-
PDF
- Split View
-
Views
-
Cite
Cite
Que Kong, Wei Ma, Haibing Yang, Guojie Ma, Jenny J Mantyla, Christoph Benning, The Arabidopsis WRINKLED1 transcription factor affects auxin homeostasis in roots, Journal of Experimental Botany, Volume 68, Issue 16, 3 October 2017, Pages 4627–4634, https://doi.org/10.1093/jxb/erx275
Close - Share Icon Share
Abstract
WRINKLED1 (WRI1) is a key transcriptional regulator of fatty acid biosynthesis genes in diverse oil-containing tissues. Loss of function of Arabidopsis WRI1 leads to a reduction in the expression of genes for fatty acid biosynthesis and glycolysis, and concomitant strong reduction of seed oil content. The wri1-1 loss-of-function mutant shows reduced primary root growth and decreased acidification of the growth medium. The content of a conjugated form of the plant growth hormone auxin, indole-3-acetic acid (IAA)-Asp, was higher in wri1-1 plants compared with the wild-type. GH3.3, a gene encoding an enzyme involved in auxin degradation, displayed higher expression in the wri1-1 mutant. EMSAs demonstrated that AtWRI1 bound to the promoter of GH3.3. Specific AtWRI1-binding motifs were identified in the promoter of GH3.3. In addition, wri1-1 displayed decreased auxin transport. Expression of some PIN genes, which encode IAA carrier proteins, was reduced in wri1-1 plants as well. Correspondingly, AtWRI1 bound to the promoter regions of some PIN genes. It is well known that auxin exerts its maximum effects at a specific, optimal concentration in roots requiring a finely balanced auxin homeostasis. This process appears to be disrupted when the expression of WRI1 and in turn a subset of its target genes are misregulated, highlighting a role for WRI1 in root auxin homeostasis.
Introduction
The growth regulator auxin (indole-3-acetic acid; IAA) plays an important role in plant morphogenesis and development such as growth of the shoot and root, hypocotyl elongation, etc. (Zhao et al., 2001; Takase et al., 2004; Woodward and Bartel, 2005). As part of auxin-mediated signaling pathways, expression of numerous genes is affected by tissue auxin concentration (Wang et al., 2010). Auxin signaling marker genes (auxin-responsive genes) have been classified into three major categories: Aux/IAA genes, SAUR genes (small, auxin-induced RNA), and GH3 genes (Ulmasov et al., 1995; Woodward and Bartel, 2005; Khan and Stone, 2007; Zhang et al., 2007).
Expression of GH3 genes is generally associated with plant development (e.g. root and hypocotyl growth). Nakazawa et al. (2001) found that Arabidopsis DFL1 (GH3.6) affects plant morphogenesis including the development of shoot, root, and hypocotyl (Nakazawa et al., 2001). GH3.9 was found to affect the growth of the primary root, and Arabidopsis (Arabidopsis thaliana) gh3.9 (a GH3.9 loss-of-function mutant) displays an elongated root compared with wild-type (WT) seedlings (Tatematsu et al., 2008). Arabidopsis GH3.5 (WES1) was found to mediate auxin homeostasis, and overexpression of GH3.5 led to reduced IAA levels, reduced plant growth, and disturbed adventitious root development (Park et al., 2007). GH3.2 (YDK1) overexpression in transgenic Arabidopsis plants also displayed reduced primary root growth and short hypocotyls (Takase et al., 2004).
Recently, several GH3 genes were found to encode enzymes which form IAA-amino acid conjugates (Staswick et al., 2002, 2005). Mutations of two GH3 genes in Physcomitrella patens led to elevated free auxin content and enhanced sensitivity to auxin application (Ludwig-Muller et al., 2009). This evidence suggests that GH3 proteins play a role in balancing auxin pools, which also supports the notion that the role of GH3 proteins is evolutionarily conserved in mediating auxin homeostasis (Chapman and Estelle, 2009).
GH3 genes have also been studied with regard to their transcriptional regulation by promoter analysis and identification of the transcription factors responsible. The majority of GH3 genes contain auxin-responsive elements (AuxREs) and other cis-elements associated with stresses and other hormone responses (Woodward and Bartel, 2005; Ostrowski and Jakubowska, 2013). DFL1 (GH3.6), AtGH3a, and GH3.17, but not GH3.3, are the targets of the transcription factor ARF8 (Tian et al., 2004). The Arabidopsis MYB77 transcription factor affects expression of GH3.2 and GH3.3 (Shin et al., 2007). The tobacco bZIP transcription factor NtBZI-1 binds to a G-box-related element (GRE) in the promoter region of NtGH3 which affects the expression of NtGH3 (Heinekamp et al., 2004).
Members of the PIN (PIN-FORMED) proteins have been known to play an important role in auxin transport (Galweiler et al., 1998; Luschnig et al., 1998; Zazimalova et al., 2010). Both environmental cues and plant hormones are found to be crucial factors in PIN expression (Peer et al., 2004; Kimbrough et al., 2005; Vieten et al., 2005; Keuskamp et al., 2010; Sassi et al., 2012). Although PIN expression has been investigated broadly (Aida et al., 2004; Vieten et al., 2005), not much is known regarding the transcription factors which directly control the expression of PIN genes. Recently, an Arabidopsis MADS transcription factor, XAL2, has been identified to mediate auxin transport by regulation of PIN1 and PIN4 (Garay-Arroyo et al., 2013). The Arabidopsis MYB88 transcription factor was recently found to control the expression of PIN3 and PIN7 directly (Wang et al., 2015). The Arabidopsis transcription factors IDD14 and IDD16 have been found to activate the expression of PIN1 (Cui et al., 2013).
Arabidopsis WRINKLED1 (WRI1) is an APETALA2 (AP2) transcription factor (Cernac and Benning, 2004; Masaki et al., 2005). The wri1-1 mutant displays an ~80% reduction of seed oil content (Focks and Benning, 1998) and decreased expression of genes encoding fatty acid and glycolytic enzymes (Ruuska et al., 2002). Overexpression of AtWRI1 led to increased expression of de novo fatty acid biosynthesis genes (Sanjaya et al., 2011) and increased oil content in seeds and leaves of transgenic Arabidopsis plants (Cernac and Benning, 2004; Sanjaya et al., 2011). The AW-box has been characterized as an AtWRI1-binding motif which is conserved in promoter regions of AtWRI1 target fatty acid biosynthesis genes (Maeo et al., 2009). WRI1 orthologs have been found to affect expression of de novo fatty acid biosynthesis genes and oil biosynthesis in different plant species (Liu et al., 2010; Shen et al., 2010; Pouvreau et al., 2011; Ma et al., 2013; Wu et al., 2014; Yang et al., 2015). Transient expression of AtWRI1 and WRI1 orthologs in tobacco leaves led to oil production (Vanhercke et al., 2013; Grimberg et al., 2015; Ma et al., 2015). AtWRI1 is targeted by the 26S proteasome-mediated degradation pathway (Chen et al., 2013), and the PEST motif in the C-terminal intrinsically disordered region (IDR) of AtWRI1 affects the stability of AtWRI1 and oil production (Ma et al., 2015). 14-3-3 proteins were recently found to interact with AtWRI1 and affect AtWRI1 transcriptional activity and oil biosynthesis (Ma et al., 2016).
In this study, we describe that Arabidopsis wri1-1 plants display altered development of the primary root. Auxin content and the expression of GH3 genes are affected in wri1-1. Notably, we found that AtWRI1 binds to the promoter of GH3.3 (proGH3.3). We identified a putative binding element for AtWRI1 located in proGH3.3 different from the known cis-element (AW-box) (Maeo et al., 2009).
Materials and methods
Plant materials
Plant materials and growth conditions have been previously described (Ma et al., 2013, 2015). Transgenic wri1-1 lines overexpressing an AtWRI1-TAP (line #6-5 and #7-2) were previously described (Ma et al., 2013). Seed sterilization followed the method described previously (Ma et al., 2013).
Growth medium pH assay
The pattern of pH change in the growth medium around the roots of Arabidopsis seedlings was determined as previously published (Ma et al., 2006; Yang et al., 2007). Arabidopsis seedlings were first grown vertically on normal medium for 3 d before being transferred to growth medium containing the pH indicator, bromocresol purple. Images were taken 7 d after the seedlings were transferred to pH indicator growth medium.
Quantification of IAA and IAA-Asp in Arabidopsis
One-week-old Arabidopsis seedlings grown vertically on normal medium were harvested (~100 mg FW per sample) and immediately frozen in liquid nitrogen. Samples were sent to the Danforth Center Proteomics & Mass Spectrometry core facility for acidic plant hormone analysis. The detailed protocol can be found at their website (http://www.danforthcenter.org/docs/default-source/CoreFacilities/pmsf/plant-hormone-analysis_customer-instructions_2014.pdf?sfvrsn=0, last accessed 27 July 2017).
Protein expression and EMSA
AtWRI158–240 was subcloned into the pET41a-6×His vector. The primers used for subcloning AtWRI158–240 were 5'-GCAGGATCC GCTTCTACCCGA-3' (forward) and 5'-TAACTCGAGTTACG GGAAAACACC-3' (reverse). Protein induction, extraction, and purification were performed as previously published (Kong et al., 2012; Ma et al., 2015). EMSAs were performed according to Kong et al. (2012).
RNA extraction and quantitative real-time PCR
Methods for RNA extraction, cDNA synthesis, and quantitative real-time PCR (qRT-PCR) were as previously described (Ma et al., 2015). The gene expression level was normalized to an internal standard, the Protein Phosphatase 2A (PP2A) gene. The primers used for qRT-PCR are described in Supplementary Table S1 at JXB online. The primers used for PP2A were previously published (Czechowski et al., 2005). The primers used for GH3.2, GH3.3, GH3.4, GH3.5, GH3.6, and GH3.7 were according to González-Lamothe et al. (2012), and the primers for ARF6, ARF8, and ARF17 were according to Gutierrez et al. (2009).
[3H]IAA transport assays
Ten-day-old seedlings were used for [3H]IAA transport assays. The assays were as described in Blakeslee et al. (2007).
Results
AtWRI1 affects Arabidopsis root development
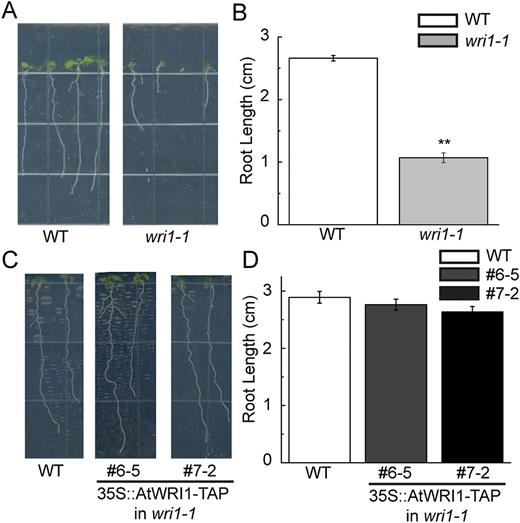
The AtWRI1 transcript is most abundant in the embryo and has generally low abundance in vegetative tissues such as leaves (Cernac and Benning, 2004). However, it should be noted that AtWRI1 has high expression in tissues such as ovaries and roots, as determined by microarray analysis (Supplementary Fig. S1). Northern blots also detected AtWRI1 at high levels in roots and flowers (Cernac and Benning, 2004). However, there is currently no report regarding wri1-related mutant phenotypes in either Arabidopsis ovaries or roots. Here, we observed the root morphology of the wri1-1 mutant and determined a possible AtWRI1 function in roots. The wri1-1 mutant displayed reduced root length compared with WT seedlings (Fig. 1A, B). In order to test whether the root phenotype in the wri1-1 mutant was due to the mutation of AtWRI1, root length in wri1-1 transgenic plants overexpressing AtWRI1-TAP in lines #6-5 and #7-2 (Ma et al., 2013) was determined to test for restoration of this phenotype. As shown in Fig. 1C and D, expression of AtWRI1-TAP restored the root growth in the wri1-1 mutant.
Primary root length of wild-type (WT), wri1-1, an AtWRI1 loss-of-function mutant, and transgenic wri1-1 plants overexpressing AtWRI1-TAP (lines 6-5 and 7-2). Root growth (A) and root length measurement (B) of 1-week-old seedlings of the WT and wri1-1. Primary root length was typically measured on 36 seedlings to calculate a mean value. Results are shown as means ±SE (n=36). Root growth (C) and root length measurement (D) of 10-day-old seedlings of the WT and transgenic wri1-1 overexpressing AtWRI1-TAP (lines 6-5 and 7-2). Primary root length was typically measured on 31 seedlings for the WT or 22 seedlings for lines 6-5 and 7-2 to calculate a mean value. Results are shown as means ±SE (n=31 for the WT and n=22 for lines 6-5 and 7-2). Primary root length of the wri1-1 mutant displayed statistically significant differences compared with the WT (P<0.01, t-test), as indicated by **. (This figure is available in colour at JXB online.)
Previous studies indicated that proton (H+) efflux is correlated with root growth. For example, Arabidopsis mutants or transgenic plants with increased root growth have been found to show enhanced H+ efflux (Ma et al., 2006; Yang et al., 2007). Hence, we investigated the H+ efflux out of wri1-1 mutant roots employing a medium acidification assay. As shown in Supplementary Fig. S2, wri1-1 roots displayed decreased H+ efflux.
The wri1-1 mutant accumulates an auxin metabolite
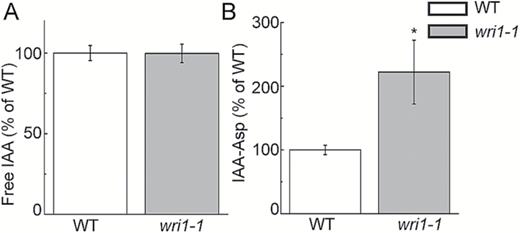
Suspecting that auxin homeostasis was altered, we quantified IAA metabolites (free IAA and IAA-Asp) using LC-MS/MS. These IAA metabolites have been shown to be associated with Arabidopsis root growth (Quint et al., 2009; Ludwig-Muller, 2011; Pencik et al., 2013). Our result indicated that free IAA was not altered in the wri1-1 mutant compared with the WT (Fig. 2A). However, tissue levels of IAA-Asp were higher in wri1-1 (Fig. 2B).
Quantification of IAA and IAA-Asp in WT and wri1-1 plants by LC-MS/MS (1-week-old seedlings). Results are shown as means ±SE (n=4–5). * indicates a significant difference (P<0.05, t-test) from the WT.
Expression of GH3 genes is altered in the wri1-1 mutant
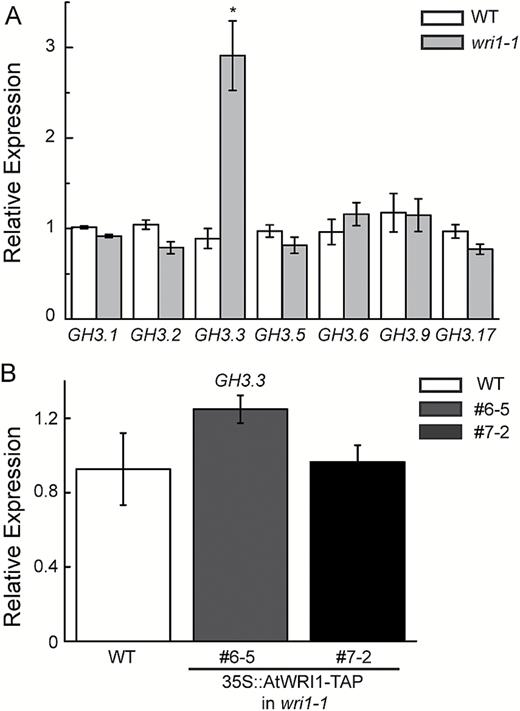
We hypothesized that AtWRI1 might play a role in modulating the expression of genes involved in auxin signaling. To test this hypothesis, we surveyed the transcriptional profiles of roots of wri1-1 and WT plants by microarray analysis. Notably, the expression of GH3.3, a member of the GH3 gene family of auxin-responsive genes, was elevated in the mutant (Supplementary Table S2). To validate the microarray data, we determined the expression of GH3.3 and other GH3 genes by qRT-PCR. The expression of GH3.3 was reproducibly elevated in wri1-1 (Fig. 3A). Overexpressing AtWRI1-TAP in wri1-1 reverted the increased expression of GH3.3 in wri1-1. The expression of GH3.3 in AtWRI1-TAP transgenic lines (#6-5 and #7-2) compared with WT plants was not significantly different (P>0.05, t-test) (Fig. 3B). This result indicates that the elevation of GH3.3 expression in the wri1-1 mutant is due to the mutation of AtWRI1. Expression of GH3.15 was also elevated in wri1-1 (Supplementary Fig. S3). However, expression of GH3.10, GH3.12, and GH3.13 in wri1-1 was reduced compared with the WT (Supplementary Fig. S3). In addition, we determined the expression of additional auxin-responsive genes of other families, including IAA7, IAA17, IAA28, ARF6, ARF8, and ARF17. Compared with WT plants, no obvious alteration was detected in the expression of the three IAA (Supplementary Fig. S4A) or ARF (Supplementary Fig. S4B) genes tested in the wri1-1 mutant.
Expression analysis of the genes encoding indole-3 acetyl-aspartate synthetases (in the GH3 gene family). (A) GH3 transcripts of WT and wri1-1 plants were measured by quantitative real-time PCR. Results are shown as means ±SE (n=3). * indicates a significant difference (P<0.05, t-test) compared with the WT. (B) GH3.3 transcripts of the WT and transgenic wri1-1 overexpressing AtWRI1-TAP (lines 6-5 and 7-2) were determined by quantitative real-time PCR. Results are shown as means ±SE (n=3).
AtWRI1 binds to the promoter of GH3.3
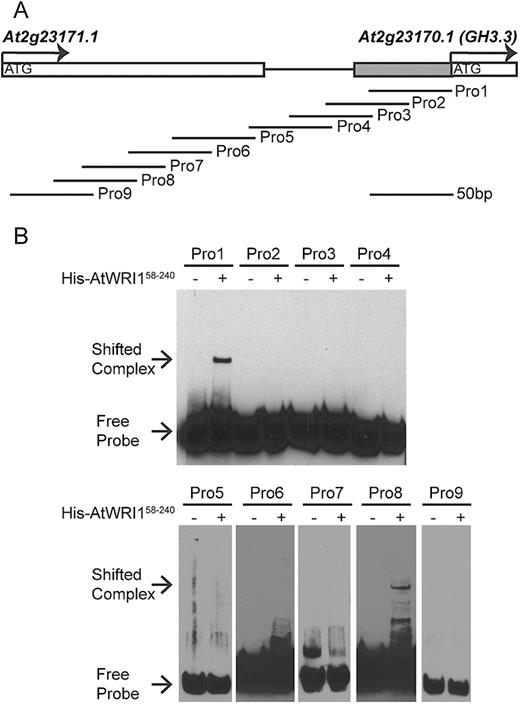
To understand how AtWRI1 affects the expression of the GH3.3 gene, we first performed bioinformatic analysis of the 1.0 kb promoter region of GH3.3 (proGH3.3). However, there was no recognizable AW-box in proGH3.3. To examine whether the AtWRI1 protein can bind to proGH3.3, we designed a series of DNA probes covering the region 268 bp upstream of the translational start site (Fig. 4A; Supplementary Fig. S5A). An EMSA indicated that the AtWRI1 protein was able to bind DNA probes Pro1 and Pro8 (Fig. 4B). Further alignment of the DNA sequences of Pro1 and Pro8 showed some similarity (Supplementary Fig. S5). Hence, they might contain the putative binding site(s) for AtWRI1 in proGH3.3.
AtWRI1 binds to the promoter of GH3.3. (A) Schematic diagram of the promoter region of the GH3.3 gene. A total of nine DNA probes have been designed from proGH3.3 for the EMSA. (B) Binding of the AP2 domain of AtWRI1 (amino acids 58–240) to various probes using EMSA.
Auxin sensitivity and transport are altered in the wri1-1 mutant
We also examined the root elongation of WT and wri1-1 seedlings in response to applied IAA. There was no apparent difference when seedlings were grown in the presence of high concentrations of IAA (10–7, 10–6, and 10–5 M). Both WT and wri1-1 seedlings showed similar inhibition in response to IAA (Supplemenatry Fig. S6). When seedlings were grown on medium with relatively low concentrations of IAA (10–9 M and 10–8 M), wri1-1 displayed more inhibition compared with the WT (Supplementary Fig. S6). This result suggests that wri1-1 had a slightly higher sensitivity to applied IAA at least at some concentrations. This result was somewhat surprising because mutants with increased GH3 gene expression and an elevated IAA-Asp level had been reported to display increased resistance to applied IAA (Nakazawa et al., 2001; Staswick et al., 2005; Park et al., 2007). We hypothesized that mutation of AtWRI1 might affect additional processes that mediate auxin homeostasis. Hence, we examined auxin transport in the WT and the wri1-1 mutant. As shown in Supplementary Fig. S7, auxin transport was reduced in the wri1-1 mutant. Compared with the WT, auxin transport was reduced 41% in roots of wri1-1.
Expression of PIN genes is altered in the wri1-1 mutant
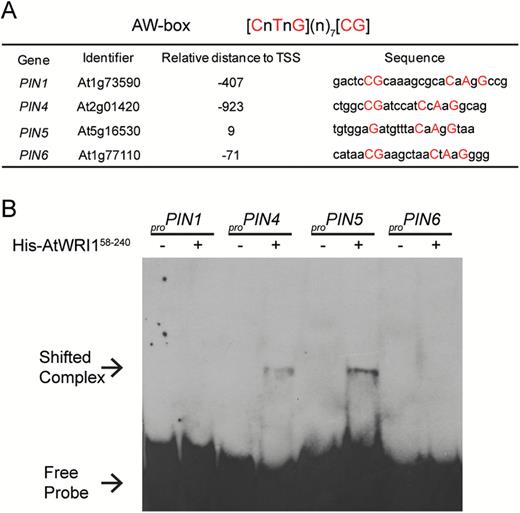
Previous studies have shown that expression of PIN genes is correlated with changes in auxin transport in Arabidopsis roots (Peer et al., 2004). Thus, we hypothesized that the decreased auxin transport that we observed in the wri1-1 mutant might be due to the altered expression of PIN genes. A qRT-PCR assay was performed, and the result indicated that the expression level of some PIN genes (PIN1, PIN3, PIN5, and PIN6) was reduced in wri1-1 (Supplementary Fig. S8). In addition, we searched the presumed promoter regions (1 kb upstream of the transcription start site) of PIN genes for the AtWRI1 binding motif AW-box (Maeo et al., 2009). An AW-box (or AW-box-like) motif was identified in the 1 kb sequences 5' of the transcriptional start of PIN1, PIN4, PIN5, and PIN6 (Fig. 5A). Therefore, AtWRI1 might control the expression of PIN genes through binding to the AW-box motif. To test this hypothesis further, we performed EMSA experiments to probe whether AtWRI1 binds to proPIN genes. The EMSA result indicates that AtWRI1 was able to bind to the AW-box within the context of the presumed proPIN4 and proPIN5 (Fig. 5B), but AtWRI1 did not appear to bind to the presumed proPIN1 and proPIN6 sequences (Fig. 5B).
Examination of AtWRI1 binding to promoters of PIN genes. (A) In silico analysis of the AW-box in the promoter region of PIN genes. The AtWRI1-binding site AW-box was identified by AthaMap (http://www.athamap.de/index.php, last accessed 27 July 2017). The region from 1 kb upstream of the TSS (transcription start site) to 100 bp downstream of the TSS was screened for each gene. Nucleotides that match to the AW-box are shown with upper case letters. (B) Binding of the AP2 domain of AtWRI1 (amino acids 58–240) to the AW-box or AW-box-like sequence in the promoters of PIN1, PIN4, PIN5, and PIN6. The sequences of probes used in EMSA are listed in (A). (This figure is available in colour at JXB online.)
Discussion
Results presented above suggest that AtWRI1 plays a role in mediating auxin homeostasis by binding to the promoter of the auxin-responsive gene GH3, providing a molecular mechanism for how AtWRI1 might affect auxin homeostasis. It should be noted that AtWRI1 binds to proGH3.3 at a non-canonical motif. In addition, AtWRI1 might play a role in controlling the expression of auxin carrier genes by binding to the canonical AtWRI1 cis-element AW-box in the regulatory regions of select PIN genes.
GH3.3 is a genuine target gene of the transcription factor AtWRI1
Our data suggest that AtWRI1 is a transcriptional repressor for GH3.3. Previous studies have indicated that GH genes have potential roles in mediating auxin homeostasis and root development (Park et al., 2007; Gutierrez et al., 2012). Thus, aside from the well-established function of AtWRI1 in oil biosynthesis and the regulation of expression of fatty acid biosynthesis genes, AtWRI1 has a set of target genes probably unrelated to oil biosynthesis, which could cause potential side effects during the WRI1-based engineering of oil content.
A 50 bp DNA fragment in proGH3.3 is critical for AtWRI1 binding to proGH3.3 (Fig. 4). Further in silico analysis indicated that it is different from AuxREs or AW-box elements. Therefore, AtWRI1 binds to this cis-element in the auxin-responsive gene GH3.3 in addition to its binding to the classical AtWRI1 AW-box-binding motif in genes related to fatty acid metabolism (Maeo et al., 2009).
Expression of GH3.3 was increased when the AtWRI1 protein was lost (wri1-1; Fig. 3A). In Arabidopsis, it has been suggested that GH3.3, GH3.5, and GH3.6 have functional redundancy with regard to their effects on lateral root growth (Gutierrez et al., 2012). We did not observe lateral root phenotypes in the wri1-1 mutant (Fig. 1A). Thus, it was unclear whether the increased transcript of GH3.3 in wri1-1 led to the reduced root growth of the wri1-1 plants. However, the endogenous level of IAA-Asp was elevated in wri1-1 compared with the WT (Fig. 2B). GH3.3 was suggested to be involved in auxin homeostasis including the biosynthesis of IAA-Asp (Gutierrez et al., 2012). Therefore, we suggest that this might be the possible molecular mechanism explaining the reduced root growth that we observed in wri1-1 plants.
In addition to the altered expression of GH3.3, wri1-1 also had changes in the expression of other GH3 genes such as GH3.10, GH3.12, GH3.13, and GH3.15 (Supplementary Fig. S3). GH3.10 antisense transgenic plants show primary root growth inhibition only under certain light conditions (Takase et al., 2003). The roles of GH3.12, GH3.13, and GH3.15 in root growth and auxin homeostasis still remain unknown at present. Therefore, we cannot rule out that the alterations of the expression of these GH3 genes also result in modulating auxin homeostasis, which in turn affects the primary root growth of wri1-1 plants.
PIN genes might be another set of AtWRI1 target genes
Plant transcription factors such as XAL2, IDD14/15/16, and MYB88 have been shown to play an important role in controlling the expression of PIN genes (Cui et al., 2013; Garay-Arroyo et al., 2013; Wang et al., 2015). Here, our data suggest that AtWRI1 might also be a transcription factor mediating the expression of PIN genes. Expression of four PIN genes was reduced in wri1-1 compared with the WT (Supplementary Fig. S8), which might explain why wri1-1 displayed reduced auxin transport in roots. In silico analysis (Fig. 5A) revealed that the AW-box (or AW-box like) motif was found in the promoter regions of three PIN genes (PIN1, PIN5, and PIN6) also showing reduced expression in wri1-1 plants (Supplementary Fig. S8), which suggests that PIN1, PIN5, and PIN6 might be AtWRI1 target genes. Although the AW-box in proPIN6 matches the AW-box identified previously (Maeo et al., 2009), we did not detect the AtWRI1 binding to the AW-box in proPIN6 in the EMSA (Fig. 5B). It has been suggested that there is a 7 bp random sequence within the AW-box (Maeo et al., 2009). Thus, the difference of a 7 bp random sequence between proPIN6 and promoters of AtWRI1 classical target genes (e.g. BCCP2) may cause the altered binding affinity of AtWRI1 in this case. Regarding the AW-box-like motif in proPIN1, it contains a 9 bp random sequence instead of 7 bp, which might explain also why AtWRI1 did not bind to proPIN1 (Fig. 5B). Although AtWRI1 bound to the AW-box in the presumed proPIN4 (Fig. 5B), it should be noted that the AW-box-like motif in the presumed proPIN4 was located ~1 kb upstream from the transcriptional start site (Fig. 5A) and probably too far away to be directly involved in the regulation of PIN4 expression. Previously identified AW-boxes in promoter regions of AtWRI1 target genes are usually 300 bp upstream from the transcriptional start site (Maeo et al., 2009), which might explain why expression of PIN4 was not altered in the wri1-1 mutant compared with the WT (Supplementary Fig. S8). In addition, imaging of proPIN4::PIN4-GFP or immunolocalization of PIN4 protein in roots of wri1-1 plants might need to be pursued in the future to clarify the effect of WRI1’s mutation on the expression of PIN4. There is no AW-box in the presumed proPIN3, so AtWRI1 might bind to this promoter through an as yet unknown cis-element. In addition, a previous study indicated that PIN5 plays a role in mediating auxin homeostasis. Induction of PIN5 expression in BY-2 cells results in reduced free IAA and increased IAA-Asp contents (Mravec et al., 2009). In line with that, the endogenous IAA-Asp content was increased in wri1-1 compared with the WT (Fig. 2B), while expression of PIN5 was reduced in wri1-1 (Supplementary Fig. S8). Taken together, mutation of AtWRI1 led to expression changes in the genes encoding both protein families related to auxin homeostasis, GH3 and PIN genes, probably leading to altered IAA metabolite levels and IAA sensitivity, and ultimately the observed root growth defect in the wri1-1 mutant. While we do not yet fully understand how the misregulation of these genes causes the observed growth phenotype, recognizing the mere fact that WRI1 affects hormone homeostasis is of importance for the engineering of crops for increased oil content using the WRI1 transcription factor in the future.
Supplementary data
Supplementary data are available at JXB online.
Fig. S1. Expression profiles of AtWRI1 in Arabidopsis root.
Fig. S2. Growth phenotype of the WT and wri1-1 on pH indicator plates.
Fig. S3. Gene expression of GH3 genes of unknown function.
Fig. S4. Gene expression of IAA and ARF auxin- responsive genes.
Fig. S5. Probes for the promoter of GH3.3 used in EMSA.
Fig. S6. Primary root growth inhibition in response to different concentrations of IAA.
Fig. S7. Auxin transport in roots of the WT and wri1-1.
Fig. S8. Expression of PIN genes.
Table S1. Quantitative real-time PCR primers used in this study.
Table S2. Genes expressed differently in roots of wri1-1 compared with WT plants in microarray analysis.
Acknowledgements
We thank Dr Curtis Wilkerson (Michigan State University) for microarray analysis and Dr Yang Yang (Michigan State University) for assistance with taking pictures of root growth. We thank Anna Hurlock (Michigan State University) for critical reading of the manuscript. We also thank the proteomics and mass spectrometry facility at the Danforth Center for acidic plant hormone analysis using the 4000 QTRAP LC-MS/MS system, which is based upon work supported by the National Science Foundation under grant no. DBI-0521250 for acquisition of the QTRAP LC-MS/MS. This work was supported by the Department of Energy–Great Lakes Bioenergy Research Center Cooperative Agreement DE-FC02-07ER64494 (CB).
References
Author notes
These authors contributed equally to this work.




Comments