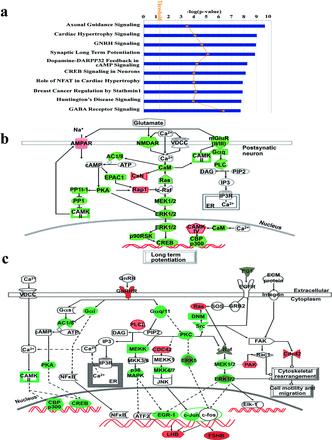
Pathway analyses of the neonatally-thimerosal-dysregulated genes in adult male mouse brains. The pathway data represent the logarithm of p-values calculated by Fisher's exact test, with a threshold for statistical significance, p < 0.05. Canonical pathways with p < 0.05 were defined as significant. Genes involved in each pathway were provided in Supplementary table 7. (a) Top 10 canonical pathways affected by up-/down-regulated genes/isoforms in thimerosal-treated male mice brain samples. The pathway analysis of genes that affected by thimerosal was identified by Ingenuity Pathway Analysis, and the expanded list of the top 10 canonical pathways were presented in the supplementary data (Supplementary table 7). (b and c) Synaptic long-term potentiation pathway and GNRH signaling pathway. Dysregulated genes in neonatally thimerosal-treated male mice brains are enriched in synaptic long-term potentiation pathway (b) and GNRH signaling pathway (c). Color indicates the strength of fold change of the indicated gene expression. Red represents the gene expression increase in thimerosal-treated mouse brains; Green represents gene expression decrease in thimerosal-treated mice brains. Each gene involved in the signaling pathways is listed in Supplementary table 7.