower neutralizing antibody titers [8].
mRNA vaccines elicit strong SARS-CoV-2–specific T-cell responses [9, 10]. Studies have suggested that SARS-CoV-2–specific T cells play a protective role in natural infection and in vaccinated individuals [11]. Direct evidence for a role of T cells in protection comes from studies showing that depletion of CD8+ T cells in convalescent rhesus macaques partially abrogates the resistance of these animals to SARS-CoV-2 reinfection [2]. However, it is not known how long vaccine-induced T cells remain in circulation. Naive T cells normally expand in response to recognition of cognate antigen and develop into effector and memory T cells. The frequency of effector T cells will decline with time, but a low percentage of memory cells will persist and expand if the same antigen is encountered again. Here we showed a modest decline in total and SARS-CoV-2–specific T cells 6 months after vaccination. These cells expanded to much higher frequencies following a 10-day culture period, which is consistent with a memory T-cell phenotype. We also show that vaccine-generated T cells recognize spike peptides from the Delta variant as efficiently as spike peptides from the vaccine strain, which is consistent with prior studies showing that T cells recognize many epitopes in the spike protein that are not affected by mutations present in the Alpha and Beta variants [3–5]. The persistence of T cells that can recognize variants of concern could potentially explain the protection from severe disease that has been reported in vaccine breakthrough infections [12].
Our study is limited by a relatively small number of study participants, but by analyzing longitudinal samples we show preservation of T-cell responses in vaccinated individuals at 6 months and we also demonstrate efficient T-cell recognition of the Delta variant spike peptides. Our data further the understanding of the immune response to the SARS-CoV-2 mRNA vaccines. The robust expansion of T cells in response to stimulation with spike peptides suggests that booster shots should successfully increase the frequency of SARS-CoV-2–specific T cells in circulation.
Supplementary Data
Supplementary materials are available at Clinical Infectious Diseases online. Consisting of data provided by the authors to benefit the reader, the posted materials are not copyedited and are the sole responsibility of the authors, so questions or comments should be addressed to the corresponding author.
Notes
Acknowledgments. We thank Evan Beck for performing the Bio-Rad Platelia SARS-CoV-2 Total Antibody assay and Saphira Cherfils (Hunter College, City University of New York) for her input.
Financial support. This work was supported by the Johns Hopkins COVID-19 Vaccine-related Research Fund.
Potential conflicts of interest. The authors: No reported conflicts of interest. All authors have submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest. Conflicts that the editors consider relevant to the content of the manuscript have been disclosed.
References
Author notes
B. A. W. and C. C. G. contributed equally to this manuscript.





Comments